
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 3,304,951 – 3,305,071 |
| Length | 120 |
| Max. P | 0.828697 |

| Location | 3,304,951 – 3,305,071 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.55 |
| Mean single sequence MFE | -27.31 |
| Consensus MFE | -25.80 |
| Energy contribution | -26.11 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.71 |
| SVM RNA-class probability | 0.828697 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
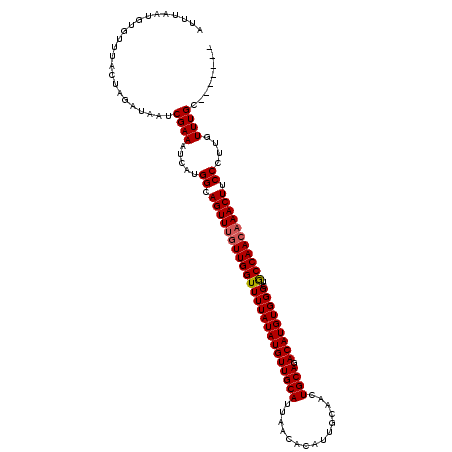
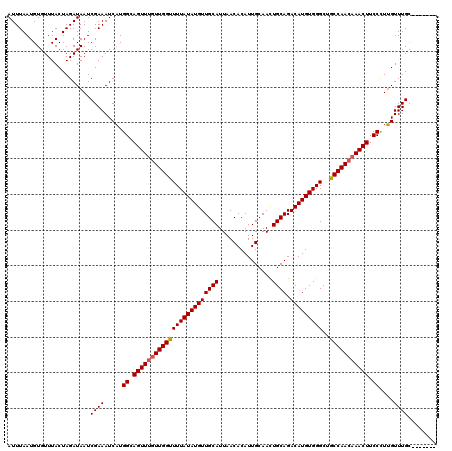
>3R_DroMel_CAF1 3304951 120 + 27905053 AUUUAAUGUGUUUACUAGAUAAUCGAAAUCUUGGCAGUUUGUUGGUUUUAUAUGUUGCAUUAACACAUUGCAACUGCAGACAUGUGGGCUGCCAACAAACUUCCCUAGUUUGCCACCCAA ......((.((..(((((......(....)..((.((((((((((((((((((((((((...............)))).)))))))))..))))))))))).)))))))..))))..... ( -32.06) >DroSec_CAF1 9025 113 + 1 AUUUAAUGUGUUUACUAGAUAAUCGAAAUCAUGGCAGUUUGUUGGUUUUAUAUGUUGCAUUAACACAUUGCAACUGCAGACAUGUGGGCUACCAACAAACUUCCCUUGUUUGC------- .......................((((.....((.((((((((((((((((((((((((...............)))).)))))))))..))))))))))).))....)))).------- ( -26.76) >DroSim_CAF1 12634 113 + 1 AUUUAAUGUGUUUACUAGAUAAUCGAAAUCAUGGCAGUUUGUUGGUUUUAUAUGUUGCAUUAACACAUUGCAACUGCAGACAUGUGGGCUGCCAACAAACUUCCCUUGUUUGC------- .......................((((.....((.((((((((((((((((((((((((...............)))).)))))))))..))))))))))).))....)))).------- ( -27.06) >DroYak_CAF1 19222 119 + 1 AUUUAAUGUGUUUACUAGAUAAUCGAAAUCAUGGCAGUUUCUUGGUUUUAUAUGUUGCAUUAACACAUUGCAAUUGCAGACAUGUGGGCUGCCAACCAACUUCCUUUGUUUGUC-CCUUA .................(((((.((((.....((.((((..((((((((((((((((((...............)))).)))))))))..)))))..)))).)))))).)))))-..... ( -23.36) >consensus AUUUAAUGUGUUUACUAGAUAAUCGAAAUCAUGGCAGUUUGUUGGUUUUAUAUGUUGCAUUAACACAUUGCAACUGCAGACAUGUGGGCUGCCAACAAACUUCCCUUGUUUGC_______ .......................((((.....((.((((((((((((((((((((((((...............)))).)))))))))..))))))))))).))....))))........ (-25.80 = -26.11 + 0.31)


| Location | 3,304,951 – 3,305,071 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.55 |
| Mean single sequence MFE | -25.68 |
| Consensus MFE | -21.64 |
| Energy contribution | -22.20 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.706741 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
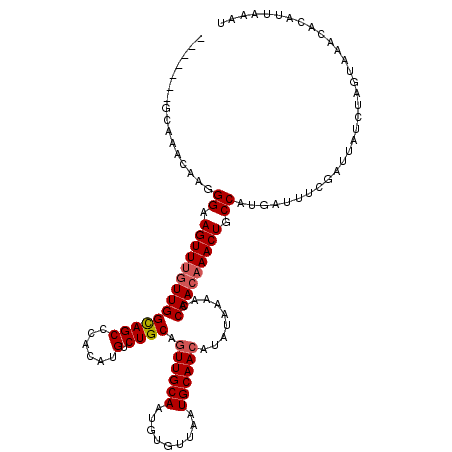
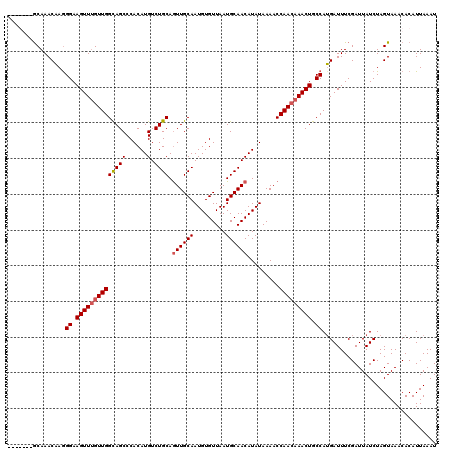
>3R_DroMel_CAF1 3304951 120 - 27905053 UUGGGUGGCAAACUAGGGAAGUUUGUUGGCAGCCCACAUGUCUGCAGUUGCAAUGUGUUAAUGCAACAUAUAAAACCAACAAACUGCCAAGAUUUCGAUUAUCUAGUAAACACAUUAAAU ....(((....(((((((.((((((((((((((......).)))).((((((.........)))))).........))))))))).))..(....)......)))))...)))....... ( -31.70) >DroSec_CAF1 9025 113 - 1 -------GCAAACAAGGGAAGUUUGUUGGUAGCCCACAUGUCUGCAGUUGCAAUGUGUUAAUGCAACAUAUAAAACCAACAAACUGCCAUGAUUUCGAUUAUCUAGUAAACACAUUAAAU -------.....((.((..(((((((((((.......((((.((((...((.....))...)))))))).....))))))))))).)).))............................. ( -25.30) >DroSim_CAF1 12634 113 - 1 -------GCAAACAAGGGAAGUUUGUUGGCAGCCCACAUGUCUGCAGUUGCAAUGUGUUAAUGCAACAUAUAAAACCAACAAACUGCCAUGAUUUCGAUUAUCUAGUAAACACAUUAAAU -------.....((.((..((((((((((((((......).)))).((((((.........)))))).........))))))))).)).))............................. ( -25.50) >DroYak_CAF1 19222 119 - 1 UAAGG-GACAAACAAAGGAAGUUGGUUGGCAGCCCACAUGUCUGCAAUUGCAAUGUGUUAAUGCAACAUAUAAAACCAAGAAACUGCCAUGAUUUCGAUUAUCUAGUAAACACAUUAAAU .....-...........((((((...((((((..........(((....)))(((((((.....)))))))............)))))).))))))........................ ( -20.20) >consensus _______GCAAACAAGGGAAGUUUGUUGGCAGCCCACAUGUCUGCAGUUGCAAUGUGUUAAUGCAACAUAUAAAACCAACAAACUGCCAUGAUUUCGAUUAUCUAGUAAACACAUUAAAU ................((.((((((((((((((......).)))).((((((.........)))))).........))))))))).))................................ (-21.64 = -22.20 + 0.56)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:00:15 2006