
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 3,770,391 – 3,770,511 |
| Length | 120 |
| Max. P | 0.843542 |

| Location | 3,770,391 – 3,770,511 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.11 |
| Mean single sequence MFE | -39.43 |
| Consensus MFE | -34.39 |
| Energy contribution | -34.83 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.02 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.670619 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3770391 120 + 27905053 GCUGUGGACUGAGAAAUGCAACGGGAUGCAUCCCUGGCGAUAUAUUUUGGCAGCAAGUAUUCGGUGGGUGUAUUUUUAGCUGGCCAAAAAUGAAUUUGCAUCGAAUGCAUCCGUCAGCAC ((((((((........(((...((((....))))((.(((......))).))))).((((((((((.(..(((((((.(....).)))))))....).)))))))))).)))).)))).. ( -39.40) >DroSim_CAF1 3595 120 + 1 GCUGUGGACUGAGAAAUGCCACGGGAUGCAUCCCUGGCGAUACAUUUUGGCAGCAAGAAUUCGGAGGGUGUAUUUUUAGCUGGCCAAAAAUGAAUUUGCAUCGAAUGCAUCCGUCAGCAC ((((((((........(((((.((((....)))))))))...((((((((((((((((((.((.....)).)))))).))).))).))))))....(((((...))))))))).)))).. ( -40.40) >DroYak_CAF1 2074 120 + 1 GCUGUGGACUGAGAAAUGCCACGGGAUGCAUCCCUGGCGAAAUAUUUUGGCAGCAAGUAUUCGGAGGAUGUAUUUUUAGCUGGCCAAAAAUGAAUUUGAAUCGAAUGCAUCCGUCAGCAC (((((((((((.(((((((((.((((....)))))))).....)))))..)))...((((((((.((((.(((((((.(....).))))))).))))...)))))))).)))).)))).. ( -38.50) >consensus GCUGUGGACUGAGAAAUGCCACGGGAUGCAUCCCUGGCGAUAUAUUUUGGCAGCAAGUAUUCGGAGGGUGUAUUUUUAGCUGGCCAAAAAUGAAUUUGCAUCGAAUGCAUCCGUCAGCAC (((((((((((.(((((((((.((((....)))))))).....)))))..)))...((((((((.((((.(((((((.(....).))))))).))))...)))))))).)))).)))).. (-34.39 = -34.83 + 0.45)


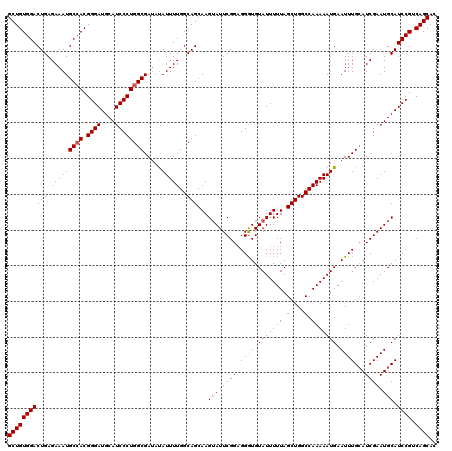
| Location | 3,770,391 – 3,770,511 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.11 |
| Mean single sequence MFE | -34.41 |
| Consensus MFE | -30.63 |
| Energy contribution | -30.96 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.36 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.76 |
| SVM RNA-class probability | 0.843542 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
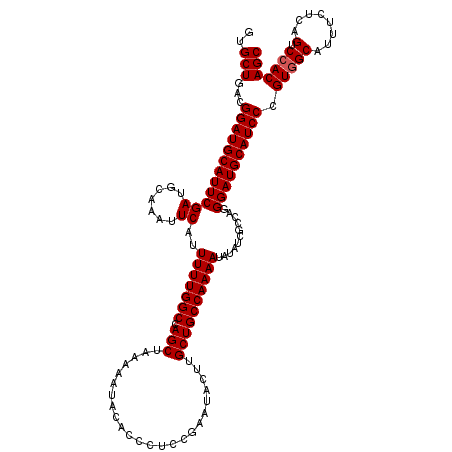
>3R_DroMel_CAF1 3770391 120 - 27905053 GUGCUGACGGAUGCAUUCGAUGCAAAUUCAUUUUUGGCCAGCUAAAAAUACACCCACCGAAUACUUGCUGCCAAAAUAUAUCGCCAGGGAUGCAUCCCGUUGCAUUUCUCAGUCCACAGC (..((((..((((((...((((((.....((((((((....))))))))...(((..((((((.((.......)).))).)))...))).))))))....))))))..))))..)..... ( -33.90) >DroSim_CAF1 3595 120 - 1 GUGCUGACGGAUGCAUUCGAUGCAAAUUCAUUUUUGGCCAGCUAAAAAUACACCCUCCGAAUUCUUGCUGCCAAAAUGUAUCGCCAGGGAUGCAUCCCGUGGCAUUUCUCAGUCCACAGC ..((((..((((......((((((.......(((((((.(((........................))))))))))))))))((((((((....)))).))))........)))).)))) ( -36.57) >DroYak_CAF1 2074 120 - 1 GUGCUGACGGAUGCAUUCGAUUCAAAUUCAUUUUUGGCCAGCUAAAAAUACAUCCUCCGAAUACUUGCUGCCAAAAUAUUUCGCCAGGGAUGCAUCCCGUGGCAUUUCUCAGUCCACAGC ..(((...((((((((((((.......))..(((((((.(((........................))))))))))...........)))))))))).(((((........).))))))) ( -32.76) >consensus GUGCUGACGGAUGCAUUCGAUGCAAAUUCAUUUUUGGCCAGCUAAAAAUACACCCUCCGAAUACUUGCUGCCAAAAUAUAUCGCCAGGGAUGCAUCCCGUGGCAUUUCUCAGUCCACAGC ..(((...((((((((((((.......))..(((((((.(((........................))))))))))...........)))))))))).(((((........).))))))) (-30.63 = -30.96 + 0.33)

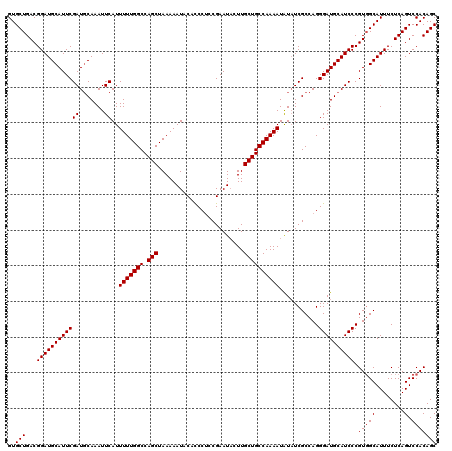
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:04:49 2006