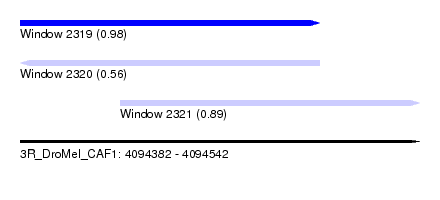
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 4,094,382 – 4,094,542 |
| Length | 160 |
| Max. P | 0.976063 |

| Location | 4,094,382 – 4,094,502 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.17 |
| Mean single sequence MFE | -41.57 |
| Consensus MFE | -38.00 |
| Energy contribution | -38.52 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.76 |
| SVM RNA-class probability | 0.976063 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4094382 120 + 27905053 UAGCAGUUUAGUUAGCUCCACUUUCUGGUUGAUCUUCAGGAAGUGCACAUAGGCAUCCUCGCUGGGCAGGAGAAACACCAGAGCGUUGCCCUCGUUUCCCUGCCGAGCCGUGCGACCUAC ............(((.((.(((((((((.......)))))))))((((..(((...))).(((.((((((.(((((...((.((...)).)).))))))))))).))).)))))).))). ( -40.90) >DroSec_CAF1 6300 120 + 1 UAGCAGUUCAGUUAGCUCCACUUUCUGGUUGAUCUUCAGGAAGUGCACAUAGGCAUCCUCGCUGGGCAGGAGAAACACAAGGGCGUUGCCUUCGUUCCCCUGCCGAGCAGUGCGUCCGAC ((((......)))).....(((((((((.......)))))))))((((..(((...))).(((.((((((.(((.....(((((...)))))..))).)))))).))).))))....... ( -41.80) >DroSim_CAF1 6337 120 + 1 UAACAGUUCAGUUAGCUCCACUUUCUGGUUAAUCUUCAGGAAGUGCACAUAGGCAUCCUCGCUGGGCAGGAGAAACACCAGGGCGUUGCCUUCGUUCCCCUGCCGAGCAGUGCGGCCUAC ((((......))))....((((((((((.......))))))))))....(((((...((.(((.((((((.(((.....(((((...)))))..))).)))))).)))))....))))). ( -45.10) >DroEre_CAF1 6344 120 + 1 GAGCAGUUUAGUUAGCUCCACCUUCUGAUUGAUCUUCAGGAAGUGCACAUAAGCAUCUUCGCUGGGCAGGAGAAACACCAGAGCGUUGCCCUCGUUUCCCUGUCGAGCAGUGCGUCCCAC ((((..........))))(((...((((.......))))(((((((......))).))))(((.((((((.(((((...((.((...)).)).))))))))))).))).)))........ ( -37.00) >DroYak_CAF1 6301 120 + 1 AAGCAGUUCAGUUAACUCCACCUUUUGGUUGAUCUUCAAGAAAUGCACAUAAGCAUCCUCGCUGGGCAGGAGAAACACCAGGGCAUUGCCCUCGUUUCCCUGCCGAGCAGUGCGUCCCAC ..(((.(((.......((.(((....))).)).......))).)))......((((....(((.((((((.(((((...(((((...))))).))))))))))).))).))))....... ( -43.04) >consensus UAGCAGUUCAGUUAGCUCCACUUUCUGGUUGAUCUUCAGGAAGUGCACAUAGGCAUCCUCGCUGGGCAGGAGAAACACCAGGGCGUUGCCCUCGUUUCCCUGCCGAGCAGUGCGUCCCAC ..................((((((((((.......)))))))))).......((((....(((.((((((.(((((...(((((...))))).))))))))))).))).))))....... (-38.00 = -38.52 + 0.52)


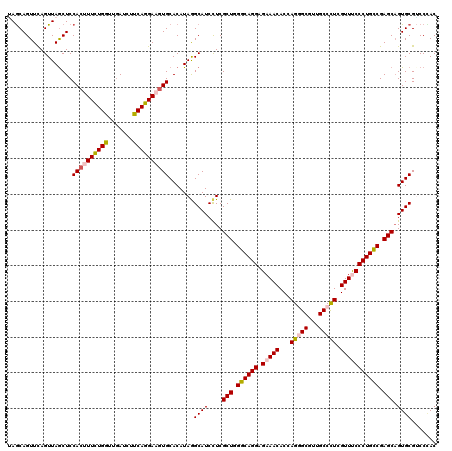
| Location | 4,094,382 – 4,094,502 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.17 |
| Mean single sequence MFE | -41.69 |
| Consensus MFE | -37.87 |
| Energy contribution | -37.23 |
| Covariance contribution | -0.64 |
| Combinations/Pair | 1.23 |
| Mean z-score | -1.16 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.558760 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4094382 120 - 27905053 GUAGGUCGCACGGCUCGGCAGGGAAACGAGGGCAACGCUCUGGUGUUUCUCCUGCCCAGCGAGGAUGCCUAUGUGCACUUCCUGAAGAUCAACCAGAAAGUGGAGCUAACUAAACUGCUA ((((.......((((((((((((((((((((((...)))))..)))))).))))))(((.((((.(((......))))))))))..................))))).......)))).. ( -43.14) >DroSec_CAF1 6300 120 - 1 GUCGGACGCACUGCUCGGCAGGGGAACGAAGGCAACGCCCUUGUGUUUCUCCUGCCCAGCGAGGAUGCCUAUGUGCACUUCCUGAAGAUCAACCAGAAAGUGGAGCUAACUGAACUGCUA .((((..((((((((.(((((((((((((.(((...))).))))...))))))))).))))(((...)))..))))((((.(((.........))).))))........))))....... ( -40.90) >DroSim_CAF1 6337 120 - 1 GUAGGCCGCACUGCUCGGCAGGGGAACGAAGGCAACGCCCUGGUGUUUCUCCUGCCCAGCGAGGAUGCCUAUGUGCACUUCCUGAAGAUUAACCAGAAAGUGGAGCUAACUGAACUGUUA ((((((..(..((((.((((((((((((.(((......)))..)))))).)))))).))))..)..))))))((.(((((.(((.........))).)))))..))((((......)))) ( -43.60) >DroEre_CAF1 6344 120 - 1 GUGGGACGCACUGCUCGACAGGGAAACGAGGGCAACGCUCUGGUGUUUCUCCUGCCCAGCGAAGAUGCUUAUGUGCACUUCCUGAAGAUCAAUCAGAAGGUGGAGCUAACUAAACUGCUC (..(........((((.((((((((((((((((...)))))..)))))).)))...(((.((((.(((......))))))))))...............)).))))........)..).. ( -36.39) >DroYak_CAF1 6301 120 - 1 GUGGGACGCACUGCUCGGCAGGGAAACGAGGGCAAUGCCCUGGUGUUUCUCCUGCCCAGCGAGGAUGCUUAUGUGCAUUUCUUGAAGAUCAACCAAAAGGUGGAGUUAACUGAACUGCUU (((((.((..(((((.(((((((((((((((((...)))))..)))))).)))))).))).))..)))))))..(((.(((.......((.(((....))).)).......))).))).. ( -44.44) >consensus GUAGGACGCACUGCUCGGCAGGGAAACGAGGGCAACGCCCUGGUGUUUCUCCUGCCCAGCGAGGAUGCCUAUGUGCACUUCCUGAAGAUCAACCAGAAAGUGGAGCUAACUGAACUGCUA ((((........(((((((((((((((((((((...)))))..)))))).))))))(((.((((.(((......))))))))))..................))))........)))).. (-37.87 = -37.23 + -0.64)


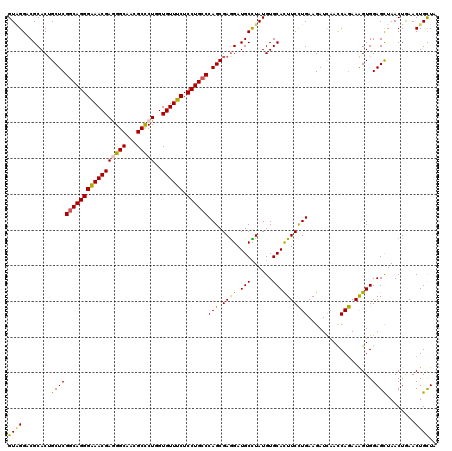
| Location | 4,094,422 – 4,094,542 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.58 |
| Mean single sequence MFE | -45.30 |
| Consensus MFE | -42.88 |
| Energy contribution | -42.76 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.29 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.96 |
| SVM RNA-class probability | 0.890447 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4094422 120 + 27905053 AAGUGCACAUAGGCAUCCUCGCUGGGCAGGAGAAACACCAGAGCGUUGCCCUCGUUUCCCUGCCGAGCCGUGCGACCUACACGGUGUACAAAGCUGGAGGCAGUUGAUGGUGGAUCCCAC .......(((..((..(((((((.((((((.(((((...((.((...)).)).))))))))))).))).((((.(((.....))))))).......))))..))..)))((((...)))) ( -42.90) >DroSec_CAF1 6340 120 + 1 AAGUGCACAUAGGCAUCCUCGCUGGGCAGGAGAAACACAAGGGCGUUGCCUUCGUUCCCCUGCCGAGCAGUGCGUCCGACACGGUGUACAAAGCUGGAGGCAGUUGAUGGUGGAUCCCAC .......(((..((..(((((((.((((((.(((.....(((((...)))))..))).)))))).))).(((((.(((...)))))))).......))))..))..)))((((...)))) ( -44.90) >DroSim_CAF1 6377 120 + 1 AAGUGCACAUAGGCAUCCUCGCUGGGCAGGAGAAACACCAGGGCGUUGCCUUCGUUCCCCUGCCGAGCAGUGCGGCCUACACGGUGUACAAAGCUGGAGGCAGUUGAUGGUGGAUCCCAC ..((((((.(((((...((.(((.((((((.(((.....(((((...)))))..))).)))))).)))))....)))))....))))))..(((((....)))))....((((...)))) ( -46.80) >DroEre_CAF1 6384 120 + 1 AAGUGCACAUAAGCAUCUUCGCUGGGCAGGAGAAACACCAGAGCGUUGCCCUCGUUUCCCUGUCGAGCAGUGCGUCCCACACGGUGCACAAAGCUGGAGGCAGUUGAUGGUGGAUCCCAU ((.(((.....(((......(((.((((((.(((((...((.((...)).)).))))))))))).))).(((((.((.....)))))))...)))....))).)).((((......)))) ( -40.70) >DroYak_CAF1 6341 120 + 1 AAAUGCACAUAAGCAUCCUCGCUGGGCAGGAGAAACACCAGGGCAUUGCCCUCGUUUCCCUGCCGAGCAGUGCGUCCCACACGAUGUACAAAGCUGGAGGCAGUUGAUGGUGGAUCCCAU ..((.(((...(((..(((((((.((((((.(((((...(((((...))))).))))))))))).))).(((((((......))))))).......))))..)))....))).))..... ( -51.20) >consensus AAGUGCACAUAGGCAUCCUCGCUGGGCAGGAGAAACACCAGGGCGUUGCCCUCGUUUCCCUGCCGAGCAGUGCGUCCCACACGGUGUACAAAGCUGGAGGCAGUUGAUGGUGGAUCCCAC ..((.(((...(((..(((((((.((((((.(((((...(((((...))))).))))))))))).))).(((((.((.....))))))).......))))..)))....))).))..... (-42.88 = -42.76 + -0.12)


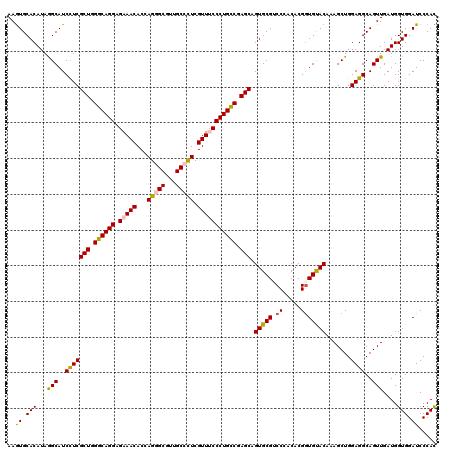
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:09:22 2006