
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 4,011,481 – 4,011,669 |
| Length | 188 |
| Max. P | 0.941805 |

| Location | 4,011,481 – 4,011,589 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 86.22 |
| Mean single sequence MFE | -27.66 |
| Consensus MFE | -19.25 |
| Energy contribution | -20.75 |
| Covariance contribution | 1.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.635950 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4011481 108 + 27905053 CCCCUUCGGUUACUCCGCCCCUGGAUCAGCCAAUC----CCCC-----CUAGCCCCCUUUGGCCAACAACUGUUACCAACUGCAGUACACACCGCCAAUGGCGGCACAUCCUGUGCA .......((((..((((....))))..))))....----....-----...(((....(((((.....(((((........))))).......))))).))).((((.....)))). ( -27.90) >DroSec_CAF1 17158 108 + 1 CCCCUUGGGUUACUCCGCCCCUGGAUCUGCCAACC----CCCC-----CUAGCCCCCUUCGGCCAACAACUGUUACCAACUGCAGUACACACCGCCAAUGGCGGCACAUCCUGUGCA ..((..((((......))))..))...(((((...----....-----...(((......))).....(((((........)))))............)))))((((.....)))). ( -28.00) >DroSim_CAF1 17734 103 + 1 C-CCUUGGGUUACUCC--CCCUGGAUCUGCCAACC----C-CC-----CUAGCCC-CUUUGGCCAACAACUGUUACCAACUGCAGUACACACCGCCAAUGGCGGCACAUCCUGUGCA .-((..(((......)--))..))...........----.-..-----...(((.-..(((((.....(((((........))))).......))))).))).((((.....)))). ( -27.30) >DroYak_CAF1 17393 116 + 1 CCCCCUGGGUUACUCCGCCCCUGGAUUUGACCCCCGUGCCCCGCUCUUCGAA-CCCCUUUAGCCACCAACUGUUAGCGACUGCAGUACACACCGCCAAUGGCGGCACAUCCUGUGCA ......((((((.((((....))))..))))))..(((....(((.......-.......))).....(((((........))))).)))..((((...))))((((.....)))). ( -27.44) >consensus CCCCUUGGGUUACUCCGCCCCUGGAUCUGCCAACC____CCCC_____CUAGCCCCCUUUGGCCAACAACUGUUACCAACUGCAGUACACACCGCCAAUGGCGGCACAUCCUGUGCA ..((..((((......))))..))...........................(((......))).....(((((........)))))......((((...))))((((.....)))). (-19.25 = -20.75 + 1.50)


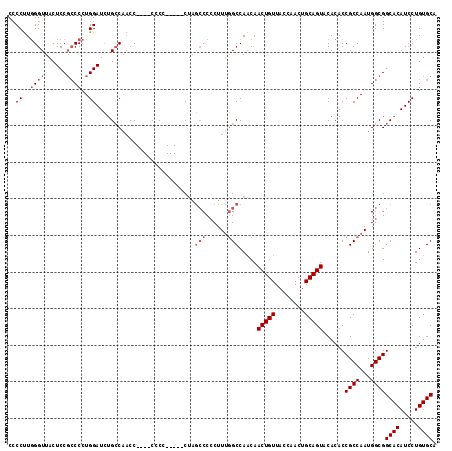
| Location | 4,011,517 – 4,011,629 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 92.58 |
| Mean single sequence MFE | -29.73 |
| Consensus MFE | -25.94 |
| Energy contribution | -26.25 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.32 |
| SVM RNA-class probability | 0.941805 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4011517 112 + 27905053 CCC-----CUAGCCCCCUUUGGCCAACAACUGUUACCAACUGCAGUACACACCGCCAAUGGCGGCACAUCCUGUGCAGCUCGUAAGCACCUCAUGCGCCAGGCUAUAAAUGUAAAUU ...-----.(((((....(((((.....(((((........))))).......)))))(((((((((.....)))).(((....)))........))))))))))............ ( -32.70) >DroSec_CAF1 17194 112 + 1 CCC-----CUAGCCCCCUUCGGCCAACAACUGUUACCAACUGCAGUACACACCGCCAAUGGCGGCACAUCCUGUGCAGCUCGUAAGCACCUCAUGCGCCAGACUAUAAAUGUAAAUU ...-----...(((......)))..((((((((........)))))............(((((((((.....)))).(((....)))........))))).........)))..... ( -24.90) >DroSim_CAF1 17767 110 + 1 -CC-----CUAGCCC-CUUUGGCCAACAACUGUUACCAACUGCAGUACACACCGCCAAUGGCGGCACAUCCUGUGCAGCUCGUAAGCACCUCAUGCGCCAGGCUAUAAAUGUAAAUU -..-----.(((((.-..(((((.....(((((........))))).......)))))(((((((((.....)))).(((....)))........))))))))))............ ( -32.70) >DroYak_CAF1 17433 116 + 1 CCGCUCUUCGAA-CCCCUUUAGCCACCAACUGUUAGCGACUGCAGUACACACCGCCAAUGGCGGCACAUCCUGUGCAGCUCGUAAGCACCUCAUGCGCCAGGCUAUAAAUGUAAAUU ............-......(((((....(((((........)))))............(((((((((.....)))).(((....)))........))))))))))............ ( -28.60) >consensus CCC_____CUAGCCCCCUUUGGCCAACAACUGUUACCAACUGCAGUACACACCGCCAAUGGCGGCACAUCCUGUGCAGCUCGUAAGCACCUCAUGCGCCAGGCUAUAAAUGUAAAUU ...................(((((....(((((........)))))............(((((((((.....)))).(((....)))........))))))))))............ (-25.94 = -26.25 + 0.31)


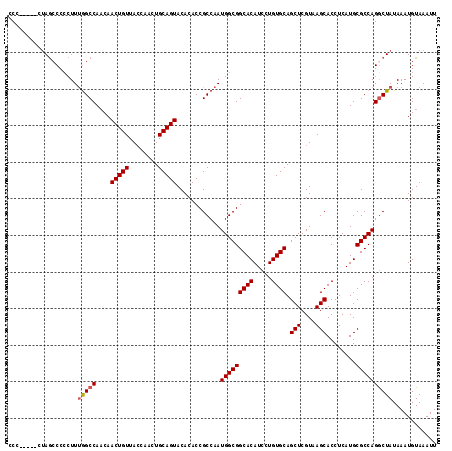
| Location | 4,011,517 – 4,011,629 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 92.58 |
| Mean single sequence MFE | -41.65 |
| Consensus MFE | -37.00 |
| Energy contribution | -37.50 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.89 |
| SVM decision value | 1.22 |
| SVM RNA-class probability | 0.932420 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4011517 112 - 27905053 AAUUUACAUUUAUAGCCUGGCGCAUGAGGUGCUUACGAGCUGCACAGGAUGUGCCGCCAUUGGCGGUGUGUACUGCAGUUGGUAACAGUUGUUGGCCAAAGGGGGCUAG-----GGG ............(((((((((.((..((.((.((((.(((((((....(((..(((((...)))))..)))..))))))).)))))).))..))))).....)))))).-----... ( -42.30) >DroSec_CAF1 17194 112 - 1 AAUUUACAUUUAUAGUCUGGCGCAUGAGGUGCUUACGAGCUGCACAGGAUGUGCCGCCAUUGGCGGUGUGUACUGCAGUUGGUAACAGUUGUUGGCCGAAGGGGGCUAG-----GGG ...............((((((.((..((.((.((((.(((((((....(((..(((((...)))))..)))..))))))).)))))).))..)).((....)).)))))-----).. ( -40.30) >DroSim_CAF1 17767 110 - 1 AAUUUACAUUUAUAGCCUGGCGCAUGAGGUGCUUACGAGCUGCACAGGAUGUGCCGCCAUUGGCGGUGUGUACUGCAGUUGGUAACAGUUGUUGGCCAAAG-GGGCUAG-----GG- ............(((((((((.((..((.((.((((.(((((((....(((..(((((...)))))..)))..))))))).)))))).))..)))))....-)))))).-----..- ( -42.70) >DroYak_CAF1 17433 116 - 1 AAUUUACAUUUAUAGCCUGGCGCAUGAGGUGCUUACGAGCUGCACAGGAUGUGCCGCCAUUGGCGGUGUGUACUGCAGUCGCUAACAGUUGGUGGCUAAAGGGG-UUCGAAGAGCGG ..............(((((((((.....)))))..((((((.(.(((.(((..(((((...)))))..))).))).((((((((.....))))))))...).))-)))).)).)).. ( -41.30) >consensus AAUUUACAUUUAUAGCCUGGCGCAUGAGGUGCUUACGAGCUGCACAGGAUGUGCCGCCAUUGGCGGUGUGUACUGCAGUUGGUAACAGUUGUUGGCCAAAGGGGGCUAG_____GGG ............(((((((((.((..((.((.((((.(((((((....(((..(((((...)))))..)))..))))))).)))))).))..))))).....))))))......... (-37.00 = -37.50 + 0.50)
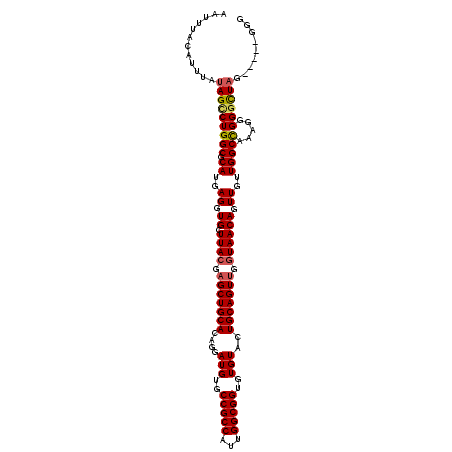



| Location | 4,011,552 – 4,011,669 |
|---|---|
| Length | 117 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.22 |
| Mean single sequence MFE | -34.92 |
| Consensus MFE | -33.12 |
| Energy contribution | -33.04 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.17 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.40 |
| SVM RNA-class probability | 0.720772 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4011552 117 + 27905053 UGCAGUACACACCGCCAAUGGC---GGCACAUCCUGUGCAGCUCGUAAGCACCUCAUGCGCCAGGCUAUAAAUGUAAAUUGCCUAAAUGGGCAAAAGCUCGAGCAUACGUUGGCCAGGAC ...........(((((((((..---.((((.....)))).(((((..(((.(((........))).............((((((....))))))..))))))))...)))))))..)).. ( -37.20) >DroSec_CAF1 17229 117 + 1 UGCAGUACACACCGCCAAUGGC---GGCACAUCCUGUGCAGCUCGUAAGCACCUCAUGCGCCAGACUAUAAAUGUAAAUUGCCUAAAUGGGCAAAAGCUCGAGCAUACGUUGGCCAGGAC ...........(((((((((..---.((((.....)))).(((((..(((..............((.......))...((((((....))))))..))))))))...)))))))..)).. ( -35.30) >DroSim_CAF1 17800 117 + 1 UGCAGUACACACCGCCAAUGGC---GGCACAUCCUGUGCAGCUCGUAAGCACCUCAUGCGCCAGGCUAUAAAUGUAAAUUGCCUAAAUGGGCAAAAGCUCGAGCAUACGUUGGCCAGGAC ...........(((((((((..---.((((.....)))).(((((..(((.(((........))).............((((((....))))))..))))))))...)))))))..)).. ( -37.20) >DroWil_CAF1 10824 109 + 1 -----------UCGCCAAUGGCUGUGGCAUAUCCUGUGCAGCUCGUAAGCACCUCAUGCGCCAGGCUAUAAAUGUAAAUUGCCUAAAUGGGUAGAAGCUCGAGCAUACGUUGCCCUCUAC -----------..((..((((((.((((.(((...((((.........))))...))).)))))))))).((((((..((((((....))))))..((....)).))))))))....... ( -27.70) >DroYak_CAF1 17472 117 + 1 UGCAGUACACACCGCCAAUGGC---GGCACAUCCUGUGCAGCUCGUAAGCACCUCAUGCGCCAGGCUAUAAAUGUAAAUUGCCUAAAUGGGCAAAAGCUCGAGCAUACGUUGGCCAGGAC ...........(((((((((..---.((((.....)))).(((((..(((.(((........))).............((((((....))))))..))))))))...)))))))..)).. ( -37.20) >consensus UGCAGUACACACCGCCAAUGGC___GGCACAUCCUGUGCAGCUCGUAAGCACCUCAUGCGCCAGGCUAUAAAUGUAAAUUGCCUAAAUGGGCAAAAGCUCGAGCAUACGUUGGCCAGGAC .............(((((((......((((.....)))).(((((..(((.(((........))).............((((((....))))))..))))))))...)))))))...... (-33.12 = -33.04 + -0.08)

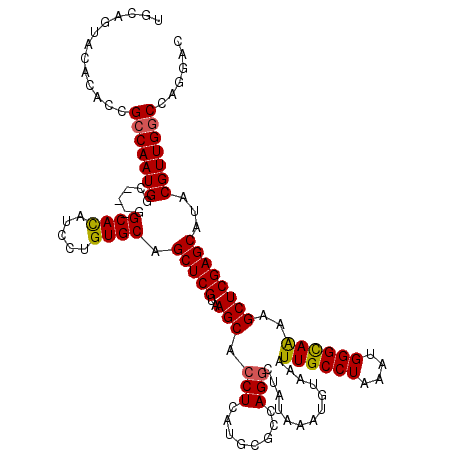

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:08:31 2006