
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 3,506,447 – 3,506,592 |
| Length | 145 |
| Max. P | 0.991255 |

| Location | 3,506,447 – 3,506,540 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 79.42 |
| Mean single sequence MFE | -31.00 |
| Consensus MFE | -21.12 |
| Energy contribution | -24.18 |
| Covariance contribution | 3.06 |
| Combinations/Pair | 1.21 |
| Mean z-score | -2.09 |
| Structure conservation index | 0.68 |
| SVM decision value | 1.23 |
| SVM RNA-class probability | 0.933226 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3506447 93 + 27905053 CCUCUUGUGUGUUCGCGGGCUCAAACUCAAUCCGGCUCAGCGAUUGUUUUCGGCCAUCCUGUGAUGGCACACAUGCACUCGUCCUCGGACGAG ......(((((((((((((((............))))).)))).........((((((....))))))))))))...((((((....)))))) ( -33.80) >DroSim_CAF1 128646 93 + 1 CCUCUUGUGUGUACGCGGGCUCAAGCUCAAUCCGGCUCAGCGAUUGUUUUGGGCCAUCCUGUGAUGGCCCACAUGCACUCGUCCUCGGACGAG .....(((((((.((((((((............))))).)))........((((((((....)))))))))))))))((((((....)))))) ( -39.70) >DroEre_CAF1 113807 90 + 1 CCUCUUGUGUGUACGCGAGCUCAAACUCAAUCCAGCUCAGCGAUUGUUUUG-GCCAUCCUCUGAUGGCCCACAUGCACUCGUCCUCGGACG-- .....(((((((.((((((((............))))).)))........(-((((((....))))))).)))))))..((((....))))-- ( -31.50) >DroYak_CAF1 169894 93 + 1 CCUCUUGUGUGUACGCGAGCUCAAACUCAAUCUGGCUCAGCGAUUGUUUUGGGCCAUCAUGUGAUGCCCCACAUGCACUCCUCCUCGGACGAG .....(((((((.(((((((.((.........)))))).)))........(((.((((....)))).))))))))))(((.((....)).))) ( -29.00) >DroAna_CAF1 98493 72 + 1 CCUCUUGUGUGUACACGAGCUCAAACUCAAUCUGG-------AUUGUUCGGGCCUGGCCUGUUCCGAGCCAC--GCCCUCG------------ ...((((((....))))))........((((....-------))))...((((.(((((......).)))).--))))...------------ ( -21.00) >consensus CCUCUUGUGUGUACGCGAGCUCAAACUCAAUCCGGCUCAGCGAUUGUUUUGGGCCAUCCUGUGAUGGCCCACAUGCACUCGUCCUCGGACGAG .....(((((((.((((((((............))))).)))........((((((((....)))))))))))))))((((((....)))))) (-21.12 = -24.18 + 3.06)



| Location | 3,506,447 – 3,506,540 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 79.42 |
| Mean single sequence MFE | -33.00 |
| Consensus MFE | -17.52 |
| Energy contribution | -19.58 |
| Covariance contribution | 2.06 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.53 |
| SVM decision value | 0.61 |
| SVM RNA-class probability | 0.798724 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3506447 93 - 27905053 CUCGUCCGAGGACGAGUGCAUGUGUGCCAUCACAGGAUGGCCGAAAACAAUCGCUGAGCCGGAUUGAGUUUGAGCCCGCGAACACACAAGAGG ((((((....))))))....(((((((((((....)))))).........((((.(.(((((((...))))).))).))))..)))))..... ( -32.60) >DroSim_CAF1 128646 93 - 1 CUCGUCCGAGGACGAGUGCAUGUGGGCCAUCACAGGAUGGCCCAAAACAAUCGCUGAGCCGGAUUGAGCUUGAGCCCGCGUACACACAAGAGG .(((((....)))))(((.((((((((((((....)))))))).........(((((((........)))).)))..)))).)))........ ( -35.80) >DroEre_CAF1 113807 90 - 1 --CGUCCGAGGACGAGUGCAUGUGGGCCAUCAGAGGAUGGC-CAAAACAAUCGCUGAGCUGGAUUGAGUUUGAGCUCGCGUACACACAAGAGG --((((....)))).((((.(((.(((((((....))))))-)...)))...((.(((((.((......)).))))))))))).......... ( -36.90) >DroYak_CAF1 169894 93 - 1 CUCGUCCGAGGAGGAGUGCAUGUGGGGCAUCACAUGAUGGCCCAAAACAAUCGCUGAGCCAGAUUGAGUUUGAGCUCGCGUACACACAAGAGG (((.(((.....)))((((...((((.((((....)))).))))........((.(((((((((...))))).))))))))))......))). ( -34.60) >DroAna_CAF1 98493 72 - 1 ------------CGAGGGC--GUGGCUCGGAACAGGCCAGGCCCGAACAAU-------CCAGAUUGAGUUUGAGCUCGUGUACACACAAGAGG ------------...((((--.(((((.......))))).)))).......-------((.....((((....))))(((....)))....)) ( -25.10) >consensus CUCGUCCGAGGACGAGUGCAUGUGGGCCAUCACAGGAUGGCCCAAAACAAUCGCUGAGCCGGAUUGAGUUUGAGCUCGCGUACACACAAGAGG .(((((....)))))(((.((((((((((((....))))))))....(((((.........)))))...........)))).)))........ (-17.52 = -19.58 + 2.06)



| Location | 3,506,500 – 3,506,592 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 77.39 |
| Mean single sequence MFE | -31.26 |
| Consensus MFE | -22.21 |
| Energy contribution | -23.46 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.71 |
| SVM decision value | 1.66 |
| SVM RNA-class probability | 0.970551 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3506500 92 + 27905053 CCAUCCUGUGAUGGCACACAUGCACUCGUCCUCGGACGAGCGGUCCGACCCUGCGCUGACCACGCCCCUUUCGGGGCGAUCUUCCGCUGACA (((((....)))))....((.((.((((((....)))))).((((((......))..)))).((((((....)))))).......))))... ( -35.10) >DroSec_CAF1 103984 71 + 1 ---------------------GCACUCGUCCUCGGACAAGCGGUCCGCCCCUGCGGUGACCACGCCCCUUUCGGGGCGAUCUUCCGCUGACA ---------------------......(((..((((.....((((.(((.....))))))).((((((....))))))....))))..))). ( -30.80) >DroSim_CAF1 128699 92 + 1 CCAUCCUGUGAUGGCCCACAUGCACUCGUCCUCGGACGAGCGGUCCGCCCCUGCGCUGACCACGCCCCUUUCGGGGCGAUCUUCCGCUGACA (((((....)))))....((.((.((((((....)))))).((((.((......)).)))).((((((....)))))).......))))... ( -38.10) >DroEre_CAF1 113859 75 + 1 CCAUCCUCUGAUGGCCCACAUGCACUCGUCCUCGGACG-----------------CCGACCACGCCCCUUUUGGGGCGCUCUCCCGCUGACA (((((....)))))....((.((...((((....))))-----------------.......((((((....)))))).......))))... ( -25.10) >DroYak_CAF1 169947 92 + 1 CCAUCAUGUGAUGCCCCACAUGCACUCCUCCUCGGACGAGCCAUCCGCCCCUGAGCUGACCACGCCCCUUUCGGGGCGAUCUUUCGCUGACA ....((((((......)))))).........((((.((((...((.((......)).))...((((((....))))))....)))))))).. ( -27.20) >consensus CCAUCCUGUGAUGGCCCACAUGCACUCGUCCUCGGACGAGCGGUCCGCCCCUGCGCUGACCACGCCCCUUUCGGGGCGAUCUUCCGCUGACA (((((....))))).......((.((((((....))))))......................((((((....)))))).......))..... (-22.21 = -23.46 + 1.25)



| Location | 3,506,500 – 3,506,592 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 77.39 |
| Mean single sequence MFE | -38.82 |
| Consensus MFE | -28.31 |
| Energy contribution | -29.60 |
| Covariance contribution | 1.29 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.35 |
| Structure conservation index | 0.73 |
| SVM decision value | 2.26 |
| SVM RNA-class probability | 0.991255 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3506500 92 - 27905053 UGUCAGCGGAAGAUCGCCCCGAAAGGGGCGUGGUCAGCGCAGGGUCGGACCGCUCGUCCGAGGACGAGUGCAUGUGUGCCAUCACAGGAUGG ......(....)((((((((....)))))(((((..(((((((......))(((((((....))))))).....))))).)))))..))).. ( -42.50) >DroSec_CAF1 103984 71 - 1 UGUCAGCGGAAGAUCGCCCCGAAAGGGGCGUGGUCACCGCAGGGGCGGACCGCUUGUCCGAGGACGAGUGC--------------------- .(((..((((.....(((((....)))))(((((..((((....)))))))))...))))..)))......--------------------- ( -38.40) >DroSim_CAF1 128699 92 - 1 UGUCAGCGGAAGAUCGCCCCGAAAGGGGCGUGGUCAGCGCAGGGGCGGACCGCUCGUCCGAGGACGAGUGCAUGUGGGCCAUCACAGGAUGG .(((..((((.((..(((((....)))))((((((.((......)).)))))))).))))..))).............(((((....))))) ( -44.30) >DroEre_CAF1 113859 75 - 1 UGUCAGCGGGAGAGCGCCCCAAAAGGGGCGUGGUCGG-----------------CGUCCGAGGACGAGUGCAUGUGGGCCAUCAGAGGAUGG .....(((.((..(((((((....)))))))..))..-----------------((((....))))..))).......(((((....))))) ( -30.30) >DroYak_CAF1 169947 92 - 1 UGUCAGCGAAAGAUCGCCCCGAAAGGGGCGUGGUCAGCUCAGGGGCGGAUGGCUCGUCCGAGGAGGAGUGCAUGUGGGGCAUCACAUGAUGG .(((.((....(((((((((....))))))..))).((((.....((((((...)))))).....))))))((((((....))))))))).. ( -38.60) >consensus UGUCAGCGGAAGAUCGCCCCGAAAGGGGCGUGGUCAGCGCAGGGGCGGACCGCUCGUCCGAGGACGAGUGCAUGUGGGCCAUCACAGGAUGG .....((....(((((((((....))))))..)))................(((((((....))))))))).......(((((....))))) (-28.31 = -29.60 + 1.29)

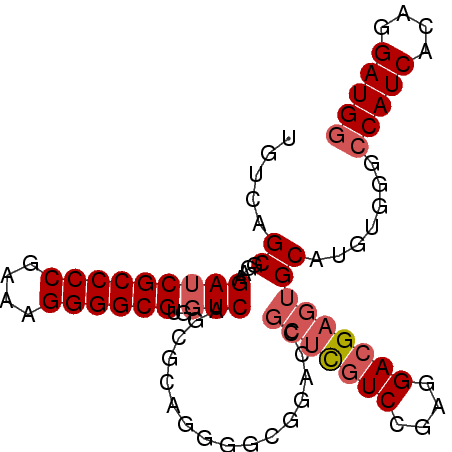

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:02:30 2006