
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 689,720 – 689,817 |
| Length | 97 |
| Max. P | 0.882592 |

| Location | 689,720 – 689,817 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 77.78 |
| Mean single sequence MFE | -19.26 |
| Consensus MFE | -17.54 |
| Energy contribution | -17.07 |
| Covariance contribution | -0.47 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.40 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.92 |
| SVM RNA-class probability | 0.882592 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 689720 97 - 27905053 GGCCAGA-GAAU--CCGA-GGGAUGA----GCAUCAUCAUCGCUCAUGUGUCAACAGCAUGCAAUGCGAUAUGAUUAUUAUUAUCUAACAAUACA--AUAACAAAAC------------- .......-....--..(.-.((((((----..(((((.(((((.(((((.......)))))....)))))))))).....))))))..)......--..........------------- ( -18.40) >DroPse_CAF1 111827 111 - 1 UGAAAGA-UAGC--AAGAGGGGAUGA----GCAGCAUCAUCGCUCAUGUGUCAACAACAUGCAAUGCGAUAUGAUUAUUAUUAUCUAACAAUACA--AUAACAAAACGGCAGCUCCCGUG .......-....--.(((...(((((----....(((.(((((.(((((.......)))))....))))))))....))))).))).........--........((((......)))). ( -21.10) >DroGri_CAF1 121424 106 - 1 AGAGAGA-GAAG--UAGUGGAGAUAAGCG-GCAUCAUCAUCGCUCAUGUGUCAACAACAUGCAAUGCGAUAUGAUUAUUAUUAUCUAACAAUACAAAACAACAAAACAAC---------- .......-...(--((.((.((((((...-..(((((.(((((.(((((.......)))))....)))))))))).....))))))..)).)))................---------- ( -22.20) >DroEre_CAF1 106297 82 - 1 -------------------AGGAUGA----GCAUCAUCAUCGCUCAUGUGUCAACAGCAUGCAAUGCGAUAUGAUUAUUAUUAUCUAACAAUACA--AUAACAAAAC------------- -------------------.((((((----..(((((.(((((.(((((.......)))))....)))))))))).....)))))).........--..........------------- ( -18.00) >DroMoj_CAF1 120089 109 - 1 AGUGAGU-GGAGUAGAGCGGAGAUAAGCGGGCAGCAUCAUUGCUCAUGUGUCAACAACAUGCAAUGCGAUAUGAUUAUUAUUAUCUAACAAUACAAAAUAACAAAACAGU---------- .......-...(((......((((((........(((.(((((.(((((.......)))))....)))))))).......)))))).....)))................---------- ( -17.26) >DroAna_CAF1 108456 96 - 1 --CCAAAUGAUA--GCAA-GGGAUGA----GCAUCAUCAUCGCUCAUGUGUCAACAACAUGCAAUGCGAUAUGAUUAUUAUUAUCUAACAAUACA--AUAACAAAAC------------- --...(((((((--(((.-..((((.----....))))(((((.(((((.......)))))....))))).)).)))))))).............--..........------------- ( -18.60) >consensus _GACAGA_GAAG__GAGA_GGGAUGA____GCAUCAUCAUCGCUCAUGUGUCAACAACAUGCAAUGCGAUAUGAUUAUUAUUAUCUAACAAUACA__AUAACAAAAC_____________ ....................((((((......(((((.(((((.(((((.......)))))....)))))))))).....)))))).................................. (-17.54 = -17.07 + -0.47)
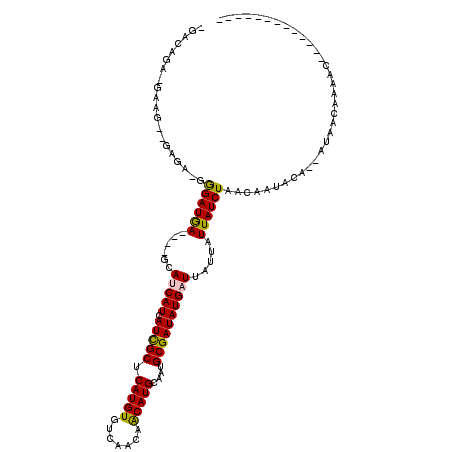


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:33:53 2006