
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 4,004,318 – 4,004,472 |
| Length | 154 |
| Max. P | 0.895509 |

| Location | 4,004,318 – 4,004,432 |
|---|---|
| Length | 114 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 85.04 |
| Mean single sequence MFE | -38.02 |
| Consensus MFE | -26.50 |
| Energy contribution | -28.12 |
| Covariance contribution | 1.62 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.99 |
| SVM RNA-class probability | 0.895509 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
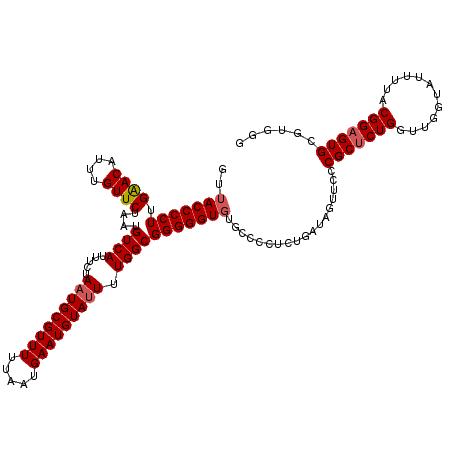
>3R_DroMel_CAF1 4004318 114 + 27905053 GCGGUG------GGCUGUUCGUGGUGGGUGGUAAUGCACGCCCACGCACUCCGUAAAAUACCAACCAGAGCGGCAACUAUCAGAGGAGCACACCCCCGCCAAAAUACAUUCAUUAAAAAC ((((.(------((.(((((((((((((((........))))))).))).((((...............))))............)))))..)))))))..................... ( -36.86) >DroSec_CAF1 9914 114 + 1 GCGGUG------GGCCACUCGUGGUGGGUGGUAAUGCACGCCCACGCACUCCGUAAAAUACCAACCAGAGCGGGAACUAUCAGAGGGGCACACCCCCGCCAAAAUACAUUCAUUAAAAAC ((((.(------((((.(((((((((((((........))))))).)))(((((...............)))))........))).)))...)).))))..................... ( -38.16) >DroSim_CAF1 10490 114 + 1 GCGGUG------GGCCACUCGUGGUGGGUGGUAAUGCACGCCCACGCACUCCGUAAAAUACCAACCAGAGCGGGAACUAUCAGAGGGACACACCCCCGCCAAAAUACAUUCAUUAAAAAC ((((.(------((((.(((((((((((((........))))))).)))(((((...............)))))........))))).....)))))))..................... ( -37.16) >DroYak_CAF1 9904 105 + 1 GCGGUUUCUGGUGGGUGGUGGUGGUGGGUGGUAAUGCACGCCCACGCACUCCGUAAAACACCAACCAGUGCGGCA---------------AACCCCCGCCAAACUACAUUCAUUAAAAAC (..(((..(((((((.((((((((((((((........))))))).))))(((((.............)))))..---------------.)))))))))))))..)............. ( -39.92) >consensus GCGGUG______GGCCACUCGUGGUGGGUGGUAAUGCACGCCCACGCACUCCGUAAAAUACCAACCAGAGCGGCAACUAUCAGAGGGGCACACCCCCGCCAAAAUACAUUCAUUAAAAAC ..((((.......(((.(((((((((((((........))))))).))).((((...............)))).........))).))).......)))).................... (-26.50 = -28.12 + 1.62)



| Location | 4,004,352 – 4,004,472 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 90.56 |
| Mean single sequence MFE | -33.58 |
| Consensus MFE | -28.74 |
| Energy contribution | -29.30 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.17 |
| SVM RNA-class probability | 0.616446 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 4004352 120 - 27905053 GUUACCCCUUGAACAUUUGUUCUAAUGUCAUUUCUAAUGCGUUUUUAAUGAAUGUAUUUUGGCGGGGGUGUGCUCCUCUGAUAGUUGCCGCUCUGGUUGGUAUUUUACGGAGUGCGUGGG ...((((((.((((....))))....((((.....(((((((((.....))))))))).))))))))))(..((((...((((.(.(((.....))).).))))....))))..)..... ( -37.70) >DroSec_CAF1 9948 120 - 1 GUUACCCCUUGAACAUUUGUUCUAAUGUCAUUUCUAAUGCGUUUUUAAUGAAUGUAUUUUGGCGGGGGUGUGCCCCUCUGAUAGUUCCCGCUCUGGUUGGUAUUUUACGGAGUGCGUGGG ..(((((((.((((....))))....((((.....(((((((((.....))))))))).)))))))))))...(((.(.((....)).(((((((............))))))).).))) ( -34.30) >DroSim_CAF1 10524 120 - 1 GUUACCCCUUGGACAUUUGUUCUAAUGUCAUUUCUAAUGCGUUUUUAAUGAAUGUAUUUUGGCGGGGGUGUGUCCCUCUGAUAGUUCCCGCUCUGGUUGGUAUUUUACGGAGUGCGUGGG .....(((.((.((.(((((...(((..((...(((((((((((.....))))))))...((((((..(.((((.....)))))..)))))).))).))..)))..))))))).)).))) ( -33.90) >DroYak_CAF1 9944 105 - 1 GUUACCCCUUGAACAUUUGUUCUAAUGUCAUUUCUAAUGCGUUUUUAAUGAAUGUAGUUUGGCGGGGGUU---------------UGCCGCACUGGUUGGUGUUUUACGGAGUGCGUGGG ((.((((((.((((....))))....((((.......(((((((.....)))))))...)))))))))).---------------.))((((((..(((........))))))))).... ( -28.40) >consensus GUUACCCCUUGAACAUUUGUUCUAAUGUCAUUUCUAAUGCGUUUUUAAUGAAUGUAUUUUGGCGGGGGUGUGCCCCUCUGAUAGUUCCCGCUCUGGUUGGUAUUUUACGGAGUGCGUGGG ..(((((((.((((....))))....((((.....(((((((((.....))))))))).)))))))))))..................(((((((............)))))))...... (-28.74 = -29.30 + 0.56)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:08:18 2006