
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 27,753,461 – 27,753,556 |
| Length | 95 |
| Max. P | 0.989419 |

| Location | 27,753,461 – 27,753,556 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 86.42 |
| Mean single sequence MFE | -32.02 |
| Consensus MFE | -29.02 |
| Energy contribution | -29.53 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.46 |
| Structure conservation index | 0.91 |
| SVM decision value | 2.16 |
| SVM RNA-class probability | 0.989419 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 27753461 95 + 27905053 GCUCACAUGCUCACAUGCUCCUGCCAGACUCUUUAGUUUGGCAGUUUGCUAGUUUCCUGGUGUGCUAGUGAG--------CUCGUGUGCUCGGUGGAUUAGUA ((.(((..((((((..((..((((((((((....))))))))))...(((((....)))))..))..)))))--------)..))).)).............. ( -37.20) >DroSec_CAF1 67641 86 + 1 GCUCA--------UAUGCUCCUGCCAGACUCC-UAGUUUGGCAGUUUGCUAGUUUCCUGGUGUGCUAGUGAG--------CUCGUGUGCUCGGUGGAUUAGUA ((...--------...))..(((((((((...-..)))))))))..((((((((..((((((..(.((....--------)).)..))).)))..)))))))) ( -32.40) >DroSim_CAF1 67884 86 + 1 GCUCA--------UAUGCUCCUGCCACACUCC-UAGUUUGGCAGUUUGCUAGUUUCCUGGUGUGCUAGUGAG--------CUCGUGUGCUCGGUGGAUUAGUA ((...--------...))..((((((.((...-..)).))))))..((((((((..((((((..(.((....--------)).)..))).)))..)))))))) ( -28.80) >DroEre_CAF1 72589 94 + 1 GCUCA--------UAUUCUCCUGCCAGUCUCG-UAGUUUGGCAGUUUGCUAGUUUCCUGGUGAGCUAGUGAGCUCGUGAGCUCGUGUGCUUGGUGGAUUAGUG .....--------.......(((((((.(...-..).)))))))...(((((((..((...((((...((((((....))))))...))))))..))))))). ( -29.70) >consensus GCUCA________UAUGCUCCUGCCAGACUCC_UAGUUUGGCAGUUUGCUAGUUUCCUGGUGUGCUAGUGAG________CUCGUGUGCUCGGUGGAUUAGUA ....................((((((((((....))))))))))..((((((((..((((.(..(...((((((....)))))).)..)))))..)))))))) (-29.02 = -29.53 + 0.50)
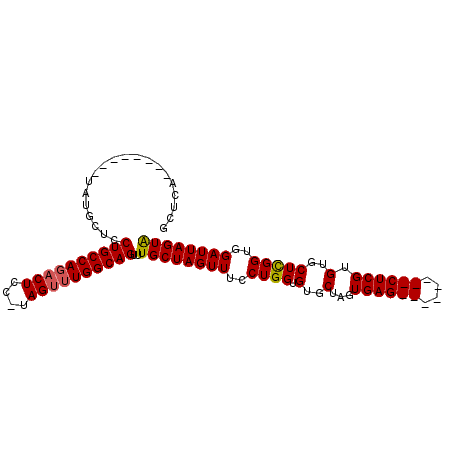



| Location | 27,753,461 – 27,753,556 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 86.42 |
| Mean single sequence MFE | -26.28 |
| Consensus MFE | -18.90 |
| Energy contribution | -19.52 |
| Covariance contribution | 0.63 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.87 |
| Structure conservation index | 0.72 |
| SVM decision value | 1.56 |
| SVM RNA-class probability | 0.963968 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 27753461 95 - 27905053 UACUAAUCCACCGAGCACACGAG--------CUCACUAGCACACCAGGAAACUAGCAAACUGCCAAACUAAAGAGUCUGGCAGGAGCAUGUGAGCAUGUGAGC ..............((.((((.(--------(((((..((.....((....))......((((((.(((....))).))))))..))..)))))).)))).)) ( -34.10) >DroSec_CAF1 67641 86 - 1 UACUAAUCCACCGAGCACACGAG--------CUCACUAGCACACCAGGAAACUAGCAAACUGCCAAACUA-GGAGUCUGGCAGGAGCAUA--------UGAGC ............((((......)--------)))....((.((..((....)).((...((((((.((..-...)).))))))..))...--------)).)) ( -23.10) >DroSim_CAF1 67884 86 - 1 UACUAAUCCACCGAGCACACGAG--------CUCACUAGCACACCAGGAAACUAGCAAACUGCCAAACUA-GGAGUGUGGCAGGAGCAUA--------UGAGC ............((((......)--------)))....((.((..((....)).((...((((((.((..-...)).))))))..))...--------)).)) ( -23.10) >DroEre_CAF1 72589 94 - 1 CACUAAUCCACCAAGCACACGAGCUCACGAGCUCACUAGCUCACCAGGAAACUAGCAAACUGCCAAACUA-CGAGACUGGCAGGAGAAUA--------UGAGC ....................(((((....)))))....(((((..((....))......((((((..(..-...)..)))))).......--------))))) ( -24.80) >consensus UACUAAUCCACCGAGCACACGAG________CUCACUAGCACACCAGGAAACUAGCAAACUGCCAAACUA_GGAGUCUGGCAGGAGCAUA________UGAGC ......(((.....((....(((((....)))))...........((....)).))....(((((.(((....))).)))))))).................. (-18.90 = -19.52 + 0.63)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:54:11 2006