
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 174,845 – 174,980 |
| Length | 135 |
| Max. P | 0.921959 |

| Location | 174,845 – 174,955 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.47 |
| Mean single sequence MFE | -33.32 |
| Consensus MFE | -17.98 |
| Energy contribution | -18.58 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.54 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.537972 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 174845 110 + 27905053 AUAGAGU----CUUCACGAGCACAUAGAUUACGACUGGACUAGGGGUAGGGGGUUUCAAUAAGCCCCCAGCUCACUAUUUGGUGAUCC----UG-GACGAA-GAUUGAAAGUUCGAACAA ....(((----((((.(.(((...........).)).).((((((((.((((((((....)))))))).))(((((....))))))))----))-)..)))-)))).............. ( -32.70) >DroSec_CAF1 11609 109 + 1 AUAGAGU----GUUCACGAGCACGUAGAUUACGAAUGGACUAAGGAUAGGGGGUUUCAAUAAGCC-CCAGCUCACUCUUUCGUGAUCC----UG-GCCGAA-GAUUCUAAGUUCGAACGA .((((((----.((((((....))).(((((((((.(((.....((...(((((((....)))))-))...))..)))))))))))).----..-...)))-.))))))........... ( -31.90) >DroSim_CAF1 10973 110 + 1 AUAGAGU----GUUCACGAGCACGUAGAUUACGAAUGGACUAAGGAUAGGGGGUUUCAAUAAGCCCCCAGCUCACUCUUUGGUGAUCC----UG-GCCGAA-GAUUCUAAGUUCGAACGA .....((----(((....)))))........((..((((((.((((..((((((((....)))))))).......((((((((.....----..-))))))-)))))).))))))..)). ( -36.30) >DroEre_CAF1 11845 116 + 1 AUAGAAGUGUGUUCAACGAUCACAUAGAUUACGA---GAUUAA-GAUAGGGGGUUUCAAUAAACCUCCAGCUCACGGUUUGGUGAUCCUUCCUCUGCCGAAGGAUUGUAAGUUCGAGAGA ......(((((.((...)).))))).........---......-....((((((((....))))))))..(((.(((....(..(((((((.......)))))))..)....))).))). ( -31.60) >DroYak_CAF1 12410 108 + 1 AUAGAGU----UUCCACGAUCACAUAGAUUACGG---GAUUAA-GAUAGGAGGUUUUAAUAAGCCUCCAGCCCACUGUUUGGUGAUCC----CGUGCCGAAAAAUUGAAACUACGUGAGA ....(((----(((...((((.....))))((((---(((((.-....((((((((....))))))))...(((.....)))))))))----)))...........)))))).(....). ( -34.10) >consensus AUAGAGU____GUUCACGAGCACAUAGAUUACGA_UGGACUAAGGAUAGGGGGUUUCAAUAAGCCCCCAGCUCACUCUUUGGUGAUCC____UG_GCCGAA_GAUUGUAAGUUCGAACGA ................(((((.....((((.((...............((((((((....))))))))...((((......))))............))...))))....)))))..... (-17.98 = -18.58 + 0.60)



| Location | 174,881 – 174,980 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 73.94 |
| Mean single sequence MFE | -30.26 |
| Consensus MFE | -17.26 |
| Energy contribution | -17.06 |
| Covariance contribution | -0.20 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.57 |
| SVM decision value | 1.14 |
| SVM RNA-class probability | 0.921959 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 174881 99 + 27905053 UAGGGGUAGGGGGUUUCAAUAAGCCCCCAGCUCACUAUUUGGUGAUCC----UG-GACGAA-GAUUGAAAGUUCGAACAAUUUCGAAAUGGUU-------UUCCUGAUUGAA (((((((.((((((((....)))))))).))(((((....))))))))----))-...(((-(((((....((((((....)))))).)))))-------)))......... ( -30.20) >DroSec_CAF1 11645 97 + 1 UAAGGAUAGGGGGUUUCAAUAAGCC-CCAGCUCACUCUUUCGUGAUCC----UG-GCCGAA-GAUUCUAAGUUCGAACGAUUUCGAAGUAGUU------UUUCCCGACCG-- ..((((...(((((((....)))))-))...((((......)))))))----)(-(.((.(-((..(((..((((((....)))))).)))..------)))..)).)).-- ( -28.10) >DroSim_CAF1 11009 98 + 1 UAAGGAUAGGGGGUUUCAAUAAGCCCCCAGCUCACUCUUUGGUGAUCC----UG-GCCGAA-GAUUCUAAGUUCGAACGAUUUCGAAGUAGUU------UUUCCCGACCG-- ..((((..((((((((....))))))))...(((((....))))))))----)(-(.((.(-((..(((..((((((....)))))).)))..------)))..)).)).-- ( -32.10) >DroEre_CAF1 11882 105 + 1 UAA-GAUAGGGGGUUUCAAUAAACCUCCAGCUCACGGUUUGGUGAUCCUUCCUCUGCCGAAGGAUUGUAAGUUCGAGAGAUUACGGAAUAGCCUACCAAUCAC--UG--U-- ...-(((.((((((((....)))))))).......(((..(((((((((((.......))))))))((((..((....))))))......))).))).)))..--..--.-- ( -31.20) >DroYak_CAF1 12443 105 + 1 UAA-GAUAGGAGGUUUUAAUAAGCCUCCAGCCCACUGUUUGGUGAUCC----CGUGCCGAAAAAUUGAAACUACGUGAGAUUACGGAAUAGCCAACCAAUCUCACGACUC-- ...-....((((((((....)))))))).........((((((.....----...))))))............(((((((((..((.....))....)))))))))....-- ( -29.70) >consensus UAAGGAUAGGGGGUUUCAAUAAGCCCCCAGCUCACUCUUUGGUGAUCC____UG_GCCGAA_GAUUGUAAGUUCGAACGAUUUCGAAAUAGUU______UUUCCCGACCG__ ...((((.((((((((....))))))))....(((......))))))).......................((((((....))))))......................... (-17.26 = -17.06 + -0.20)

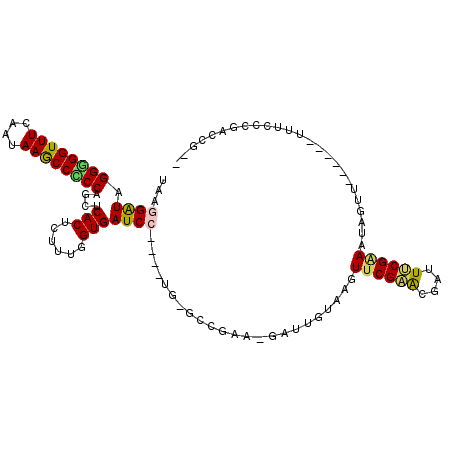

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:30:16 2006