
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 3,704,745 – 3,704,841 |
| Length | 96 |
| Max. P | 0.946293 |

| Location | 3,704,745 – 3,704,841 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 81.88 |
| Mean single sequence MFE | -16.86 |
| Consensus MFE | -10.42 |
| Energy contribution | -11.26 |
| Covariance contribution | 0.84 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.54 |
| Structure conservation index | 0.62 |
| SVM decision value | 1.36 |
| SVM RNA-class probability | 0.946293 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3704745 96 + 27905053 AAUGAAAAUUUCCGCUUAUGUAAGUACACCGAACUUACGUCAGCCAUGAUGGGAAACGAAACGCAAAGGAAAAUAAACGAAAAACAAAAUUAAAAU ........((((((((.((((((((.......)))))))).))).......(....)..........)))))........................ ( -17.90) >DroSec_CAF1 62741 96 + 1 AAUGAAAAUUUCCGCUUAUGUAAGUACACCGAACUUACGUCAGCCAUGACGUGAAACGAAACGCAAAGGAAAAGAAGCGAAAAACAAAAUUAAAAU ........((((((((.((((((((.......)))))))).)))..((.(((........)))))..)))))........................ ( -19.10) >DroSim_CAF1 65722 96 + 1 AAUGAAAAUUUCCGCUUAUGUAAGUACACCGAACUUACGUCAGCCAUGAUGGAAAACGAAACGCAAAGGAAAAGAAGCGAAAAACAAAAUUAAAAU ........((((((((.((((((((.......)))))))).)))......)))))......(((............)))................. ( -15.80) >DroEre_CAF1 64328 96 + 1 AAUGAAAAUUUCCGCUUAUAUAAGUACUCCGCACUUACGUCAUCCAUGAUGGAAAACGAAACGCAAGGGAAAACACACGAAAAACAAAAUUAAAAU ........(((((.(((...(((((.......)))))(((..(((.....)))..)))......))))))))........................ ( -13.50) >DroYak_CAF1 60917 72 + 1 AAGGAAAUUUUCCGCUUAUAAAAGUACACCGAACUUACGUCAGCCAUGAUGGAAAACGAAGUUCAAGGAAAA------------------------ .......((((((((((....)))).....((((((.(((...((.....))...)))))))))..))))))------------------------ ( -18.00) >consensus AAUGAAAAUUUCCGCUUAUGUAAGUACACCGAACUUACGUCAGCCAUGAUGGAAAACGAAACGCAAAGGAAAAGAAACGAAAAACAAAAUUAAAAU ........((((((((.((((((((.......)))))))).))).....((.....)).........)))))........................ (-10.42 = -11.26 + 0.84)

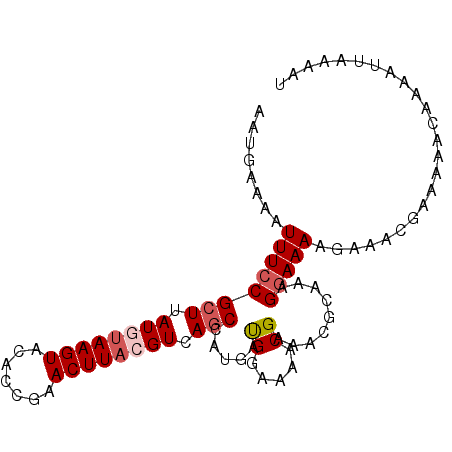
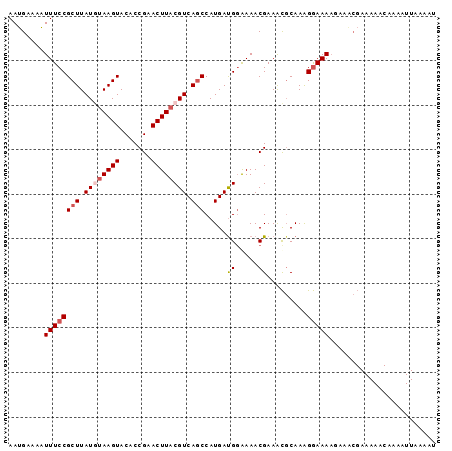
| Location | 3,704,745 – 3,704,841 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 81.88 |
| Mean single sequence MFE | -18.64 |
| Consensus MFE | -11.60 |
| Energy contribution | -11.36 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.715047 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3704745 96 - 27905053 AUUUUAAUUUUGUUUUUCGUUUAUUUUCCUUUGCGUUUCGUUUCCCAUCAUGGCUGACGUAAGUUCGGUGUACUUACAUAAGCGGAAAUUUUCAUU .............(((((((((((...............(((..((.....))..)))((((((.......)))))))))))))))))........ ( -18.60) >DroSec_CAF1 62741 96 - 1 AUUUUAAUUUUGUUUUUCGCUUCUUUUCCUUUGCGUUUCGUUUCACGUCAUGGCUGACGUAAGUUCGGUGUACUUACAUAAGCGGAAAUUUUCAUU .............(((((((((.....((...((((........))))...)).....((((((.......))))))..)))))))))........ ( -17.90) >DroSim_CAF1 65722 96 - 1 AUUUUAAUUUUGUUUUUCGCUUCUUUUCCUUUGCGUUUCGUUUUCCAUCAUGGCUGACGUAAGUUCGGUGUACUUACAUAAGCGGAAAUUUUCAUU .............(((((((((.................(((..((.....))..)))((((((.......))))))..)))))))))........ ( -18.40) >DroEre_CAF1 64328 96 - 1 AUUUUAAUUUUGUUUUUCGUGUGUUUUCCCUUGCGUUUCGUUUUCCAUCAUGGAUGACGUAAGUGCGGAGUACUUAUAUAAGCGGAAAUUUUCAUU .......((((((((...(((.((.((((...((((((((((.(((.....))).)))).)))))))))).)).)))..))))))))......... ( -20.30) >DroYak_CAF1 60917 72 - 1 ------------------------UUUUCCUUGAACUUCGUUUUCCAUCAUGGCUGACGUAAGUUCGGUGUACUUUUAUAAGCGGAAAAUUUCCUU ------------------------((((((((((((((((((..((.....))..)))).))))))))....(((....))).))))))....... ( -18.00) >consensus AUUUUAAUUUUGUUUUUCGCUUCUUUUCCUUUGCGUUUCGUUUUCCAUCAUGGCUGACGUAAGUUCGGUGUACUUACAUAAGCGGAAAUUUUCAUU ........................(((((((((......(((..((.....))..)))((((((.......)))))).)))).)))))........ (-11.60 = -11.36 + -0.24)

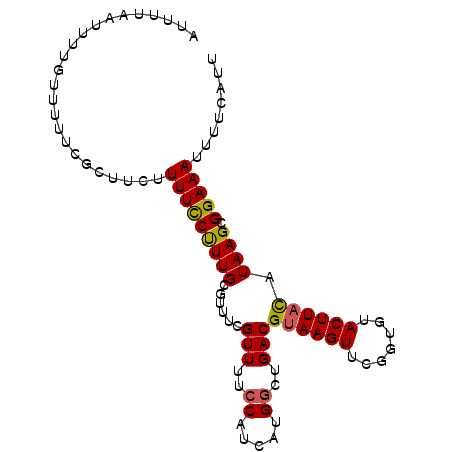
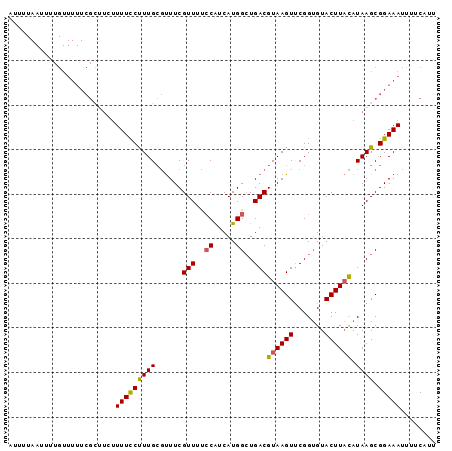
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:04:06 2006