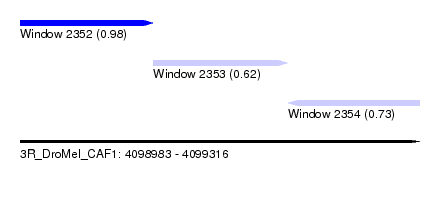
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 4,098,983 – 4,099,316 |
| Length | 333 |
| Max. P | 0.981970 |

| Location | 4,098,983 – 4,099,094 |
|---|---|
| Length | 111 |
| Sequences | 3 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 82.88 |
| Mean single sequence MFE | -31.02 |
| Consensus MFE | -28.87 |
| Energy contribution | -29.20 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.93 |
| SVM decision value | 1.90 |
| SVM RNA-class probability | 0.981970 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4098983 111 + 27905053 CCAAAUCCAGAUCCUCAUUCUUCCUUCUACUCCUACUCC---UUCUCCCGGAGGUCAGUAGCGUCCCUUUCCGGGGGUUGAAUUCAAAGGAAUUCGCUGGGUCCUCCGGCACUC .......................................---.....(((((((.(..((((.(((((....)))))..((((((....)))))))))).).)))))))..... ( -30.70) >DroSim_CAF1 10918 102 + 1 CCA------GAU------UCCUCCAACUACUCCUACUCCUCCUCCUCCCGGAGGUCAGUAGCGUCCCUUUCCGGGGGUUGAAUUCAAAGGAAUUCGCUGGGUCCUCCGGCACUC ...------(((------((((.(((((....(((((...(((((....)))))..)))))...(((.....)))))))).......))))))).(((((.....))))).... ( -32.90) >DroYak_CAF1 10894 99 + 1 ---------GAUCCUCAUUUCUCCUCCUACUCC---UUC---UUCUCGCGGAGGUCAGUAGCGUCCCUUUCCGGGGGUUGAAUUCAAAGGAAUUCGCUGGGUCCUCCGGCACUC ---------(((((((.........((((((((---(((---.......)))))..))))).).........)))))))((((((....))))))(((((.....))))).... ( -29.47) >consensus CCA______GAUCCUCAUUCCUCCUACUACUCCUACUCC___UUCUCCCGGAGGUCAGUAGCGUCCCUUUCCGGGGGUUGAAUUCAAAGGAAUUCGCUGGGUCCUCCGGCACUC ...............................................(((((((.(..((((.(((((....)))))..((((((....)))))))))).).)))))))..... (-28.87 = -29.20 + 0.33)


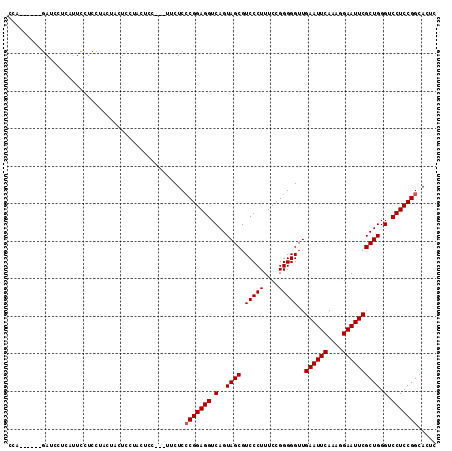
| Location | 4,099,094 – 4,099,206 |
|---|---|
| Length | 112 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.88 |
| Mean single sequence MFE | -32.52 |
| Consensus MFE | -29.11 |
| Energy contribution | -29.07 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.02 |
| Mean z-score | -1.18 |
| Structure conservation index | 0.90 |
| SVM decision value | 0.17 |
| SVM RNA-class probability | 0.618631 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4099094 112 + 27905053 UCACCUUUUCCGCGGCCAACAAAUC--------UUGAGUGCAACAAUAUGGCAAUUGUUGUGUAUUGCUACGAGAACUUGUUAAUUGUUUCGCGCUCGAGUGGCCAAGAGCUCUGAAUGG ...((..(((...(((((......(--------((((((((((((((.((((((((.(((((......))))).)).))))))))))))..)))))))))))))).((....))))).)) ( -33.60) >DroSec_CAF1 11006 112 + 1 UCACCUUUUCCGCGGCCAACAAAUC--------UUGAGUGCAACAAUAUGGCAAUUGUUGUGUAUUGCUACGAGAACUUGUUAAUUGUUUCGCGCUCGAGUGGCCAAGAGCUCUGAAUGG ...((..(((...(((((......(--------((((((((((((((.((((((((.(((((......))))).)).))))))))))))..)))))))))))))).((....))))).)) ( -33.60) >DroSim_CAF1 11020 112 + 1 UCACCUUUUCCGCGGCCAACAAAUC--------UUGAGUGCAACAAUAUGGCAAUUGUUGUGUAUUGCUACGAGAACUUGUUAAUUGUUUCGCGCUCGAGUGGCCAAGAGCUCUGAAUGG ...((..(((...(((((......(--------((((((((((((((.((((((((.(((((......))))).)).))))))))))))..)))))))))))))).((....))))).)) ( -33.60) >DroEre_CAF1 11045 120 + 1 UCACCUUUUUCGCGGCUAACAAAUCUUCAAAUCUUGAGUGCAACAAUAUGGCAAUUGUUGUGUAUUGCUACGAGAACUUGUUAAUUGUUUCGCGCUCGAGUGGCCAAGAGCUCUGAAUGG ...((...((((.(((.......((((.....(((((((((((((((.((((((((.(((((......))))).)).))))))))))))..))))))))).....))))))).)))).)) ( -32.11) >DroYak_CAF1 10993 120 + 1 UCACCUUUUUCGCGGCUAACAAAUCUUCAAAUCCUGAGUGCAACAAUAUGGCAAUUGUUGUGCGUUGCUACGAGAACUUGUUAAUUGUUUCGCGCUCGAGUGGCCAAGAGCUCUGAAUGG ......((((((..((.(((......(((.....)))(..(((((((......)))))))..))))))..))))))........(..(((.(.((((..........)))).).)))..) ( -29.70) >consensus UCACCUUUUCCGCGGCCAACAAAUC________UUGAGUGCAACAAUAUGGCAAUUGUUGUGUAUUGCUACGAGAACUUGUUAAUUGUUUCGCGCUCGAGUGGCCAAGAGCUCUGAAUGG (((.(((((....(((((...............((((((((((((((.((((((((.(((((......))))).)).))))))))))))..)))))))).))))))))))...))).... (-29.11 = -29.07 + -0.04)



| Location | 4,099,206 – 4,099,316 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.18 |
| Mean single sequence MFE | -34.14 |
| Consensus MFE | -26.53 |
| Energy contribution | -26.53 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.730319 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4099206 110 - 27905053 ACUCCAUGAAUGAAUGGAGUUGGCCACAAUGGGCCGUC-UGGAAAGUGUUUUCCUGAAAAAAAAAACCCAAGGAGCACCUAGAAGC---------ACCGCGGAAUGAAGACUGCCGCAGC (((((((......))))))).((((......)))).((-(((...((((..((((...............)))))))))))))...---------...((((...(....)..))))... ( -32.76) >DroSec_CAF1 11118 106 - 1 ACUCCAUGAAUGAAUGGAGUUGGCCACAAUGGGCCGUC-UGGAAAGUGUUUUCCUGAA----AAAACCCAAGGAGCACCUAGAACC---------ACCGUGGAAUGGAGACUGCCGCAGC (((((((......))))))).((((......)))).((-(((...((((..((((...----........)))))))))))))...---------...((((...(....)..))))... ( -35.50) >DroSim_CAF1 11132 108 - 1 ACUCCAUGAAUGAAUGGAGUUGGCCACAAUGGGCCGUC-UGGAAAGUGUUUUCCUAAA--AAAAAACCCAAGGAGCACCUAGAAGC---------ACCGCGGAAUGGAGACCGCCGCAGC (((((((......))))))).((((......)))).((-(((...((((..((((...--..........)))))))))))))...---------...((((...(....)..))))... ( -35.62) >DroEre_CAF1 11165 105 - 1 ACUCCAUGAAUGAAUGGAGCUGGCCACAAUGGGCCGUCCUGGAAAGUGUUUCGCUGA------AAACCCAAGGAGCACCAAGAAGC---------ACCGCGGAAUGGAGACAGCCGCGGC ..(((((......)))))(((((((......))))..........(((((((..((.------.....)).))))))).....)))---------.((((((..((....)).)))))). ( -39.40) >DroYak_CAF1 11113 114 - 1 ACUCCAUGAAUGAAUGGAGCUGGCCACAAUGAACCGUCCUGGAAAGUGUUUCGCAGA------AUACCCAAGGAGCACCAAGAAGCACCAAGAGCACCGCGGAAUGCAGACUGCCGCAGC .((((((......))))))..(((.........((.....))...(((((((((.(.------.....)..((.((........)).)).........))))))))).....)))..... ( -27.40) >consensus ACUCCAUGAAUGAAUGGAGUUGGCCACAAUGGGCCGUC_UGGAAAGUGUUUUCCUGAA____AAAACCCAAGGAGCACCUAGAAGC_________ACCGCGGAAUGGAGACUGCCGCAGC .((((((......))))))..((((......))))..........(((((((...................)))))))....................((((...(....)..))))... (-26.53 = -26.53 + -0.00)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:09:54 2006