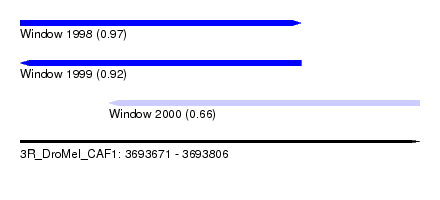
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 3,693,671 – 3,693,806 |
| Length | 135 |
| Max. P | 0.969377 |

| Location | 3,693,671 – 3,693,766 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 78.69 |
| Mean single sequence MFE | -18.57 |
| Consensus MFE | -13.40 |
| Energy contribution | -13.40 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.48 |
| Structure conservation index | 0.72 |
| SVM decision value | 1.64 |
| SVM RNA-class probability | 0.969377 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3693671 95 + 27905053 CUGUUUCACAUAUAUAUGUAAUUUAUUGU-----UUUUUGUUUU-GGU----UUUAG----UAUGAUAAAUAAAUGCUUAGCGACGACAAUUUGUUGUCGCACCUUCUU .......((((....))))..........-----..........-(((----...((----(((.........)))))..((((((((.....)))))))))))..... ( -17.70) >DroPse_CAF1 55200 86 + 1 CUUUAUCAGUUU---------UUUUUUG-------UUUUUUUUUUGGUU---UUUAG----UAUUUUAAAUAAAUGCUUAGCGACGACAAUUUGUUGUCGCACCUUCUU ............---------.......-------..........(((.---...((----(((((.....)))))))..((((((((.....)))))))))))..... ( -18.90) >DroSim_CAF1 53612 91 + 1 CUGUUUCACAUAUAU----AAUUAAUUGU-----UUUUGGUUUU-UGU----UUUAG----UAUGAUAAAUAAAUGCUUAGCGACGACAAUUUGUUGUCGCACCUUCUU ...............----..........-----....(((...-...----...((----(((.........)))))..((((((((.....)))))))))))..... ( -16.90) >DroEre_CAF1 51180 101 + 1 CUGUUUCACAUAUAU----AAUUUAUUGUACUUUUUUUUGUUUUCGGUUUUGUUUAG----UAUGAUAAAUAAAUGCUUAGCGACGACAAUUUGUUGUCGCACCUUCUU ...............----.((((((..((((.....(((....)))........))----))..)))))).........((((((((.....))))))))........ ( -20.82) >DroYak_CAF1 49662 97 + 1 CUGUUUCACAUAUAU----AAUUAAUUGUAUUUUUUU----UUUCGGUUUUGUUUAG----UAUGAUAAAUAAAUGCUUAGCGACGACAAUUUGUUGUCGCACCUUCUU ...........((((----((....))))))......----....(((((((((((.----.....))))))))......((((((((.....)))))))))))..... ( -19.20) >DroAna_CAF1 38513 88 + 1 UUGUUUCAUCACU-----------UUGGU-----UUUUGGUUUU-UUU----UUUAGUACGUAUGUGAAAUAAAUUCUUAGCGACGACAAUUUGUUGUCGCACCUUCUU ((((((((.((..-----------.((..-----..((((....-...----.))))..))..)))))))))).......((((((((.....))))))))........ ( -17.90) >consensus CUGUUUCACAUAUAU____AAUUUAUUGU_____UUUUUGUUUU_GGU____UUUAG____UAUGAUAAAUAAAUGCUUAGCGACGACAAUUUGUUGUCGCACCUUCUU ................................................................................((((((((.....))))))))........ (-13.40 = -13.40 + 0.00)



| Location | 3,693,671 – 3,693,766 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 78.69 |
| Mean single sequence MFE | -12.53 |
| Consensus MFE | -10.60 |
| Energy contribution | -10.60 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.15 |
| SVM RNA-class probability | 0.922459 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3693671 95 - 27905053 AAGAAGGUGCGACAACAAAUUGUCGUCGCUAAGCAUUUAUUUAUCAUA----CUAAA----ACC-AAAACAAAAA-----ACAAUAAAUUACAUAUAUAUGUGAAACAG .....((((((((.((.....)).)))))..........((((.....----.))))----)))-..........-----........((((((....))))))..... ( -14.00) >DroPse_CAF1 55200 86 - 1 AAGAAGGUGCGACAACAAAUUGUCGUCGCUAAGCAUUUAUUUAAAAUA----CUAAA---AACCAAAAAAAAAA-------CAAAAAA---------AAACUGAUAAAG .....((((((((.((.....)).)))))..((.((((.....)))).----))...---.)))..........-------.......---------............ ( -11.50) >DroSim_CAF1 53612 91 - 1 AAGAAGGUGCGACAACAAAUUGUCGUCGCUAAGCAUUUAUUUAUCAUA----CUAAA----ACA-AAAACCAAAA-----ACAAUUAAUU----AUAUAUGUGAAACAG .....((((((((.((.....)).)))))..........((((.....----.))))----...-...)))....-----..........----............... ( -10.80) >DroEre_CAF1 51180 101 - 1 AAGAAGGUGCGACAACAAAUUGUCGUCGCUAAGCAUUUAUUUAUCAUA----CUAAACAAAACCGAAAACAAAAAAAAGUACAAUAAAUU----AUAUAUGUGAAACAG .....((((((((((....)))))).))))..((((..((((((..((----((.......................))))..)))))).----....))))....... ( -15.00) >DroYak_CAF1 49662 97 - 1 AAGAAGGUGCGACAACAAAUUGUCGUCGCUAAGCAUUUAUUUAUCAUA----CUAAACAAAACCGAAA----AAAAAAAUACAAUUAAUU----AUAUAUGUGAAACAG .....((((((((.((.....)).)))))......(((.((((.....----.)))).))))))....----..................----............... ( -10.90) >DroAna_CAF1 38513 88 - 1 AAGAAGGUGCGACAACAAAUUGUCGUCGCUAAGAAUUUAUUUCACAUACGUACUAAA----AAA-AAAACCAAAA-----ACCAA-----------AGUGAUGAAACAA .....((((((((.((.....)).)))))...(((.....)))..............----...-...)))....-----.....-----------.((......)).. ( -13.00) >consensus AAGAAGGUGCGACAACAAAUUGUCGUCGCUAAGCAUUUAUUUAUCAUA____CUAAA____ACC_AAAACAAAAA_____ACAAUAAAUU____AUAUAUGUGAAACAG .....((((((((((....)))))).))))............................................................................... (-10.60 = -10.60 + -0.00)



| Location | 3,693,701 – 3,693,806 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 81.85 |
| Mean single sequence MFE | -18.35 |
| Consensus MFE | -14.55 |
| Energy contribution | -14.72 |
| Covariance contribution | 0.17 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.45 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.656113 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3693701 105 - 27905053 GGCUUGCCGACGGUGCAUAAAAAACGAACAACAAAUCAGAAAGAAGGUGCGACAACAAAUUGUCGUCGCUAAGCAUUUAUUU----A-UCAUA----CUAAA----ACC-AAAACAAAA .(((((.((((...((((.......((........)).........))))(((((....))))))))).)))))........----.-.....----.....----...-......... ( -18.59) >DroVir_CAF1 67220 94 - 1 GGCUUGCCAACGGUGCAUAAAAAACGUACAACAAAUCAGAAAGAAGGUGCGACAGCAAUAUGUCGUCGCUAAGCAUUUACA----UACUUAUA----CUA------------CA----- .(((((((....((((.........))))......((.....)).)))(((((.((.....)).))))).)))).......----........----...------------..----- ( -18.00) >DroGri_CAF1 24360 110 - 1 GGCUUGCCAACGGUGCAUAAAAAACGUACAACAAAUCAGAAAGAAGGUGCGACAGCAAUAUGUCGUCGCUAAGCAUUUACAUACAUACUUAUA----CUAAA----AGA-AAAAAAAAA .(((((((....((((.........))))......((.....)).)))(((((.((.....)).))))).))))...................----.....----...-......... ( -18.00) >DroYak_CAF1 49693 106 - 1 GGCUUGCCGACGGUGCAUAAAAAACGAACAACAAAUCAGAAAGAAGGUGCGACAACAAAUUGUCGUCGCUAAGCAUUUAUUU----A-UCAUA----CUAAACAAAACCGAAA----AA .(((((.((((...((((.......((........)).........))))(((((....))))))))).)))))........----.-.....----................----.. ( -18.59) >DroMoj_CAF1 72125 101 - 1 GGCAUGCCAACGGUGCAUAAAAAACGUACAACAAAUCAGAAAGAAGGUGCGACAGCAAUAUGUCGUCGCUAAGCAUUUACA----UACUUAUA----CUACA----AAA-UACA----- .((..(((....((((.........))))......((.....)).)))(((((.((.....)).)))))...)).......----........----.....----...-....----- ( -16.50) >DroAna_CAF1 38532 109 - 1 GGGUUGCCAACGGUGCAUAAAAAACGAACAACAAAUCAGAAAGAAGGUGCGACAACAAAUUGUCGUCGCUAAGAAUUUAUUU----C-ACAUACGUACUAAA----AAA-AAAACCAAA .((((......(((((.............................((((((((((....)))))).))))..(((.....))----)-......)))))...----...-..))))... ( -20.44) >consensus GGCUUGCCAACGGUGCAUAAAAAACGAACAACAAAUCAGAAAGAAGGUGCGACAACAAAAUGUCGUCGCUAAGCAUUUACAU____A_UCAUA____CUAAA____AAA_AAAA___AA ((....))...(((((...................((.....)).(((((((((......))))).))))..))))).......................................... (-14.55 = -14.72 + 0.17)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:04:01 2006