
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 4,106,318 – 4,106,441 |
| Length | 123 |
| Max. P | 0.992356 |

| Location | 4,106,318 – 4,106,420 |
|---|---|
| Length | 102 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.36 |
| Mean single sequence MFE | -34.55 |
| Consensus MFE | -22.04 |
| Energy contribution | -21.85 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.76 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.713360 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4106318 102 - 27905053 CCGA------ACGGAGCCGAACAAUUUGUGUAAUUGACAAAUUGACGGCAGCGCU-UUGUGCGGAAGCGUUCAAAGCCGUAAAUUGUCGCUAACAAGA--GCGACACU--CC-A------ ....------..(((((((..((((((((.......)))))))).)))).(((((-((((((....)))..))))).)))....(((((((......)--)))))).)--))-.------ ( -38.80) >DroPse_CAF1 12180 107 - 1 -CGA------UGGGACCCAAACAAUUUGUGUAAUUGACAAAUUGACGGCAACGCU-UUGUGCGGAAGCGUUCGAAGCCGUAAAUUGUCGCUAACAAGA--GCAAGGCG--AC-ACGGCUA -...------..((((.((((((((((((.......)))))))).((....)).)-))).((....))))))..((((((.....((((((.......--....))))--))-)))))). ( -33.20) >DroGri_CAF1 14694 104 - 1 -AGC------CGAGACCUAAACAAUUUGUGUAAUUGACAAAUUGACGGCGACGCUUUUGUGCGGAAGCGUUCAAAGCCGUAAAUUGUCGCUAACAAGAGAGCUCGAUC--CC-A------ -...------((((((.....((((((((.......))))))))((((((((((((((....)))))))))....))))).....)))(((........))))))...--..-.------ ( -30.80) >DroWil_CAF1 2637 100 - 1 ----------GGAGACCUAAACAAUUUGUGUAAUUGACAAAUUGACGGCAACGCU-UUGUGCGGAAGCGUUCAAAGCCGUAAAUUGUCGCUAACAAGA--GCGACACA--CACAC----- ----------(....).....((((((((.......))))))))(((((((((((-((.....))))))))....)))))....(((((((......)--))))))..--.....----- ( -32.90) >DroAna_CAF1 12987 110 - 1 CCAAGUCCAGGGGGACCCAAACAAUUUGUGUAAUUGACAAAUUGACGGCAGCGCU-UUGUGCGGAAGCGUUCAAAGCCGUAAAUUGUCGCUAACAAGA--GCGACACUCUCC-A------ ..........(((((......((((((((.......))))))))(((((((((((-((.....))))))))....)))))....(((((((......)--)))))).)))))-.------ ( -38.40) >DroPer_CAF1 12257 107 - 1 -CGA------UGGGACCCAAACAAUUUGUGUAAUUGACAAAUUGACGGCAACGCU-UUGUGCGGAAGCGUUCGAAGCCGUAAAUUGUCGCUAACAAGA--GCAAGGCG--AC-ACGGCUA -...------..((((.((((((((((((.......)))))))).((....)).)-))).((....))))))..((((((.....((((((.......--....))))--))-)))))). ( -33.20) >consensus _CGA______GGGGACCCAAACAAUUUGUGUAAUUGACAAAUUGACGGCAACGCU_UUGUGCGGAAGCGUUCAAAGCCGUAAAUUGUCGCUAACAAGA__GCGACACG__CC_A______ .............(((.....((((((((.......))))))))(((((((((((.((....)).))))))....))))).....)))................................ (-22.04 = -21.85 + -0.19)


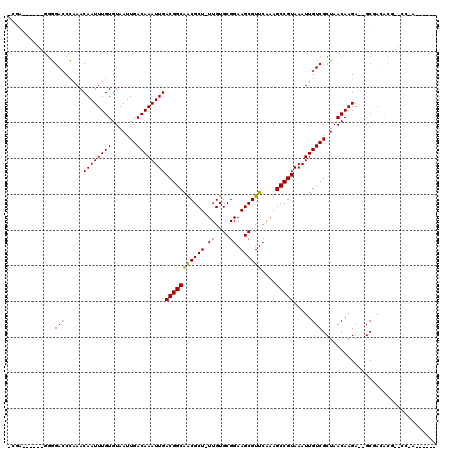
| Location | 4,106,347 – 4,106,441 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 76.45 |
| Mean single sequence MFE | -23.23 |
| Consensus MFE | -15.75 |
| Energy contribution | -16.42 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.51 |
| Structure conservation index | 0.68 |
| SVM decision value | 2.09 |
| SVM RNA-class probability | 0.987681 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4106347 94 + 27905053 ACGGCUUUGAACGCUUCCGCACAAAGCGCUGCCGUCAAUUUGUCAAUUACACAAAUUGUUCGGCUCCGU------UCGGUUCCCCCAUCUC-AAAAGCCAC ..(((((((((((....(((.....)))..((((.((((((((.......))))))))..))))..)))------))((.....)).....-.)))))).. ( -30.80) >DroPse_CAF1 12215 86 + 1 ACGGCUUCGAACGCUUCCGCACAAAGCGUUGCCGUCAAUUUGUCAAUUACACAAAUUGUUUGGGUCCCA------UCG--UCCCCCGA-------CAACAA (((((....(((((((.......)))))))))))).(((((((.......)))))))(((.(((..(..------..)--..))).))-------)..... ( -22.80) >DroWil_CAF1 2668 79 + 1 ACGGCUUUGAACGCUUCCGCACAAAGCGUUGCCGUCAAUUUGUCAAUUACACAAAUUGUUUAGGUCUCC---------------CCGA-------CAGAAU (((((....(((((((.......))))))))))))((((((((.......)))))))).....(((...---------------..))-------)..... ( -20.20) >DroMoj_CAF1 2841 90 + 1 ACGGCUUUGAACGCUUCCGCACAAUGCGUCGCCGUCAAUUUGUCAAUUACACAAAUUGUUUAGCUCUCG------ACU--CGCCACGACACCAAC---CAC ..(((.((((..(((..(((.....))).......((((((((.......))))))))...)))..)))------)..--.)))...........---... ( -17.20) >DroAna_CAF1 13018 83 + 1 ACGGCUUUGAACGCUUCCGCACAAAGCGCUGCCGUCAAUUUGUCAAUUACACAAAUUGUUUGGGUCCCCCUGGACUUGGAUCC------------------ (((((......(((((.......)))))..)))))((((((((.......))))))))((..(((((....)))))..))...------------------ ( -25.20) >DroPer_CAF1 12292 86 + 1 ACGGCUUCGAACGCUUCCGCACAAAGCGUUGCCGUCAAUUUGUCAAUUACACAAAUUGUUUGGGUCCCA------UCG--UCCCCCGA-------AACCAA (((((....(((((((.......))))))))))))((((((((.......))))))))((((((.....------...--...)))))-------)..... ( -23.20) >consensus ACGGCUUUGAACGCUUCCGCACAAAGCGUUGCCGUCAAUUUGUCAAUUACACAAAUUGUUUGGGUCCCA______UCG__UCCCCCGA________A_CAA (((((....(((((((.......))))))))))))((((((((.......))))))))........................................... (-15.75 = -16.42 + 0.67)



| Location | 4,106,347 – 4,106,441 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 76.45 |
| Mean single sequence MFE | -31.35 |
| Consensus MFE | -21.36 |
| Energy contribution | -21.17 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.10 |
| Mean z-score | -3.01 |
| Structure conservation index | 0.68 |
| SVM decision value | 2.32 |
| SVM RNA-class probability | 0.992356 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4106347 94 - 27905053 GUGGCUUUU-GAGAUGGGGGAACCGA------ACGGAGCCGAACAAUUUGUGUAAUUGACAAAUUGACGGCAGCGCUUUGUGCGGAAGCGUUCAAAGCCGU ..((((((.-......((....))((------(((..((((..((((((((.......)))))))).))))..(((.....)))....))))))))))).. ( -35.00) >DroPse_CAF1 12215 86 - 1 UUGUUG-------UCGGGGGA--CGA------UGGGACCCAAACAAUUUGUGUAAUUGACAAAUUGACGGCAACGCUUUGUGCGGAAGCGUUCGAAGCCGU ..((((-------((((((..--(..------..)..)))...((((((((.......)))))))).)))))))(((((((((....)))..))))))... ( -35.30) >DroWil_CAF1 2668 79 - 1 AUUCUG-------UCGG---------------GGAGACCUAAACAAUUUGUGUAAUUGACAAAUUGACGGCAACGCUUUGUGCGGAAGCGUUCAAAGCCGU ....((-------((((---------------(....))....((((((((.......)))))))).)))))..(((((((((....)))..))))))... ( -28.60) >DroMoj_CAF1 2841 90 - 1 GUG---GUUGGUGUCGUGGCG--AGU------CGAGAGCUAAACAAUUUGUGUAAUUGACAAAUUGACGGCGACGCAUUGUGCGGAAGCGUUCAAAGCCGU ..(---(((.(((((((.(..--((.------(....)))...((((((((.......)))))))).).))))))).....((....))......)))).. ( -28.60) >DroAna_CAF1 13018 83 - 1 ------------------GGAUCCAAGUCCAGGGGGACCCAAACAAUUUGUGUAAUUGACAAAUUGACGGCAGCGCUUUGUGCGGAAGCGUUCAAAGCCGU ------------------((((....)))).((....))....((((((((.......))))))))(((((((((((((.....))))))))....))))) ( -29.10) >DroPer_CAF1 12292 86 - 1 UUGGUU-------UCGGGGGA--CGA------UGGGACCCAAACAAUUUGUGUAAUUGACAAAUUGACGGCAACGCUUUGUGCGGAAGCGUUCGAAGCCGU ..((((-------((((((..--(..------..)..)))...((((((((.......)))))))).....((((((((.....))))))))))))))).. ( -31.50) >consensus GUG_U________UCGGGGGA__CGA______GGGGACCCAAACAAUUUGUGUAAUUGACAAAUUGACGGCAACGCUUUGUGCGGAAGCGUUCAAAGCCGU ...........................................((((((((.......))))))))(((((((((((((.....))))))))....))))) (-21.36 = -21.17 + -0.19)


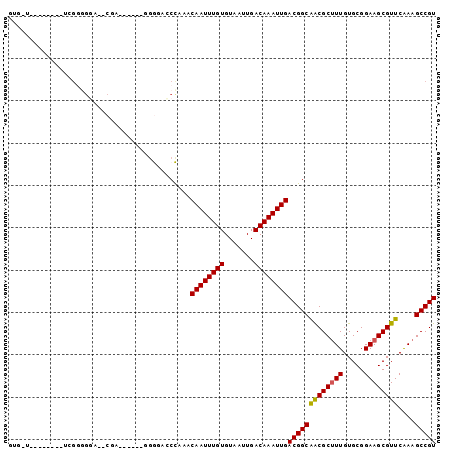
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:10:06 2006