
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 3,927,072 – 3,927,178 |
| Length | 106 |
| Max. P | 0.999190 |

| Location | 3,927,072 – 3,927,178 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 66.47 |
| Mean single sequence MFE | -18.65 |
| Consensus MFE | -8.32 |
| Energy contribution | -9.07 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.35 |
| Structure conservation index | 0.45 |
| SVM decision value | 1.53 |
| SVM RNA-class probability | 0.961504 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3927072 106 + 27905053 AUAAUCGUUUU-----UAUAUUCACAUUGACCUUUGUAGCC-AGCCGCUACAAAGUGGUAUAUAGUCGUUAGACAAUUUUAAGUUAUAAAUCCAUUUCUUUGUUAACACACC-------- ......(..((-----((((..........((((((((((.-....))))))))).).......(((....)))..........))))))..)...................-------- ( -18.10) >DroSec_CAF1 1917 107 + 1 AUAAUCAUUUG-----CAUAUUCACAUUGACCUUUGUAGCCCAUCCGCUACAAAGUAAUAUAUAGUCGUUAGACAAUUAUAAAUUAUAAACCCAUUUCUUUGUUAACACACC-------- ...........-----.......(((..((.(((((((((......)))))))))...((((..(((....)))...))))...............))..))).........-------- ( -16.70) >DroSim_CAF1 1951 107 + 1 AUAAUCAUUUG-----CAUAUUCACAUUGACCUUUGUAAAC-AGUCACUACAAAGUGGUAUAUAGU-------CAAUUAUAAAUUAUAAACCCAUUUCUUUGUUAACACACUUGUUAACA ...........-----...............(((((((...-......))))))).(((.((((((-------.........)))))).)))........((((((((....)))))))) ( -14.60) >DroEre_CAF1 1915 90 + 1 AUUACCAUAUGGGUAUUAUAUUUACAUAGACCUUUGUAGCU-AAGCGCCACAAAGUGCUAUGUGAUUGUUAUUCAGUAAUAAAUUACU-------------------GC----------A ..((((.....))))..(((.((((((((..((((((.((.-....)).))))))..)))))))).)))....((((((....)))))-------------------).----------. ( -25.20) >consensus AUAAUCAUUUG_____CAUAUUCACAUUGACCUUUGUAGCC_AGCCGCUACAAAGUGGUAUAUAGUCGUUAGACAAUUAUAAAUUAUAAACCCAUUUCUUUGUUAACACACC________ ...............................(((((((((......)))))))))................................................................. ( -8.32 = -9.07 + 0.75)

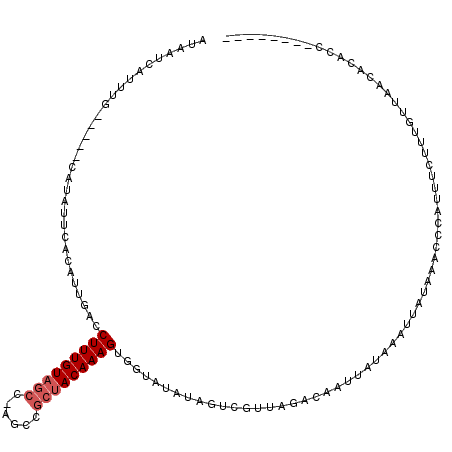

| Location | 3,927,072 – 3,927,178 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 66.47 |
| Mean single sequence MFE | -23.88 |
| Consensus MFE | -12.94 |
| Energy contribution | -13.00 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.26 |
| Mean z-score | -2.58 |
| Structure conservation index | 0.54 |
| SVM decision value | 3.42 |
| SVM RNA-class probability | 0.999190 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3927072 106 - 27905053 --------GGUGUGUUAACAAAGAAAUGGAUUUAUAACUUAAAAUUGUCUAACGACUAUAUACCACUUUGUAGCGGCU-GGCUACAAAGGUCAAUGUGAAUAUA-----AAAACGAUUAU --------..((((((.(((..((..(((.((((.....))))...(((....)))......)))(((((((((....-.))))))))).))..))).))))))-----........... ( -23.40) >DroSec_CAF1 1917 107 - 1 --------GGUGUGUUAACAAAGAAAUGGGUUUAUAAUUUAUAAUUGUCUAACGACUAUAUAUUACUUUGUAGCGGAUGGGCUACAAAGGUCAAUGUGAAUAUG-----CAAAUGAUUAU --------.(((((((.(((..((..........((((.((((...(((....))))))).))))(((((((((......))))))))).))..))).))))))-----).......... ( -25.00) >DroSim_CAF1 1951 107 - 1 UGUUAACAAGUGUGUUAACAAAGAAAUGGGUUUAUAAUUUAUAAUUG-------ACUAUAUACCACUUUGUAGUGACU-GUUUACAAAGGUCAAUGUGAAUAUG-----CAAAUGAUUAU ((((((((....))))))))........(((.((((.((.......)-------).)))).))).((((((((.....-..))))))))((((.(((......)-----))..))))... ( -19.60) >DroEre_CAF1 1915 90 - 1 U----------GC-------------------AGUAAUUUAUUACUGAAUAACAAUCACAUAGCACUUUGUGGCGCUU-AGCUACAAAGGUCUAUGUAAAUAUAAUACCCAUAUGGUAAU .----------.(-------------------(((((....))))))..........((((((..(((((((((....-.)))))))))..))))))........((((.....)))).. ( -27.50) >consensus ________GGUGUGUUAACAAAGAAAUGGGUUUAUAAUUUAUAAUUGUCUAACGACUAUAUACCACUUUGUAGCGGCU_GGCUACAAAGGUCAAUGUGAAUAUA_____CAAAUGAUUAU ........................................(((((((((....)))(((((....(((((((((......)))))))))....)))))................)))))) (-12.94 = -13.00 + 0.06)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:06:57 2006