
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 3,949,433 – 3,949,553 |
| Length | 120 |
| Max. P | 0.952395 |

| Location | 3,949,433 – 3,949,553 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 78.78 |
| Mean single sequence MFE | -42.08 |
| Consensus MFE | -21.47 |
| Energy contribution | -21.28 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.38 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.51 |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.594781 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3949433 120 + 27905053 AAGGAGGGGGAAACGUCACCGCCUAUUCCAAAUCCAGUUUGGACCUUCGAGCAAUGCUUUGCCGAAUAUCCAGAUAUGCUUGAGGAGAUUACCAAAAUGGGCUUCUCCAAGCCCUCGCCC ...(((((((((..((....))...)))).......(((((((..((((.((((....))))))))..)))))))........(((((...((.....))...)))))...))))).... ( -38.80) >DroVir_CAF1 17300 120 + 1 AAGGAUGGCGAAACCCCAACGCCCAUACCCAAUCCUGUAUGGACUUUCGAGCAAUGCUUUGCCGAAUAUCCUGAUUUGCUGGGCGAAAUACAGAAGCAGGGCUUUGCGCAUCCAUCGCCC ..((((((((((.(((...(((((((((........))))(((..((((.((((....))))))))..))).........))))).....(....)..))).))))).)))))....... ( -40.00) >DroPse_CAF1 15879 120 + 1 AAGGAGGGCGAAGAAAUCCCACCCAUACCGAAUCCGGUAUGGACAUUCGAGCAAUGCUUUGCCGAAUAUCCCGAUCUGUUGGGCGAAAUCGAGAAGCAGGGCUUCCCUAAGCCCUCGCCC ...((((((..((.........((((((((....))))))))......((((..((((((..(((....(((((....))))).....))).))))))..))))..))..)))))).... ( -47.90) >DroGri_CAF1 15566 120 + 1 AAGGAGGGAGAAACUUUGCCAGCAAUUCCGAAUCCCGUGUGGAGUUUCGAGCAGUGCUUUGACGAGUAUCCCGAUCUGCUAGCCGAAGUGCAGAAGCAGGGUUUCGCGAAGCCCUCGCCC ...((((..(((((((..((................).)..))))))).......((((((.((((...(((..(((((..........)))))....))).)))))))))))))).... ( -38.49) >DroMoj_CAF1 17896 120 + 1 CAGGAGGGCGAAGUGUUUGCGCCCAUACCGAAUCCUGUAUGGAAGUUCGAGCAAUGCUUUGCCGAGUACCCCGAUUUGCUGGCCGAAAUACAGAAGCAGGGAUUUACGAAGCCUUCACCC .....(((.((((((((((.((((((((.(....).))))))..)).))))))..((((((..((((.(((.(.....(((.........)))...).))))))).)))))))))).))) ( -36.00) >DroPer_CAF1 15868 120 + 1 AAGGAGGGCGAAGAAGUCCCACCCAUACCGAAUCCGGUAUGGACAUUCGAGCAAUGCUUUGCCGAAUAUCCCGAUCUGUUGGGUGAAAUCGAGAAGCAGGGCUUCCCUAAGCCCUCGCCC ...((((((...(((((((...((((((((....))))))))............((((((..(((....(((((....))))).....))).))))))))))))).....)))))).... ( -51.30) >consensus AAGGAGGGCGAAACAUUACCACCCAUACCGAAUCCGGUAUGGACAUUCGAGCAAUGCUUUGCCGAAUAUCCCGAUCUGCUGGGCGAAAUACAGAAGCAGGGCUUCCCGAAGCCCUCGCCC ...(((((..............((((((........))))))...((((.((((....))))))))........((((((...(........).))))))...........))))).... (-21.47 = -21.28 + -0.19)


| Location | 3,949,433 – 3,949,553 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 78.78 |
| Mean single sequence MFE | -47.03 |
| Consensus MFE | -25.96 |
| Energy contribution | -27.10 |
| Covariance contribution | 1.14 |
| Combinations/Pair | 1.38 |
| Mean z-score | -2.61 |
| Structure conservation index | 0.55 |
| SVM decision value | 1.42 |
| SVM RNA-class probability | 0.952395 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
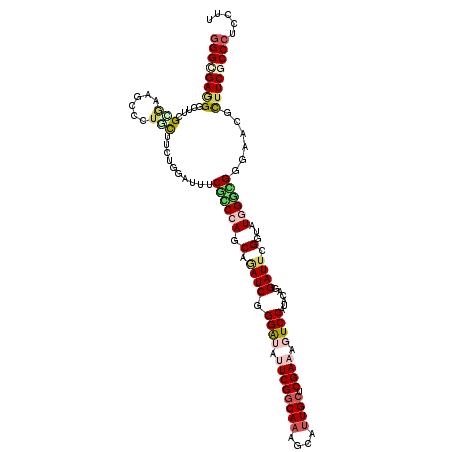
>3R_DroMel_CAF1 3949433 120 - 27905053 GGGCGAGGGCUUGGAGAAGCCCAUUUUGGUAAUCUCCUCAAGCAUAUCUGGAUAUUCGGCAAAGCAUUGCUCGAAGGUCCAAACUGGAUUUGGAAUAGGCGGUGACGUUUCCCCCUCCUU .((.(((((((((((((.(((......)))..)))))..)))).....(((((.((((((((....)))).)))).)))))..........((((...(((....))))))).)))))). ( -40.40) >DroVir_CAF1 17300 120 - 1 GGGCGAUGGAUGCGCAAAGCCCUGCUUCUGUAUUUCGCCCAGCAAAUCAGGAUAUUCGGCAAAGCAUUGCUCGAAAGUCCAUACAGGAUUGGGUAUGGGCGUUGGGGUUUCGCCAUCCUU .......(((((.((.(((((((((....))....((((((((.((((.((((.((((((((....)))).)))).))))......))))..)).))))))..))))))).))))))).. ( -45.50) >DroPse_CAF1 15879 120 - 1 GGGCGAGGGCUUAGGGAAGCCCUGCUUCUCGAUUUCGCCCAACAGAUCGGGAUAUUCGGCAAAGCAUUGCUCGAAUGUCCAUACCGGAUUCGGUAUGGGUGGGAUUUCUUCGCCCUCCUU (((((((((....(((((((...)))))))(((..(((((.........(((((((((((((....)))).)))))))))((((((....)))))))))))..))))))))))))..... ( -61.40) >DroGri_CAF1 15566 120 - 1 GGGCGAGGGCUUCGCGAAACCCUGCUUCUGCACUUCGGCUAGCAGAUCGGGAUACUCGUCAAAGCACUGCUCGAAACUCCACACGGGAUUCGGAAUUGCUGGCAAAGUUUCUCCCUCCUU ((((((((((...(((......)))....)).)))))((((((((.((.((((.(((((...(((...))).(......)..))))))))).)).)))))))).........)))..... ( -35.80) >DroMoj_CAF1 17896 120 - 1 GGGUGAAGGCUUCGUAAAUCCCUGCUUCUGUAUUUCGGCCAGCAAAUCGGGGUACUCGGCAAAGCAUUGCUCGAACUUCCAUACAGGAUUCGGUAUGGGCGCAAACACUUCGCCCUCCUG ((((((((((((.((..(((((((((.(((.....)))..))))....))))).....)).))))...(((((.((((((.....)))...))).))))).......))))))))..... ( -39.30) >DroPer_CAF1 15868 120 - 1 GGGCGAGGGCUUAGGGAAGCCCUGCUUCUCGAUUUCACCCAACAGAUCGGGAUAUUCGGCAAAGCAUUGCUCGAAUGUCCAUACCGGAUUCGGUAUGGGUGGGACUUCUUCGCCCUCCUU (((((((((....(((((((...)))))))..(..(((((.........(((((((((((((....)))).)))))))))((((((....)))))))))))..)..)))))))))..... ( -59.80) >consensus GGGCGAGGGCUUCGCGAAGCCCUGCUUCUGGAUUUCGCCCAGCAGAUCGGGAUAUUCGGCAAAGCAUUGCUCGAAAGUCCAUACAGGAUUCGGUAUGGGCGGGAACGCUUCGCCCUCCUU ((((((((.....(((......)))..........((((((.(.((((.((((.((((((((....)))).)))).))))......)))).)...))))))......))))))))..... (-25.96 = -27.10 + 1.14)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:07:09 2006