
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 4,317,899 – 4,318,169 |
| Length | 270 |
| Max. P | 0.909575 |

| Location | 4,317,899 – 4,318,015 |
|---|---|
| Length | 116 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.92 |
| Mean single sequence MFE | -24.00 |
| Consensus MFE | -20.40 |
| Energy contribution | -20.74 |
| Covariance contribution | 0.34 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.537663 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4317899 116 - 27905053 AAUUUUGUUUGCGUAUUUCAAUUCGGAUUUUGAUUUCGCAUUUGAUUGCCAAAAGCGAUUUUUACAGUUAGAGUUCCACUCACUUGUGCACUCGAACAAAAUA----AAAAAAAUUGCAG .((((((((((.((..........(((((((((((((((.((((.....)))).)))).......)))))))))))(((......))).)).)))))))))).----............. ( -25.31) >DroSec_CAF1 5269 119 - 1 AAUUUUGUUUGCGUAUUUCAAUUCGGAUUUUGAUUUCGCAUUUGAUUGCCAAAAGCGAUUUUU-CAGUUAGAGUUCCACUCACUCGUGCACUCAAACAAAAUAUAAAAAAAAAAUUACAG .((((((((((.((..........(((((((((((((((.((((.....)))).)))).....-.)))))))))))(((......))).)).)))))))))).................. ( -25.20) >DroEre_CAF1 5294 109 - 1 AAUUUUGUUUGCGUAUUUCAAUUCGGAUUUUGAUUUCGCAUUUGAUUGCCAAAAGCGAUUUUUACAGUUAGAGUUUCACUCACUUAUGCACUCGGAAAAAACC----GAAAAA------- .........((((((..........((((.(((..((((.((((.....)))).))))...))).)))).(((.....)))...)))))).((((......))----))....------- ( -21.50) >consensus AAUUUUGUUUGCGUAUUUCAAUUCGGAUUUUGAUUUCGCAUUUGAUUGCCAAAAGCGAUUUUUACAGUUAGAGUUCCACUCACUUGUGCACUCGAACAAAAUA____AAAAAAAUU_CAG .((((((((((.((..........(((((((((((((((.((((.....)))).)))).......)))))))))))(((......))).)).)))))))))).................. (-20.40 = -20.74 + 0.34)



| Location | 4,317,975 – 4,318,095 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 99.44 |
| Mean single sequence MFE | -25.93 |
| Consensus MFE | -25.17 |
| Energy contribution | -25.51 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.51 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.584702 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4317975 120 - 27905053 CCAAUUCCGGUAGCCAACUUUUGGCAGCCGGCGGAUGCACAAACGAAUUUUAAACACUUUCGACAUCAAAUCCACUUUACAAUUUUGUUUGCGUAUUUCAAUUCGGAUUUUGAUUUCGCA ......(((((.((((.....)))).))))).((((.......((((...........)))).......))))................((((.(..((((........))))..))))) ( -27.44) >DroSec_CAF1 5348 120 - 1 CCAAUUCCGGUAGCCAACUUUUGGCAGCCGGCGGAUGCACAAACGAAUUUUAAACACUUUCGACAUCAAAUCCACUUUACAAUUUUGUUUGCGUAUUUCAAUUCGGAUUUUGAUUUCGCA ......(((((.((((.....)))).))))).((((.......((((...........)))).......))))................((((.(..((((........))))..))))) ( -27.44) >DroEre_CAF1 5363 120 - 1 CCAAUUCCGAUAGCCAACUUUUGGCAGCCGGCGGAUGCACAAACGAAUUUUAAACACUUUCGACAUCAAAUCCACUUUACAAUUUUGUUUGCGUAUUUCAAUUCGGAUUUUGAUUUCGCA .(((.(((((..((((.....)))).......((((((.((((((((....................................)))))))).))))))....)))))..)))........ ( -22.92) >consensus CCAAUUCCGGUAGCCAACUUUUGGCAGCCGGCGGAUGCACAAACGAAUUUUAAACACUUUCGACAUCAAAUCCACUUUACAAUUUUGUUUGCGUAUUUCAAUUCGGAUUUUGAUUUCGCA ......(((((.((((.....)))).))))).((((.......((((...........)))).......))))................((((.(..((((........))))..))))) (-25.17 = -25.51 + 0.33)


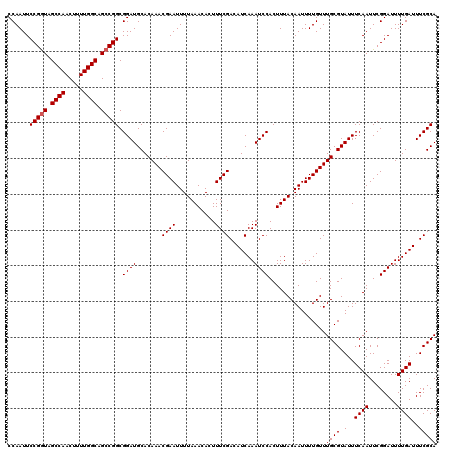
| Location | 4,318,015 – 4,318,129 |
|---|---|
| Length | 114 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.09 |
| Mean single sequence MFE | -26.22 |
| Consensus MFE | -20.92 |
| Energy contribution | -21.25 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.86 |
| Structure conservation index | 0.80 |
| SVM decision value | 1.07 |
| SVM RNA-class probability | 0.909575 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
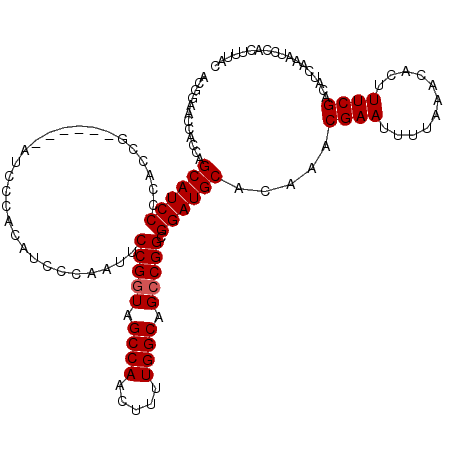
>3R_DroMel_CAF1 4318015 114 - 27905053 ACGGAACCACCAGCAUCCCCACCG------AUACCACAUCCCAAUUCCGGUAGCCAACUUUUGGCAGCCGGCGGAUGCACAAACGAAUUUUAAACACUUUCGACAUCAAAUCCACUUUAC ..(((.......((((((.....(------((.....)))......(((((.((((.....)))).))))).)))))).....((((...........))))........)))....... ( -29.90) >DroSec_CAF1 5388 120 - 1 ACUGAACCACCGGCAUCCCCACCAUCCCACAUCCCACAUCCCAAUUCCGGUAGCCAACUUUUGGCAGCCGGCGGAUGCACAAACGAAUUUUAAACACUUUCGACAUCAAAUCCACUUUAC ..(((.......((((((............................(((((.((((.....)))).))))).)))))).....((((...........))))...)))............ ( -23.95) >DroEre_CAF1 5403 112 - 1 ACGGCACCACCAGCAUCC--ACCG------AUCCCACAUCCCAAUUCCGAUAGCCAACUUUUGGCAGCCGGCGGAUGCACAAACGAAUUUUAAACACUUUCGACAUCAAAUCCACUUUAC ..((........((((((--.(((------.......(((........))).((((.....))))...))).)))))).....((((...........)))).........))....... ( -24.80) >consensus ACGGAACCACCAGCAUCCCCACCG______AUCCCACAUCCCAAUUCCGGUAGCCAACUUUUGGCAGCCGGCGGAUGCACAAACGAAUUUUAAACACUUUCGACAUCAAAUCCACUUUAC ............((((((............................(((((.((((.....)))).))))).)))))).....((((...........)))).................. (-20.92 = -21.25 + 0.33)

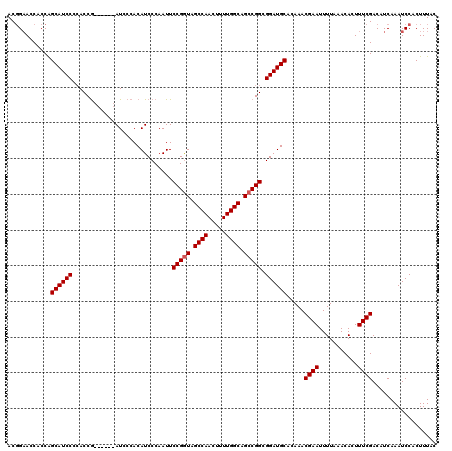
| Location | 4,318,055 – 4,318,169 |
|---|---|
| Length | 114 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 90.68 |
| Mean single sequence MFE | -42.23 |
| Consensus MFE | -37.72 |
| Energy contribution | -37.83 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.740512 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
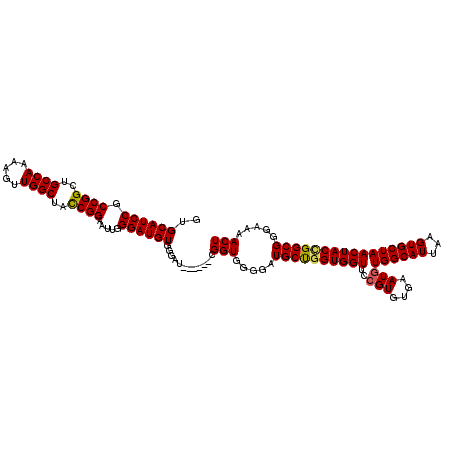
>3R_DroMel_CAF1 4318055 114 + 27905053 GUGCAUCCGCCGGCUGCCAAAAGUUGGCUACCGGAAUUGGGAUGUGGUAU------CGGUGGGGAUGCUGGUGGUUCCGUGUGAAUGUGGCAUUAAGUGCUAACUACUGGCGUGAAAACU ..((((((.((((..((((.....))))..)))).....)))))).....------.(((....((((..(((((..(((....)))((((((...)))))))))))..))))....))) ( -39.60) >DroSec_CAF1 5428 120 + 1 GUGCAUCCGCCGGCUGCCAAAAGUUGGCUACCGGAAUUGGGAUGUGGGAUGUGGGAUGGUGGGGAUGCCGGUGGUUCAGUGUGAAUGUGGCAUUAAGUGCUAACUACCGGCGGGAAAACU ..((((((.((((..((((.....))))..)))).....))))))............(((.....(((((((((((.((..(.((((...))))..)..))))))))))))).....))) ( -43.90) >DroEre_CAF1 5443 112 + 1 GUGCAUCCGCCGGCUGCCAAAAGUUGGCUAUCGGAAUUGGGAUGUGGGAU------CGGU--GGAUGCUGGUGGUGCCGUGUGAAUGUGGCAUUAAGUGCUAACUACGGGCGGGAAAACU ..(((((((((((((((((.....))))((((........))))..)).)------))))--)))))).(((..(.((((.((....((((((...))))))....)).)))).)..))) ( -43.20) >consensus GUGCAUCCGCCGGCUGCCAAAAGUUGGCUACCGGAAUUGGGAUGUGGGAU______CGGUGGGGAUGCUGGUGGUUCCGUGUGAAUGUGGCAUUAAGUGCUAACUACCGGCGGGAAAACU ..((((((.((((..((((.....))))..)))).....))))))............(((.....((((((((((..(((....)))((((((...)))))))))))))))).....))) (-37.72 = -37.83 + 0.12)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:12:18 2006