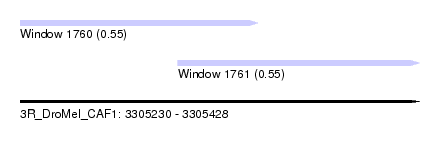
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 3,305,230 – 3,305,428 |
| Length | 198 |
| Max. P | 0.553571 |

| Location | 3,305,230 – 3,305,348 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.23 |
| Mean single sequence MFE | -29.89 |
| Consensus MFE | -25.67 |
| Energy contribution | -26.05 |
| Covariance contribution | 0.38 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.42 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.552982 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
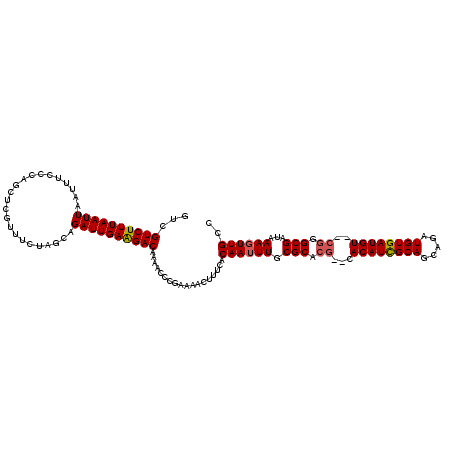
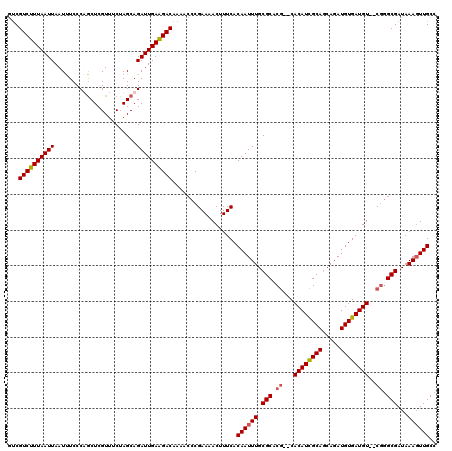
>3R_DroMel_CAF1 3305230 118 + 27905053 CUCGUCUUUAAUUAAUUUCCCUACUCGUUUCUAGCAGAUUGAGGACAAAACCCGAAAACUUUCACAAUUUGCGCACG--CACAUCGCAACAGAUGUGAUGUGGCGGGCGAUAAAGUUGCC ...((((((((((.......................))))))))))..................((((((.(((.((--(((((((((.....)))))))).))).)))...)))))).. ( -31.60) >DroSec_CAF1 9290 116 + 1 GUCGUCUUUAAUUAAUUUCCCAGCUCGUUUCUAGCAGAUUGAAGACAAAACCCGAAAACUUUCACAAUUUGCGCACG--CACAUCGCAGCAGAUGUGAUGU--CGGGCGAUAAAGUUGCC ...((((((((((.........(((.......))).))))))))))..................((((((.(((.((--.((((((((.....))))))))--)).)))...)))))).. ( -29.90) >DroSim_CAF1 12899 116 + 1 GUCGUCUUUAAUUAAUUUCCCAGCUCGUUUCUAGCAGAUUGAAGACAAAACCCGAAAACUUUCACAAUUUGCGCACG--CACAUCGCAGCAGAUGUGAUGU--CGGGCGAUAAAGUUGCC ...((((((((((.........(((.......))).))))))))))..................((((((.(((.((--.((((((((.....))))))))--)).)))...)))))).. ( -29.90) >DroYak_CAF1 19497 118 + 1 GCCGUCUUUAAUUAAUCUCCCUGCUCGUUUCUAGCAGAUUGAAGACGAACCCCGAAAACUUUCACAAUUUGCGCACAUUCACAUUGCAGCAGAUGUGAUGU--CAUGCGAUAAACUUGCC ((((((((((((........(((((.......))))))))))))))).....((.......(((((.((((((((.........))).))))))))))...--....))........)). ( -28.14) >consensus GUCGUCUUUAAUUAAUUUCCCAGCUCGUUUCUAGCAGAUUGAAGACAAAACCCGAAAACUUUCACAAUUUGCGCACG__CACAUCGCAGCAGAUGUGAUGU__CGGGCGAUAAAGUUGCC ...((((((((((.......................))))))))))..................((((((.(((.((...((((((((.....))))))))..)).)))...)))))).. (-25.67 = -26.05 + 0.38)

| Location | 3,305,308 – 3,305,428 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.70 |
| Mean single sequence MFE | -30.88 |
| Consensus MFE | -24.27 |
| Energy contribution | -25.40 |
| Covariance contribution | 1.13 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.553571 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
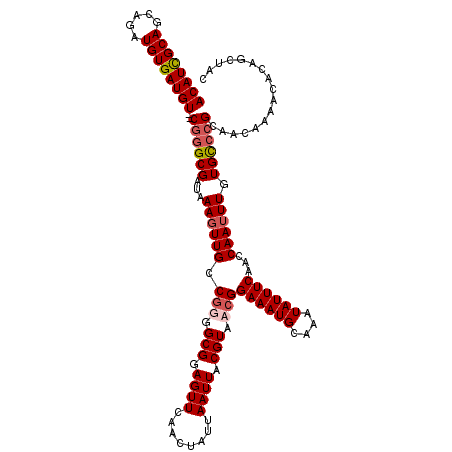
>3R_DroMel_CAF1 3305308 120 + 27905053 ACAUCGCAACAGAUGUGAUGUGGCGGGCGAUAAAGUUGCCGGGGCGGAGUUCAACUAUUAAUUACGUAACGGAAAUGCAAAUAUUUCAACCAAUUUGUGCCCGCAACAAGACACAGCUAC ((((((((.....)))))))).(((((((...((((((.((..(((.((((........)))).)))..))((((((....))))))...)))))).)))))))................ ( -36.20) >DroSec_CAF1 9368 118 + 1 ACAUCGCAGCAGAUGUGAUGU--CGGGCGAUAAAGUUGCCGUAGCGGAGUUCAACUAUUAAUUACGUAACGGAAAUGCAAAUAUUUCAACCAAUUUGUGUCCGCAACAAAACACAGAUAC ((((((((.....))))))))--((((((...((((((.(((.(((.((((........)))).))).)))((((((....))))))...)))))).))))))................. ( -28.70) >DroSim_CAF1 12977 118 + 1 ACAUCGCAGCAGAUGUGAUGU--CGGGCGAUAAAGUUGCCGUUGCGGAGUUCAACUAUUAAUUACGUAACGGAAAUGCAAAUAUUUCAACCAAUUUGUGCCCGCAACAAAACACAGCUAC ((((((((.....))))))))--((((((...((((((.(((((((.((((........)))).)))))))((((((....))))))...)))))).))))))................. ( -35.00) >DroYak_CAF1 19577 118 + 1 ACAUUGCAGCAGAUGUGAUGU--CAUGCGAUAAACUUGCCAGGGCGGAGUUCAACUAUUAAUUACGUAACGGAAAUGCAAAUAUUUCAACCAAUUUGUGCCCGCAACAAAACACAUCUAC ..........(((((((.(((--(....))))...((((..(((((.((...............((...))((((((....)))))).......)).))))))))).....))))))).. ( -23.60) >consensus ACAUCGCAGCAGAUGUGAUGU__CGGGCGAUAAAGUUGCCGGGGCGGAGUUCAACUAUUAAUUACGUAACGGAAAUGCAAAUAUUUCAACCAAUUUGUGCCCGCAACAAAACACAGCUAC ((((((((.....))))))))..((((((...((((((.(((.(((.((((........)))).))).)))((((((....))))))...)))))).))))))................. (-24.27 = -25.40 + 1.13)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:00:17 2006