
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 4,203,600 – 4,203,759 |
| Length | 159 |
| Max. P | 0.999183 |

| Location | 4,203,600 – 4,203,700 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 78.08 |
| Mean single sequence MFE | -24.94 |
| Consensus MFE | -13.49 |
| Energy contribution | -13.22 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.54 |
| SVM decision value | 0.77 |
| SVM RNA-class probability | 0.845953 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4203600 100 + 27905053 GCGCGCAGAGCAUUAAAAAAUUCAUAACCAAGGGGGGAGGGGGGUGA--AGGUGGUAAGAAUGAUGUCGGG-GAAAACUGAGAGCUCUGUGCUCUCAUGAAUA ..(((((((((.(((..(..(((((..((..........))..))))--)..)..)))......(.((((.-.....)))).))))))))))........... ( -30.00) >DroSec_CAF1 14946 91 + 1 GCGCGCAGAGCAUUAAAAAAUUCAUAACGAUGGGGGGAGUGGGGAGA--G--------GAUUGCUGACG-G-GAAAACUGAGAGCUCUGCGCUCUCAUGAAUA ..(((((((((.........(((((....)))))(..((..(.....--.--------..)..))..).-.-...........)))))))))........... ( -24.40) >DroSim_CAF1 15703 84 + 1 GCGCGCAGAGCAUUAAAAAAUUCAUAACGAGGGGGG-------GACA--G--------GAGUGCUGACG-G-GAGAACUGAGAGCUCUGCGCUCUCAUGAAUA ..(((((((((..........(((..((.....(..-------..).--.--------..))..)))((-(-.....)))...)))))))))........... ( -25.60) >DroEre_CAF1 15663 82 + 1 CAGCACAGAGCAUUAAAAAAUUCAUAACGAUGGGG---------AGA--G--------GAGUGCUGACG-G-GAAAACUGAGAGCACUGCGCUCUCAUGAAUA ..((.....))........((((((.......(((---------((.--(--------.((((((..((-(-.....)))..)))))).).))))))))))). ( -24.11) >DroYak_CAF1 15303 83 + 1 CUGCGCAGAGCAUUAAAAAAUUCAUAACGAUGGGG---------AGA--G--------GAGUGCUGACG-GGGAAAACUGAGAGCACUGCGCUUUCAUGAAUA .(((.....))).......((((((.......(..---------((.--(--------.((((((..((-(......)))..)))))).).))..))))))). ( -22.91) >DroAna_CAF1 17158 88 + 1 GCGCACUGAGCUUUAAAAAAUUCAUAACGUUUGGGCGG-----GAAAAUG--------GGGCAGUGGCG-G-GAUAACUGGGAGCGCAGAGCUCUCAUGAAUA ((.((((..((((((.....(((....(((....))).-----)))..))--------)))))))))).-.-......(((((((.....)))))))...... ( -22.60) >consensus GCGCGCAGAGCAUUAAAAAAUUCAUAACGAUGGGGG_______GAGA__G________GAGUGCUGACG_G_GAAAACUGAGAGCUCUGCGCUCUCAUGAAUA ..((.....))........((((((..(.....).............................................((((((.....)))))))))))). (-13.49 = -13.22 + -0.28)


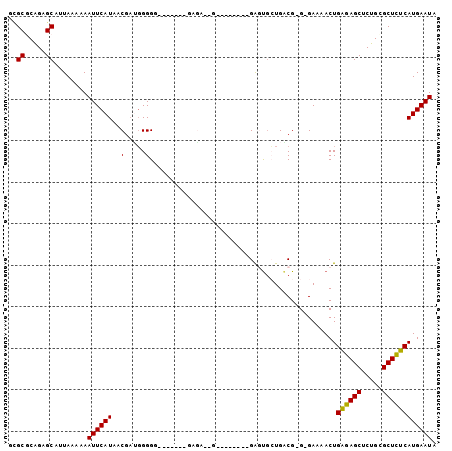
| Location | 4,203,637 – 4,203,727 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 82.97 |
| Mean single sequence MFE | -17.84 |
| Consensus MFE | -14.37 |
| Energy contribution | -15.01 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.701708 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4203637 90 + 27905053 AGGGGGGUGAAGGUGGUAAGAAUGAUGUCGGG-GAAAACUGAGAGCUCUGUGCUCUCAUGAAUAAAUCAGAACUCAAA-AAAAUAUUGGGGG ....((((...(((.......((((..((((.-.....))))((((.....))))))))......)))...))))...-............. ( -16.62) >DroSec_CAF1 14983 81 + 1 AGUGGGGAGAG--------GAUUGCUGACG-G-GAAAACUGAGAGCUCUGCGCUCUCAUGAAUAAAUCAGAACUCAAA-AAAAUAUGGGGGG ........(((--------.....((((..-.-......(((((((.....)))))))........))))..)))...-............. ( -16.83) >DroSim_CAF1 15739 75 + 1 ------GACAG--------GAGUGCUGACG-G-GAGAACUGAGAGCUCUGCGCUCUCAUGAAUAAAUCAGAACUCAAA-AAAAUAUGGGGGG ------.....--------((((.(((((.-.-..)...(((((((.....)))))))........)))).))))...-............. ( -19.80) >DroEre_CAF1 15698 73 + 1 -------AGAG--------GAGUGCUGACG-G-GAAAACUGAGAGCACUGCGCUCUCAUGAAUAAAUCAGAACUCAAA-AAAAUAUGG-GGG -------....--------((((.((((..-.-......(((((((.....)))))))........)))).))))...-.........-... ( -19.33) >DroYak_CAF1 15338 75 + 1 -------AGAG--------GAGUGCUGACG-GGGAAAACUGAGAGCACUGCGCUUUCAUGAAUAAAUCAGAACUCAAAAAAAAUAUGG-GGG -------....--------((((.((((.(-(......))((((((.....)))))).........)))).)))).............-... ( -16.60) >consensus ______GAGAG________GAGUGCUGACG_G_GAAAACUGAGAGCUCUGCGCUCUCAUGAAUAAAUCAGAACUCAAA_AAAAUAUGGGGGG ...................((((.((((...........(((((((.....)))))))........)))).))))................. (-14.37 = -15.01 + 0.64)



| Location | 4,203,637 – 4,203,727 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 82.97 |
| Mean single sequence MFE | -17.59 |
| Consensus MFE | -15.29 |
| Energy contribution | -15.97 |
| Covariance contribution | 0.68 |
| Combinations/Pair | 1.06 |
| Mean z-score | -3.35 |
| Structure conservation index | 0.87 |
| SVM decision value | 3.42 |
| SVM RNA-class probability | 0.999183 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4203637 90 - 27905053 CCCCCAAUAUUUU-UUUGAGUUCUGAUUUAUUCAUGAGAGCACAGAGCUCUCAGUUUUC-CCCGACAUCAUUCUUACCACCUUCACCCCCCU .............-..((((...((((.......(((((((.....)))))))(((...-...)))))))..))))................ ( -14.30) >DroSec_CAF1 14983 81 - 1 CCCCCCAUAUUUU-UUUGAGUUCUGAUUUAUUCAUGAGAGCGCAGAGCUCUCAGUUUUC-C-CGUCAGCAAUC--------CUCUCCCCACU .............-...(((..(((((.......(((((((.....)))))))......-.-.))))).....--------)))........ ( -17.36) >DroSim_CAF1 15739 75 - 1 CCCCCCAUAUUUU-UUUGAGUUCUGAUUUAUUCAUGAGAGCGCAGAGCUCUCAGUUCUC-C-CGUCAGCACUC--------CUGUC------ .............-...((((.(((((.......(((((((.....)))))))......-.-.))))).))))--------.....------ ( -21.76) >DroEre_CAF1 15698 73 - 1 CCC-CCAUAUUUU-UUUGAGUUCUGAUUUAUUCAUGAGAGCGCAGUGCUCUCAGUUUUC-C-CGUCAGCACUC--------CUCU------- ...-.........-...((((.(((((.......(((((((.....)))))))......-.-.))))).))))--------....------- ( -20.66) >DroYak_CAF1 15338 75 - 1 CCC-CCAUAUUUUUUUUGAGUUCUGAUUUAUUCAUGAAAGCGCAGUGCUCUCAGUUUUCCC-CGUCAGCACUC--------CUCU------- ...-.............((((.(((((.......(((.(((.....))).)))........-.))))).))))--------....------- ( -13.89) >consensus CCCCCCAUAUUUU_UUUGAGUUCUGAUUUAUUCAUGAGAGCGCAGAGCUCUCAGUUUUC_C_CGUCAGCACUC________CUCUC______ .................((((.(((((.......(((((((.....)))))))..........))))).))))................... (-15.29 = -15.97 + 0.68)

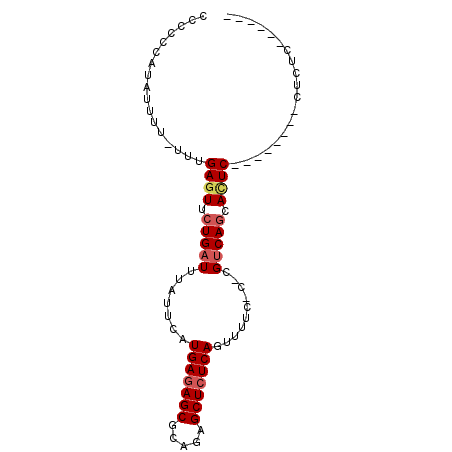

| Location | 4,203,668 – 4,203,759 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 90.16 |
| Mean single sequence MFE | -20.17 |
| Consensus MFE | -17.19 |
| Energy contribution | -17.52 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.85 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4203668 91 - 27905053 ---GGAUGGAUAUGGAACAAUAAAACAUGUUU-UUCCCCCCAAUAUUUU-UUUGAGUUCUGAUUUAUUCAUGAGAGCACAGAGCUCUCAGUUUUC-C ---((((((((.((((((...((((.(((((.-........))))).))-))...)))))))))))))).(((((((.....)))))))......-. ( -22.40) >DroSec_CAF1 15005 91 - 1 ---GGAUGGAUAUGGAACAAUAAAACAUGUUU-UUCCCCCCCAUAUUUU-UUUGAGUUCUGAUUUAUUCAUGAGAGCGCAGAGCUCUCAGUUUUC-C ---(((..((((((...........)))))).-.)))............-..(((((........)))))(((((((.....)))))))......-. ( -22.20) >DroSim_CAF1 15755 92 - 1 ---GGAUGGAUAUGGAACAAUAAAACAUGUUUUUUCCCCCCCAUAUUUU-UUUGAGUUCUGAUUUAUUCAUGAGAGCGCAGAGCUCUCAGUUCUC-C ---((..(((...((((((........)))))).)))..))........-...(((..((((.....)))(((((((.....))))))))..)))-. ( -22.20) >DroEre_CAF1 15713 90 - 1 ---GGAUGGAUAUGGAACAAUAAAACAUGUUU-UUCCCC-CCAUAUUUU-UUUGAGUUCUGAUUUAUUCAUGAGAGCGCAGUGCUCUCAGUUUUC-C ---(((..((((((...........)))))).-.)))..-.........-..(((((........)))))(((((((.....)))))))......-. ( -21.10) >DroYak_CAF1 15353 92 - 1 ---GGAUGGAUAUGGAACAAUAAAACAUGUUU-UUCCCC-CCAUAUUUUUUUUGAGUUCUGAUUUAUUCAUGAAAGCGCAGUGCUCUCAGUUUUCCC ---(((..((((((...........)))))).-.)))..-.............((((.(((.(((.......)))...))).))))........... ( -15.10) >DroAna_CAF1 17214 85 - 1 UACGGAUGGAUAUGGAACAAUAAAACAUGUUU-UUCG---------UUU-UUUCAGUUCUGAUUUAUUCAUGAGAGCUCUGCGCUCCCAGUUAUC-C ...((((((((.((((((...(((((......-...)---------)))-)....)))))))))))))).((.((((.....)))).))......-. ( -18.00) >consensus ___GGAUGGAUAUGGAACAAUAAAACAUGUUU_UUCCCC_CCAUAUUUU_UUUGAGUUCUGAUUUAUUCAUGAGAGCGCAGAGCUCUCAGUUUUC_C ...((((((((.((((((.....................................)))))))))))))).(((((((.....)))))))........ (-17.19 = -17.52 + 0.33)

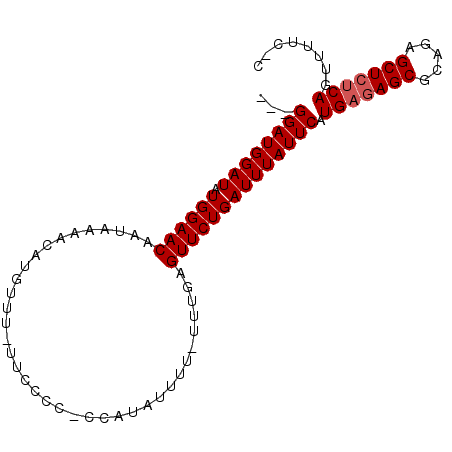

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:11:10 2006