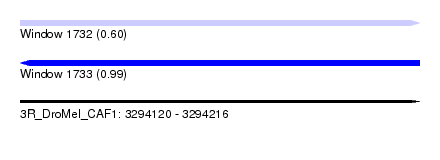

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 3,294,120 – 3,294,216 |
| Length | 96 |
| Max. P | 0.993617 |

| Location | 3,294,120 – 3,294,216 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 93.29 |
| Mean single sequence MFE | -19.73 |
| Consensus MFE | -16.08 |
| Energy contribution | -16.32 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.41 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.599852 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
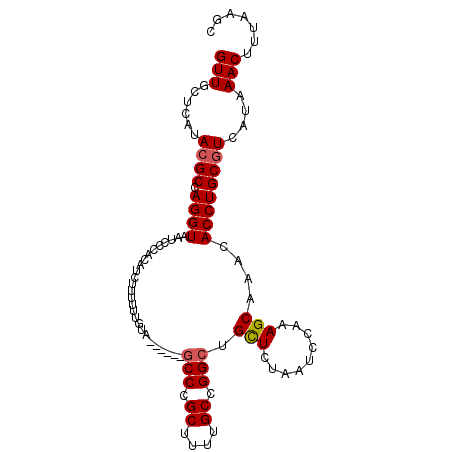
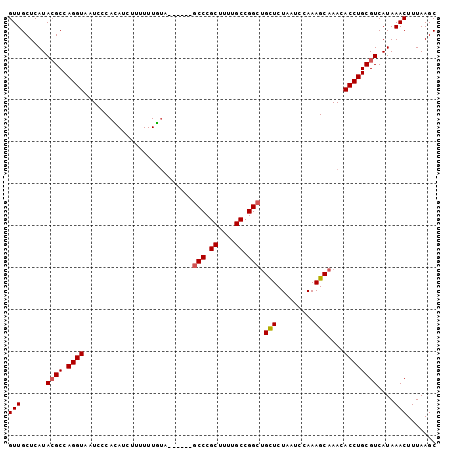
>3R_DroMel_CAF1 3294120 96 + 27905053 GUUGCUCAUACGCCAGGUAAUCCCACAUCUUUUUUGUA------GCCCGCUUAUGCCGGCUGCUCUAAUCCAAAGCAAACACCUGCGUCAUAAACUUUAAGC (((......((((.((((............((...(((------(((.((....)).))))))...))............))))))))....)))....... ( -21.55) >DroSec_CAF1 46949 96 + 1 GUUGCUCAUACGCCAGGUAAUCCCACAUCUUUUUUGUA------GCCCGCUUUUGCCGGCUGCUCUAAUCCAAAGCAAACACCUGCGUCAUAAACUUUAAGC (((......((((.((((............((...(((------(((.((....)).))))))...))............))))))))....)))....... ( -21.55) >DroSim_CAF1 48125 96 + 1 GUUGCUCAUACGCCAGGUAAUCCCACAUCUUUUUUGUA------GCCCGCUUUUGCCGGCUGCUCUAAUCCAAAGCAAACACCUGCGUCAUAAACUUUAAGC (((......((((.((((............((...(((------(((.((....)).))))))...))............))))))))....)))....... ( -21.55) >DroEre_CAF1 48843 94 + 1 GUUGCUCAUACGCCAGGUAAUCCCAUAUCUUUUUUA--------CCCCGCUUUUGCCGGCUGCUCUAAUCCAAAGCAAACACCUGCGUCAUAAACUUUAAGC (((......((((.((((..................--------.((.((....)).)).((((.........))))...))))))))....)))....... ( -15.50) >DroYak_CAF1 66424 101 + 1 GUUGCUCAUAGGCCAGGUAAUC-CAUAUCUUUUUUGUACCUUUUGCCCGCUUUUGCCGGCAGUUCUAAUCCAAAGCAAACACCUGCGUCAUAAACUUUAAGC ..........((((((((....-....................((((.((....)).))))(((.........)))....))))).)))............. ( -18.50) >consensus GUUGCUCAUACGCCAGGUAAUCCCACAUCUUUUUUGUA______GCCCGCUUUUGCCGGCUGCUCUAAUCCAAAGCAAACACCUGCGUCAUAAACUUUAAGC (((......((((.((((..........................(((.((....)).))).(((.........)))....))))))))....)))....... (-16.08 = -16.32 + 0.24)

| Location | 3,294,120 – 3,294,216 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 93.29 |
| Mean single sequence MFE | -28.60 |
| Consensus MFE | -24.94 |
| Energy contribution | -25.74 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.03 |
| Structure conservation index | 0.87 |
| SVM decision value | 2.41 |
| SVM RNA-class probability | 0.993617 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
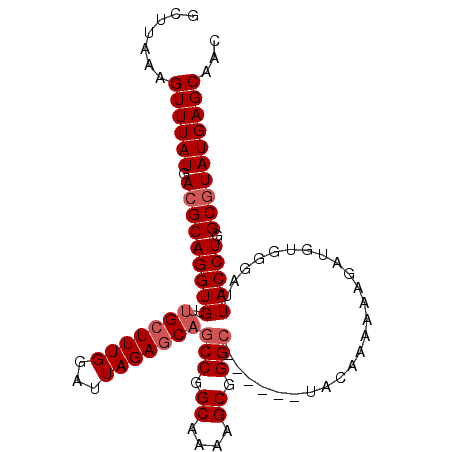
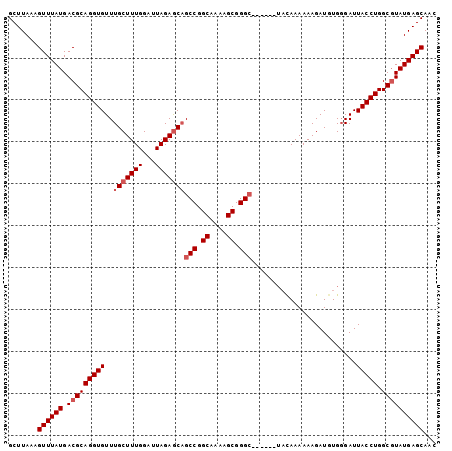
>3R_DroMel_CAF1 3294120 96 - 27905053 GCUUAAAGUUUAUGACGCAGGUGUUUGCUUUGGAUUAGAGCAGCCGGCAUAAGCGGGC------UACAAAAAAGAUGUGGGAUUACCUGGCGUAUGAGCAAC .......(((((((.((((((((...((((((...))))))((((.((....)).)))------)..................))))).))))))))))... ( -30.40) >DroSec_CAF1 46949 96 - 1 GCUUAAAGUUUAUGACGCAGGUGUUUGCUUUGGAUUAGAGCAGCCGGCAAAAGCGGGC------UACAAAAAAGAUGUGGGAUUACCUGGCGUAUGAGCAAC .......(((((((.((((((((...((((((...))))))((((.((....)).)))------)..................))))).))))))))))... ( -30.40) >DroSim_CAF1 48125 96 - 1 GCUUAAAGUUUAUGACGCAGGUGUUUGCUUUGGAUUAGAGCAGCCGGCAAAAGCGGGC------UACAAAAAAGAUGUGGGAUUACCUGGCGUAUGAGCAAC .......(((((((.((((((((...((((((...))))))((((.((....)).)))------)..................))))).))))))))))... ( -30.40) >DroEre_CAF1 48843 94 - 1 GCUUAAAGUUUAUGACGCAGGUGUUUGCUUUGGAUUAGAGCAGCCGGCAAAAGCGGGG--------UAAAAAAGAUAUGGGAUUACCUGGCGUAUGAGCAAC .......(((((((.((((((((.((((((((...))))))))((.((....)).)).--------.................))))).))))))))))... ( -27.10) >DroYak_CAF1 66424 101 - 1 GCUUAAAGUUUAUGACGCAGGUGUUUGCUUUGGAUUAGAACUGCCGGCAAAAGCGGGCAAAAGGUACAAAAAAGAUAUG-GAUUACCUGGCCUAUGAGCAAC (((((..((((.((..((((...((((........)))).))))...)).))))((((...(((((.............-...))))).)))).)))))... ( -24.69) >consensus GCUUAAAGUUUAUGACGCAGGUGUUUGCUUUGGAUUAGAGCAGCCGGCAAAAGCGGGC______UACAAAAAAGAUGUGGGAUUACCUGGCGUAUGAGCAAC .......((((((.(((((((((..(((((((...)))))))(((.((....)).))).........................))))).))))))))))... (-24.94 = -25.74 + 0.80)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:59:49 2006