
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 3,304,227 – 3,304,335 |
| Length | 108 |
| Max. P | 0.715149 |

| Location | 3,304,227 – 3,304,335 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 83.97 |
| Mean single sequence MFE | -27.27 |
| Consensus MFE | -18.66 |
| Energy contribution | -18.35 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.715149 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
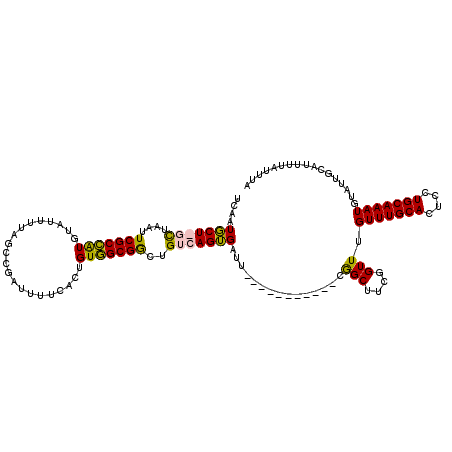
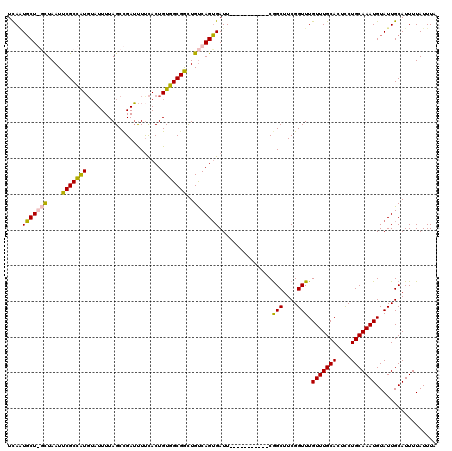
>3R_DroMel_CAF1 3304227 108 + 27905053 UCAAUGCU-UCUAAUUCGCCAUGUAUUUUAGCCGAUUUUCACUGUGGCGGCUGUCAGUGAUU-----------CAGCUUCGGUUUGUUUGCACUCCUGCAAAUGUAUUGCAUUUUAUUUA .((((((.-....................((((((...((((((..........))))))..-----------.....)))))).(((((((....)))))))))))))........... ( -22.90) >DroSec_CAF1 8304 109 + 1 UCAUUGCUGGCUAAUUCGCCAUGUAUUUUAGCCGAUUUUCACUGUGGCGGCUGUCAGUGAUU-----------CGGCUUCGGUUUGUUUGCACUCCUGCAAAUGUAUUGCAUUUUAUUUA ....(((((((......)))).)))....((((((...((((((..........)))))).)-----------))))).((((.((((((((....)))))))).))))........... ( -30.70) >DroSim_CAF1 11913 109 + 1 UCAGUGCUGGCUAAUUCGCCGUGUAUUUUAGCCGAUUUCCACUGUGGCGGCUGUCAGUGAUU-----------CGGCUUCGGUUUGUUUGCACUCCUGCAAAUGUAUUGCAUUUUAUUUU ..(((((..((.((((.((((..((((.((((((....((.....))))))))..))))...-----------))))...)))).))..)))))..(((((.....)))))......... ( -29.50) >DroYak_CAF1 18516 118 + 1 UUAAUUCU-CUUAAUUCGCAAUU-UUUUUAGCUGAACUUUACUGUUGCGAUUGCCAGGGAUUCGGUAUCGUUUCGGCUUCGGUUUGUUUGCACUUCUGCAAAUGUAUUGCAUUUUAUUUA ........-........(((((.-.....(((((((.....(((..(((((.(((........))))))))..))).))))))).(((((((....)))))))..))))).......... ( -26.00) >consensus UCAAUGCU_GCUAAUUCGCCAUGUAUUUUAGCCGAUUUUCACUGUGGCGGCUGUCAGUGAUU___________CGGCUUCGGUUUGUUUGCACUCCUGCAAAUGUAUUGCAUUUUAUUUA ....(((((((....(((((((.....................)))))))..)))))))....................((((.((((((((....)))))))).))))........... (-18.66 = -18.35 + -0.31)

| Location | 3,304,227 – 3,304,335 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.97 |
| Mean single sequence MFE | -25.82 |
| Consensus MFE | -13.77 |
| Energy contribution | -15.83 |
| Covariance contribution | 2.06 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.79 |
| Structure conservation index | 0.53 |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.652271 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
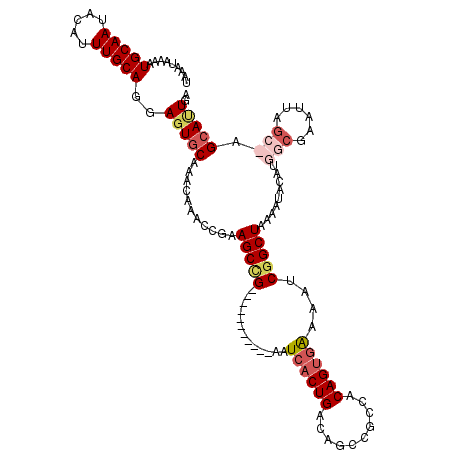
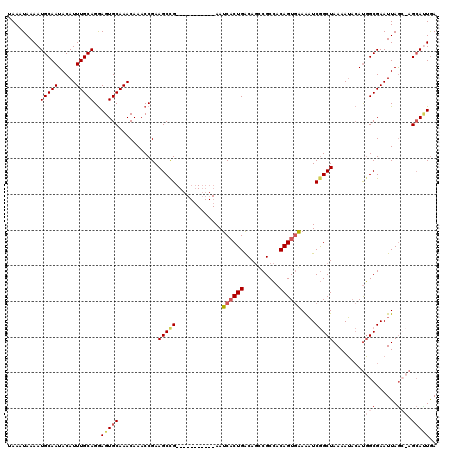
>3R_DroMel_CAF1 3304227 108 - 27905053 UAAAUAAAAUGCAAUACAUUUGCAGGAGUGCAAACAAACCGAAGCUG-----------AAUCACUGACAGCCGCCACAGUGAAAAUCGGCUAAAAUACAUGGCGAAUUAGA-AGCAUUGA .......(((((.....((((((.((..((....))..))..(((((-----------(.((((((..........))))))...))))))..........))))))....-.))))).. ( -25.20) >DroSec_CAF1 8304 109 - 1 UAAAUAAAAUGCAAUACAUUUGCAGGAGUGCAAACAAACCGAAGCCG-----------AAUCACUGACAGCCGCCACAGUGAAAAUCGGCUAAAAUACAUGGCGAAUUAGCCAGCAAUGA .........(((((.....)))))....(((...........(((((-----------(.((((((..........))))))...))))))........((((......))))))).... ( -29.60) >DroSim_CAF1 11913 109 - 1 AAAAUAAAAUGCAAUACAUUUGCAGGAGUGCAAACAAACCGAAGCCG-----------AAUCACUGACAGCCGCCACAGUGGAAAUCGGCUAAAAUACACGGCGAAUUAGCCAGCACUGA .........(((((.....)))))..(((((...........(((((-----------(.((((((..........))))))...)))))).........(((......))).))))).. ( -32.40) >DroYak_CAF1 18516 118 - 1 UAAAUAAAAUGCAAUACAUUUGCAGAAGUGCAAACAAACCGAAGCCGAAACGAUACCGAAUCCCUGGCAAUCGCAACAGUAAAGUUCAGCUAAAAA-AAUUGCGAAUUAAG-AGAAUUAA .........((((((...((((((....))))))........(((.(((.((((.(((......)))..))))...........))).))).....-.)))))).......-........ ( -16.10) >consensus UAAAUAAAAUGCAAUACAUUUGCAGGAGUGCAAACAAACCGAAGCCG___________AAUCACUGACAGCCGCCACAGUGAAAAUCGGCUAAAAUACAUGGCGAAUUAGC_AGCAUUGA .........(((((.....)))))..(((((...........(((((.............((((((..........))))))....))))).........(((......))).))))).. (-13.77 = -15.83 + 2.06)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:00:11 2006