
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 3,247,923 – 3,248,043 |
| Length | 120 |
| Max. P | 0.984452 |

| Location | 3,247,923 – 3,248,043 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 79.33 |
| Mean single sequence MFE | -35.50 |
| Consensus MFE | -22.64 |
| Energy contribution | -22.58 |
| Covariance contribution | -0.05 |
| Combinations/Pair | 1.22 |
| Mean z-score | -3.48 |
| Structure conservation index | 0.64 |
| SVM decision value | 1.97 |
| SVM RNA-class probability | 0.984452 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
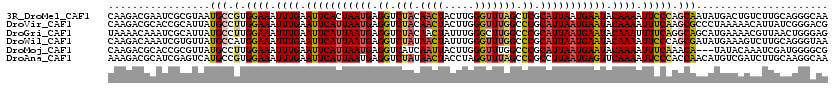
>3R_DroMel_CAF1 3247923 120 + 27905053 CAAGACGAAUCGCGUAAUGCCGUGGAAAUUUGAAUUCACUAAUGAGGUCUACAACUACUUGGGUUUAGCUCGCAUUAAUGAAUACAAAAUUCCCAGCAAUAUGACUGUCUUGCAGGGCAA (((((((.....((((.(((.(.((((.((((.(((((.(((((.(..(((.((((.....)))))))..).))))).))))).)))).))))).))).))))..)))))))........ ( -34.40) >DroVir_CAF1 686 120 + 1 CAAGACGCACCGCAUUAUGCCUUGGAAAUUUGAAUUCAUUAAUGAGGUCUACAACUACUUGGGUUUGGCCCGCAUUAAUGAAUACAAAAUUUCAAGCGCCCUAAAAACAUUAUCGGGACG .....((..(((......(((((((((.((((.(((((((((((.((.(((.((((.....))))))).)).))))))))))).)))).))))))).))..(((.....))).)))..)) ( -33.30) >DroGri_CAF1 692 120 + 1 UAAAACAAAUCGCAUUAUGCCUUGGAAAUUUGAAUUCAUUAAUGAGGUCUACAACUAUUUGGGCUUGGCCCGCAUUAAUGAAUACAAAUUUUCAGGCAGCAUGAAAACGUUAACUGGGAG .........((.((((((((((((((((((((.(((((((((((((........))....((((...)))).))))))))))).))))))))))))..))))))..........)).)). ( -36.40) >DroWil_CAF1 109751 120 + 1 CAAGACAAAUCGUGUUAUGCCAUGGAAAUUUGAAUUCAUUAAUGAGGUCUAUAACUAUUUGGGUUUGGCCCGCAUUAAUGAAUACAAAAUUCCCAGCGAUAUGAAAGUCUUGCAGGGUAA ((((((...(((((((..((...((((.((((.(((((((((((.((((...((((.....)))).))))..))))))))))).)))).))))..)))))))))..))))))........ ( -39.30) >DroMoj_CAF1 581 117 + 1 CAAGACGCACCGCGUUAUGCCUUGGAAAUUUGAAUUCAUUAAUGAGGUCAUCAAUUACUUGGGUUUGGCCCGCAUUAAUGAAUACAAAAUUUCAAACA---UAUACAAAUCGAUGGGGCG .....(((.(((((.((((..((((((.((((.(((((((((((.(((((((((....))))...)))))..))))))))))).)))).)))))).))---)).......)).))).))) ( -33.60) >DroAna_CAF1 10155 120 + 1 AAAGACGCAUCGAGUCAUGCCGUGGAAAUUUGAAUUCAUUAAUGAGGUCUAUAACUACCUAGGUUUAGCCCGCCUUAAUGAGUUCAAAAUUCCCACCAACAUGUCGAUCUUGCAAGGCAA ......(((((((..((((..((((((.((((((((((((((.(.((.(((.((((.....))))))).)).).)))))))))))))).)).))))...))))))))...)))....... ( -36.00) >consensus CAAGACGAAUCGCGUUAUGCCUUGGAAAUUUGAAUUCAUUAAUGAGGUCUACAACUACUUGGGUUUGGCCCGCAUUAAUGAAUACAAAAUUCCAAGCAACAUGAAAAUCUUGCAGGGCAA .................(((.(.((((.((((.(((((((((((.((.(((.((((.....))))))).)).))))))))))).)))).))))).)))...................... (-22.64 = -22.58 + -0.05)
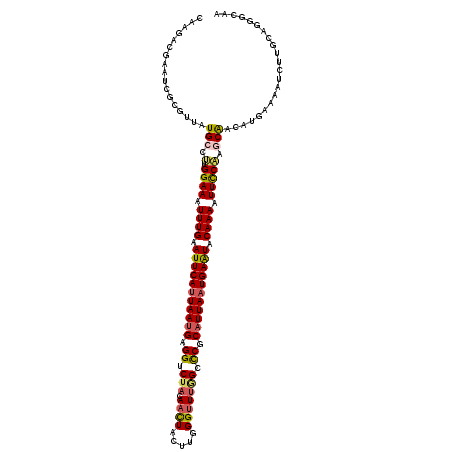

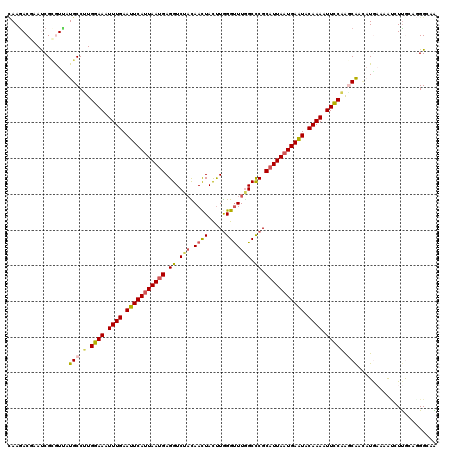
| Location | 3,247,923 – 3,248,043 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 79.33 |
| Mean single sequence MFE | -33.47 |
| Consensus MFE | -20.08 |
| Energy contribution | -19.58 |
| Covariance contribution | -0.50 |
| Combinations/Pair | 1.25 |
| Mean z-score | -2.81 |
| Structure conservation index | 0.60 |
| SVM decision value | 1.60 |
| SVM RNA-class probability | 0.966935 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3247923 120 - 27905053 UUGCCCUGCAAGACAGUCAUAUUGCUGGGAAUUUUGUAUUCAUUAAUGCGAGCUAAACCCAAGUAGUUGUAGACCUCAUUAGUGAAUUCAAAUUUCCACGGCAUUACGCGAUUCGUCUUG ........((((((((((.((.(((((((((.((((.(((((((((((.(..((((((.......))).)))..).))))))))))).)))).)))).))))).))...)))).)))))) ( -38.80) >DroVir_CAF1 686 120 - 1 CGUCCCGAUAAUGUUUUUAGGGCGCUUGAAAUUUUGUAUUCAUUAAUGCGGGCCAAACCCAAGUAGUUGUAGACCUCAUUAAUGAAUUCAAAUUUCCAAGGCAUAAUGCGGUGCGUCUUG (((((..(........)..)))))((((.((.((((.(((((((((((.((....(((.......))).....)).))))))))))).)))).)).))))((((......))))...... ( -32.30) >DroGri_CAF1 692 120 - 1 CUCCCAGUUAACGUUUUCAUGCUGCCUGAAAAUUUGUAUUCAUUAAUGCGGGCCAAGCCCAAAUAGUUGUAGACCUCAUUAAUGAAUUCAAAUUUCCAAGGCAUAAUGCGAUUUGUUUUA .........((((..(((((..(((((..(((((((.(((((((((((.((((...)))).....(((...)))..))))))))))).)))))))...)))))..))).))..))))... ( -31.90) >DroWil_CAF1 109751 120 - 1 UUACCCUGCAAGACUUUCAUAUCGCUGGGAAUUUUGUAUUCAUUAAUGCGGGCCAAACCCAAAUAGUUAUAGACCUCAUUAAUGAAUUCAAAUUUCCAUGGCAUAACACGAUUUGUCUUG ........((((((..((.(((.((((((((.((((.(((((((((((.((....(((.......))).....)).))))))))))).)))).)))).)))))))....))...)))))) ( -32.00) >DroMoj_CAF1 581 117 - 1 CGCCCCAUCGAUUUGUAUA---UGUUUGAAAUUUUGUAUUCAUUAAUGCGGGCCAAACCCAAGUAAUUGAUGACCUCAUUAAUGAAUUCAAAUUUCCAAGGCAUAACGCGGUGCGUCUUG .....(((((....((.((---((((((((((((.(.(((((((((((.(((.((....(((....))).)).)))))))))))))).))))))))..))))))))).)))))....... ( -29.00) >DroAna_CAF1 10155 120 - 1 UUGCCUUGCAAGAUCGACAUGUUGGUGGGAAUUUUGAACUCAUUAAGGCGGGCUAAACCUAGGUAGUUAUAGACCUCAUUAAUGAAUUCAAAUUUCCACGGCAUGACUCGAUGCGUCUUU .......(((...(((((((((((...((((.((((((.(((((((...(((((((((.......))).))).)))..))))))).)))))).)))).)))))))..)))))))...... ( -36.80) >consensus CUCCCCUGCAAGAUUUUCAUGUUGCUGGAAAUUUUGUAUUCAUUAAUGCGGGCCAAACCCAAGUAGUUGUAGACCUCAUUAAUGAAUUCAAAUUUCCAAGGCAUAACGCGAUGCGUCUUG .......................((((((((.((((.(((((((((((.((....(((.......))).....)).))))))))))).))))))))..)))).................. (-20.08 = -19.58 + -0.50)
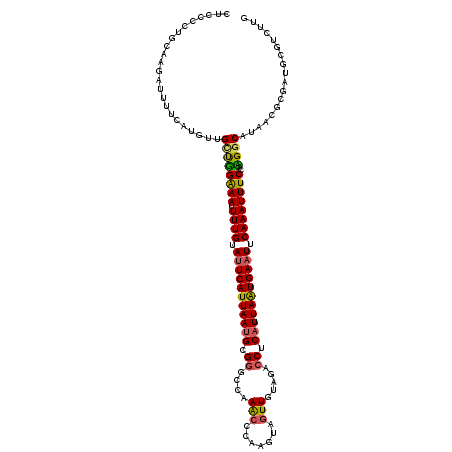
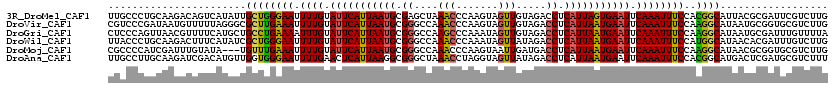


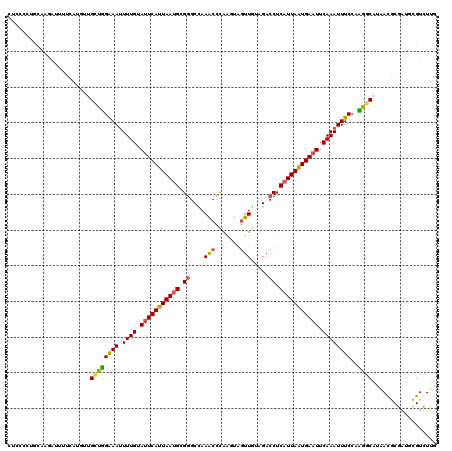
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:59:14 2006