
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 27,880,388 – 27,880,478 |
| Length | 90 |
| Max. P | 0.987174 |

| Location | 27,880,388 – 27,880,478 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 91 |
| Reading direction | forward |
| Mean pairwise identity | 72.42 |
| Mean single sequence MFE | -19.38 |
| Consensus MFE | -14.52 |
| Energy contribution | -12.84 |
| Covariance contribution | -1.68 |
| Combinations/Pair | 1.47 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.75 |
| SVM decision value | 2.07 |
| SVM RNA-class probability | 0.987174 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 27880388 90 + 27905053 AAAAGGACACUAAGCCAAACUUUGGUAAAAAACCAUUUAACAAGCGCCCGGCACAAGAGAAUGGUGGUAAAUC-GUUUGUUAAAAGGGCAA .............(((......((((.....))))(((((((((((.(((.((........)).))).....)-))))))))))..))).. ( -22.10) >DroPse_CAF1 285120 76 + 1 ACA---------------AAUUCGGCAAGAAGCCCUUCAACAAACGGCCAGCCCAGGACAACGGUGCAAAGCCUGCUUGGAAGAAGGUCAA ...---------------.....(((.....)))...........((((..((((((.((..(((.....))))))))))..)..)))).. ( -17.30) >DroSec_CAF1 190289 90 + 1 AAAAAGAUACUAAGCCUAACUUUGGCAAAAAACCAUUUAACAAGCGCCCGGCACAAGAGAAUGGUGGUAAAUC-GUUUGUGAAAAGGGCAA .............(((.......)))...................((((..(((((....(((((.....)))-)))))))....)))).. ( -20.10) >DroSim_CAF1 192893 90 + 1 AAAAAGAUACUAAGCCUAACUUUGGCAAAAAACCAUUUAACAAGCGCCCGGCACAAGAGAAUGGUGGUAAAUC-GUUUGUGAAAAGGGCAA .............(((.......)))...................((((..(((((....(((((.....)))-)))))))....)))).. ( -20.10) >DroPer_CAF1 276529 76 + 1 ACA---------------AAUUCGGCAAGAAGCCCUUCAACAAACGGCCAGCCCAGGACAACGGUGCAAAGCCUGCUUGGAAGAAGGUCAA ...---------------.....(((.....)))...........((((..((((((.((..(((.....))))))))))..)..)))).. ( -17.30) >consensus AAAA_GA_ACUAAGCC_AACUUUGGCAAAAAACCAUUUAACAAGCGCCCGGCACAAGAGAAUGGUGGUAAAUC_GUUUGUAAAAAGGGCAA .......................(((.....)))...........((((...(((((.....(((.....)))..))))).....)))).. (-14.52 = -12.84 + -1.68)


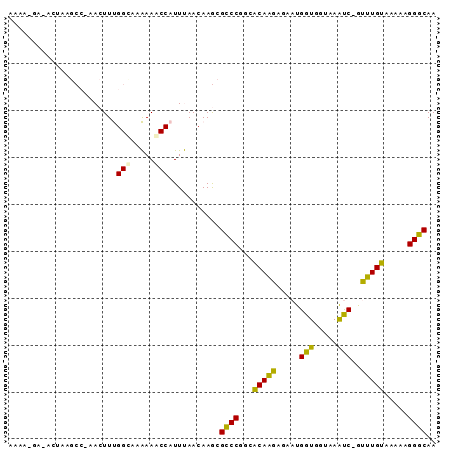
| Location | 27,880,388 – 27,880,478 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 91 |
| Reading direction | reverse |
| Mean pairwise identity | 72.42 |
| Mean single sequence MFE | -20.90 |
| Consensus MFE | -11.84 |
| Energy contribution | -11.08 |
| Covariance contribution | -0.76 |
| Combinations/Pair | 1.40 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.57 |
| SVM decision value | 0.22 |
| SVM RNA-class probability | 0.639897 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 27880388 90 - 27905053 UUGCCCUUUUAACAAAC-GAUUUACCACCAUUCUCUUGUGCCGGGCGCUUGUUAAAUGGUUUUUUACCAAAGUUUGGCUUAGUGUCCUUUU ...........((((..-((...........))..))))...(((((((.((((((((((.....))))...))))))..))))))).... ( -17.90) >DroPse_CAF1 285120 76 - 1 UUGACCUUCUUCCAAGCAGGCUUUGCACCGUUGUCCUGGGCUGGCCGUUUGUUGAAGGGCUUCUUGCCGAAUU---------------UGU .....((((...(((((.((((..((.(((......))))).)))))))))..))))(((.....))).....---------------... ( -24.20) >DroSec_CAF1 190289 90 - 1 UUGCCCUUUUCACAAAC-GAUUUACCACCAUUCUCUUGUGCCGGGCGCUUGUUAAAUGGUUUUUUGCCAAAGUUAGGCUUAGUAUCUUUUU ..((((....(((((..-((...........))..)))))..))))(((((.....((((.....))))....)))))............. ( -19.10) >DroSim_CAF1 192893 90 - 1 UUGCCCUUUUCACAAAC-GAUUUACCACCAUUCUCUUGUGCCGGGCGCUUGUUAAAUGGUUUUUUGCCAAAGUUAGGCUUAGUAUCUUUUU ..((((....(((((..-((...........))..)))))..))))(((((.....((((.....))))....)))))............. ( -19.10) >DroPer_CAF1 276529 76 - 1 UUGACCUUCUUCCAAGCAGGCUUUGCACCGUUGUCCUGGGCUGGCCGUUUGUUGAAGGGCUUCUUGCCGAAUU---------------UGU .....((((...(((((.((((..((.(((......))))).)))))))))..))))(((.....))).....---------------... ( -24.20) >consensus UUGCCCUUUUCACAAAC_GAUUUACCACCAUUCUCUUGUGCCGGGCGCUUGUUAAAUGGUUUUUUGCCAAAGUU_GGCUUAGU_UC_UUUU ..((((....(((((...(((........)))...)))))..))))...........(((.....)))....................... (-11.84 = -11.08 + -0.76)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:54:42 2006