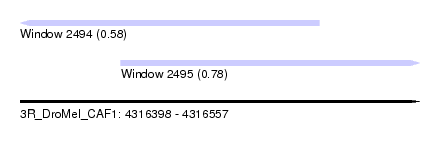

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 4,316,398 – 4,316,557 |
| Length | 159 |
| Max. P | 0.777817 |

| Location | 4,316,398 – 4,316,517 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.66 |
| Mean single sequence MFE | -35.83 |
| Consensus MFE | -32.12 |
| Energy contribution | -31.90 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.90 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.576222 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4316398 119 - 27905053 UGACAACAUUGACAAAAUGUCAAUGCUUUUCGC-CGGGCCGAGGAAAACUGGUUUUCUGGAGAAUGCUCCGCAUAAUUAGCCUCAAACUCUUGACAGUUGUUUCGCAAGAGGGGGCAUUU .((.(((((((((.....))))))).)).))((-(...((((((....((((((...(((((....)))))...))))))))))...((((((...(......).))))))))))).... ( -35.90) >DroSec_CAF1 3796 120 - 1 UGACAACAUUGACAAAAUGUCAAUGCCUUUCGUGCGGGCCGAGGAAAACUGGUUUUCUGGAGAAUGCUCCGCAUAAUUAGCCUCAAACUCUUGACAGUUGUUUCGCAAGAGGGGACAUUU ......(((((((.....)))))))((((((.(((((..(((((....((((((...(((((....)))))...)))))))))).((((......)))))..))))).))))))...... ( -37.70) >DroEre_CAF1 3788 118 - 1 UGACAACAUUGACAAAAUGUCAAUGCUUUUCGC-CGGGCCGAGGA-AACUGGUUUUCUGGAGAAUCCUCCGCAUAAUUAGCCUCAAACUCUUGACAGUUGUUUCGCAAGAGGGGACAUUU .((.(((((((((.....))))))).)).))..-.(..((((((.-..((((((...(((((....)))))...))))))))))...((((((...(......).))))))))..).... ( -33.90) >consensus UGACAACAUUGACAAAAUGUCAAUGCUUUUCGC_CGGGCCGAGGAAAACUGGUUUUCUGGAGAAUGCUCCGCAUAAUUAGCCUCAAACUCUUGACAGUUGUUUCGCAAGAGGGGACAUUU ...((((((((((.....)))))))..........(((..((((....((((((...(((((....)))))...))))))))))...))))))...(((.((((....)))).))).... (-32.12 = -31.90 + -0.22)


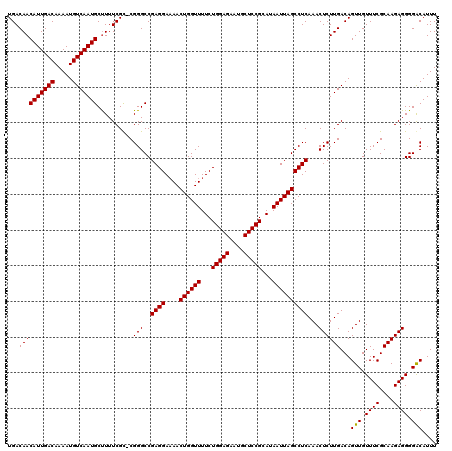
| Location | 4,316,438 – 4,316,557 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.99 |
| Mean single sequence MFE | -32.17 |
| Consensus MFE | -28.81 |
| Energy contribution | -28.82 |
| Covariance contribution | 0.01 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.90 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.777817 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
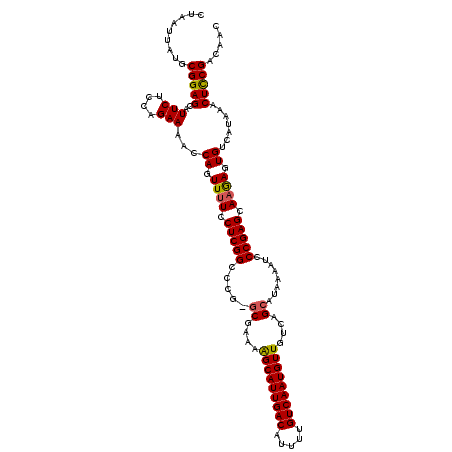
>3R_DroMel_CAF1 4316438 119 + 27905053 CUAAUUAUGCGGAGCAUUCUCCAGAAAACCAGUUUUCCUCGGCCCG-GCGAAAAGCAUUGACAUUUUGUCAAUGUUGUCAGCAUAAAAUCCCGAGCAAGAGUGUCAUAAACUCCGACAAC .......(((((((((((((...(((((....)))))(((((....-((....(((((((((.....)))))))))....))........)))))..)))))).......))))).)).. ( -33.81) >DroSec_CAF1 3836 120 + 1 CUAAUUAUGCGGAGCAUUCUCCAGAAAACCAGUUUUCCUCGGCCCGCACGAAAGGCAUUGACAUUUUGUCAAUGUUGUCAGCAUAAAAUCCCGAGCAAGAGUGUCAUAAACUUCGACAAC ....(((((.((((....)))).......((.((((.(((((......(....)((((((((.....))))))))...............))))).)))).)).)))))........... ( -28.90) >DroEre_CAF1 3828 118 + 1 CUAAUUAUGCGGAGGAUUCUCCAGAAAACCAGUU-UCCUCGGCCCG-GCGAAAAGCAUUGACAUUUUGUCAAUGUUGUCAGCAUAAAAUCCCGAGCAAAAGUGUCAUAAACUCCGGGAAC ....((((((.(((((..((..........))..-))))).....(-(((....((((((((.....)))))))))))).))))))..(((((.(....(((.......))))))))).. ( -33.80) >consensus CUAAUUAUGCGGAGCAUUCUCCAGAAAACCAGUUUUCCUCGGCCCG_GCGAAAAGCAUUGACAUUUUGUCAAUGUUGUCAGCAUAAAAUCCCGAGCAAGAGUGUCAUAAACUCCGACAAC .........(((((..(((....)))...((.((((.(((((.....((....(((((((((.....)))))))))....))........))))).)))).)).......)))))..... (-28.81 = -28.82 + 0.01)

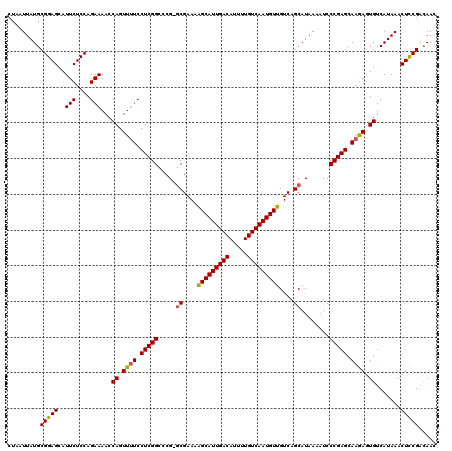
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:12:11 2006