
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 3,484,442 – 3,484,542 |
| Length | 100 |
| Max. P | 0.978589 |

| Location | 3,484,442 – 3,484,542 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 85.69 |
| Mean single sequence MFE | -21.83 |
| Consensus MFE | -16.88 |
| Energy contribution | -16.50 |
| Covariance contribution | -0.38 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.97 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.861285 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3484442 100 + 27905053 GGC-----CAAAUAGCCCGUGAAAAUACAUUUCCAAAAGAAAUAAUGUUGUCUUCGUAUACGAAAUUCCACGGCUAUUUGGGUUAAUUAAGAAUCGAAAAUGCUU ..(-----(((((((((.(((.......(((((.....))))).........((((....))))....)))))))))))))........................ ( -23.20) >DroSec_CAF1 82032 100 + 1 GGC-----CAAAUAGCCCGUGAAAAUACAUUACCAAAGGAAAUAAUGUUGUCCUCGUAUAUAAAAUUCCACGGCUAUUUGGGUUAAUUAAGAACCAAAAAUGCUC (((-----(((((((((.(((.........(((...((((.(......).)))).)))..........))))))))))))((((.......))))......))). ( -21.91) >DroSim_CAF1 105830 100 + 1 GGC-----CAAAUAGCCCGUGAAAAUACAUUUCCAAAAGAAAUAAUGUUGUCCUCGUAUAUGAAAUUCCACGGCUAUUUGGGUUAAUUAAGAACCAAAAAUGCUC (((-----(((((((((.(((.......(((((.....))))).(((.......)))...........))))))))))))((((.......))))......))). ( -20.60) >DroYak_CAF1 146728 101 + 1 GGCAAAUGCAAAUAGCUCGUGAAAAUACAUAUUCGAACGAAAUAAUGUUGUUCUCAUAUACGAAGUACCACGGCUAUUUGGGUUAAUUAAAAAUCGAA--UCG-- ....(((.((((((((.((((....(((......((((((.......))))))((......)).))).)))))))))))).)))..............--...-- ( -21.60) >consensus GGC_____CAAAUAGCCCGUGAAAAUACAUUUCCAAAAGAAAUAAUGUUGUCCUCGUAUACGAAAUUCCACGGCUAUUUGGGUUAAUUAAGAACCAAAAAUGCUC ........((((((((.((((.......(((((.....)))))....((((........)))).....))))))))))))((((.......)))).......... (-16.88 = -16.50 + -0.38)

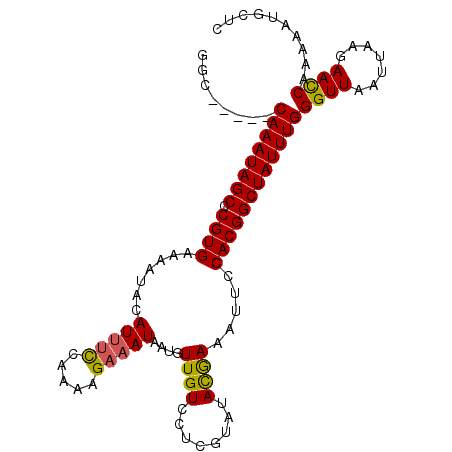

| Location | 3,484,442 – 3,484,542 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 85.69 |
| Mean single sequence MFE | -26.15 |
| Consensus MFE | -19.50 |
| Energy contribution | -18.87 |
| Covariance contribution | -0.63 |
| Combinations/Pair | 1.26 |
| Mean z-score | -2.82 |
| Structure conservation index | 0.75 |
| SVM decision value | 1.82 |
| SVM RNA-class probability | 0.978589 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3484442 100 - 27905053 AAGCAUUUUCGAUUCUUAAUUAACCCAAAUAGCCGUGGAAUUUCGUAUACGAAGACAACAUUAUUUCUUUUGGAAAUGUAUUUUCACGGGCUAUUUG-----GCC ........................(((((((((((((((((((((....))))).......((((((.....))))))...)))))).)))))))))-----).. ( -27.40) >DroSec_CAF1 82032 100 - 1 GAGCAUUUUUGGUUCUUAAUUAACCCAAAUAGCCGUGGAAUUUUAUAUACGAGGACAACAUUAUUUCCUUUGGUAAUGUAUUUUCACGGGCUAUUUG-----GCC ((((.......)))).........((((((((((((((((...((((((((((((..........)))))..)).))))).)))))).)))))))))-----).. ( -29.20) >DroSim_CAF1 105830 100 - 1 GAGCAUUUUUGGUUCUUAAUUAACCCAAAUAGCCGUGGAAUUUCAUAUACGAGGACAACAUUAUUUCUUUUGGAAAUGUAUUUUCACGGGCUAUUUG-----GCC ((((.......)))).........((((((((((((((((((((......)))).......((((((.....))))))...)))))).)))))))))-----).. ( -26.60) >DroYak_CAF1 146728 101 - 1 --CGA--UUCGAUUUUUAAUUAACCCAAAUAGCCGUGGUACUUCGUAUAUGAGAACAACAUUAUUUCGUUCGAAUAUGUAUUUUCACGAGCUAUUUGCAUUUGCC --...--..................(((((((((((((.....(((((....((((...........))))..))))).....))))).))))))))........ ( -21.40) >consensus GAGCAUUUUCGAUUCUUAAUUAACCCAAAUAGCCGUGGAAUUUCAUAUACGAGGACAACAUUAUUUCUUUUGGAAAUGUAUUUUCACGGGCUAUUUG_____GCC .........................(((((((((((((((...(((...(((((.............)))))...)))...))))))).))))))))........ (-19.50 = -18.87 + -0.63)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:02:23 2006