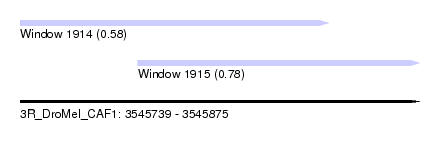
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 3,545,739 – 3,545,875 |
| Length | 136 |
| Max. P | 0.778777 |

| Location | 3,545,739 – 3,545,844 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 85.61 |
| Mean single sequence MFE | -25.52 |
| Consensus MFE | -17.21 |
| Energy contribution | -17.10 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.10 |
| Mean z-score | -3.11 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.581136 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
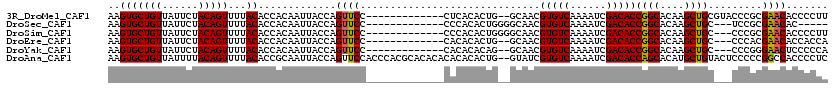
>3R_DroMel_CAF1 3545739 105 + 27905053 AAGUGCUGUUAUUCUACAGUUUUACACCACAAUUACCAGUUCC-------------CUCACACUG--GCAACGUGUCAAAAUCGACACCGGCACAAGCUGCGUACCCGCGAACACCCCUU ..((((((...........................(((((...-------------.....))))--)....(((((......)))))))))))....((((....)))).......... ( -23.60) >DroSec_CAF1 4638 99 + 1 AAGUGCUGUUAUUCUACAGUUUUACACCACAAUUACCAGUUCC-------------CCCACACUGGGGCAACGUGUCAAAAUCGACACCGGCACAAGCUGC---UCCGCGAACAC----- ..((((((..............................(((.(-------------(((.....)))).)))(((((......)))))))))))..((...---...))......----- ( -26.80) >DroSim_CAF1 4901 104 + 1 AAGUGCUGUUAUUCUACAGUUUUACACCACAAUUACCAGUUCC-------------CCCACACUGGGGCAACGUGUCAAAAUCGACACCGGCACAAGCUGC---CCCGCGAACACCCCUU ..((((((..............................(((.(-------------(((.....)))).)))(((((......)))))))))))..((...---...))........... ( -26.80) >DroEre_CAF1 4911 102 + 1 AAGUGCUGUUAUUCUACAGUUUUACACCACAAUUACCAGUUCC-------------CACACACUG--GCAACGUGUCAAAAUCGACACCGGCACAAGCUGC---CCCACGAACACCACCA ..(((.((((......((((((....((.......(((((...-------------.....))))--)....(((((......))))).))...)))))).---......)))).))).. ( -23.22) >DroYak_CAF1 5400 102 + 1 AAGUGCUGUUAUUCUACAGUUUUACACCACAAUUACCAGUUCC-------------CACACACAG--GCAACGUGUCAAAAUCGACACCGGCACAAGCUGC---CCCGGGAACUCCCCCA ..(((((((......)))))...))............((((((-------------(.......(--....)(((((......))))).((((.....)))---)..)))))))...... ( -27.10) >DroAna_CAF1 1564 118 + 1 AAGUGCUGUUAUUUUACAGUUUUACACCGCAAUUACCAGUUCCACCCACGCACACACACACACUG--GUAUCGUGUCAAAAUCGACACCAGCACAUGCUGUACUCCCCCGGCCACCCCUC ..((((((.........................(((((((.....................))))--)))..(((((......)))))))))))..((((........))))........ ( -25.60) >consensus AAGUGCUGUUAUUCUACAGUUUUACACCACAAUUACCAGUUCC_____________CACACACUG__GCAACGUGUCAAAAUCGACACCGGCACAAGCUGC___CCCGCGAACACCCCUA ..(((((((......)))))...)).............((((..............................(((((......)))))((((....)))).........))))....... (-17.21 = -17.10 + -0.11)

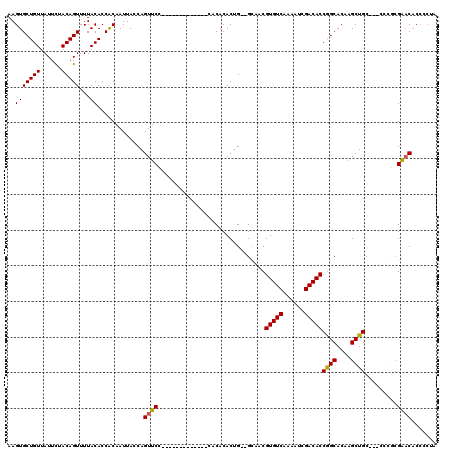
| Location | 3,545,779 – 3,545,875 |
|---|---|
| Length | 96 |
| Sequences | 3 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 82.13 |
| Mean single sequence MFE | -22.20 |
| Consensus MFE | -17.89 |
| Energy contribution | -18.67 |
| Covariance contribution | 0.78 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.778777 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
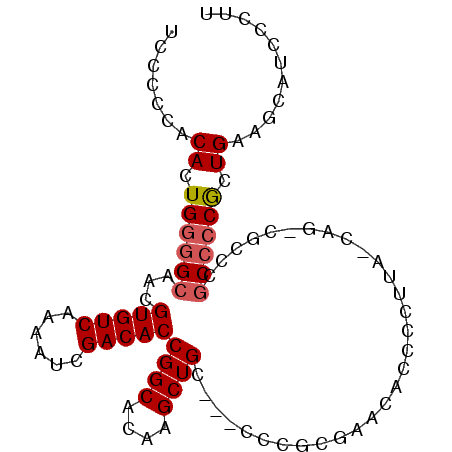
>3R_DroMel_CAF1 3545779 96 + 27905053 UCCCUCACACUG--GCAACGUGUCAAAAUCGACACCGGCACAAGCUGCGUACCCGCGAACACCCCUUAUCAGUCGCCCACCCCGCUGAAACAUCCCUU ...........(--((.(((((((......))))).(((....)))(((....)))...............)).)))..................... ( -16.40) >DroSec_CAF1 4678 81 + 1 UCCCCCACACUGGGGCAACGUGUCAAAAUCGACACCGGCACAAGCUGC---UCCGCGAACAC--------------CCGCCCCACUGAAGCAUCUCUU .......((.((((((...(((((......)))))((((....)))).---...........--------------..)))))).))........... ( -24.40) >DroSim_CAF1 4941 95 + 1 UCCCCCACACUGGGGCAACGUGUCAAAAUCGACACCGGCACAAGCUGC---CCCGCGAACACCCCUUACCAGCCGCCCGCCCCGCUGAAGCAUCCCUU .......((.((((((...(((((......))))).(((....((((.---..................)))).))).)))))).))........... ( -25.81) >consensus UCCCCCACACUGGGGCAACGUGUCAAAAUCGACACCGGCACAAGCUGC___CCCGCGAACACCCCUUA_CAG_CGCCCGCCCCGCUGAAGCAUCCCUU .......((.((((((...(((((......)))))((((....))))...............................)))))).))........... (-17.89 = -18.67 + 0.78)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:02:41 2006