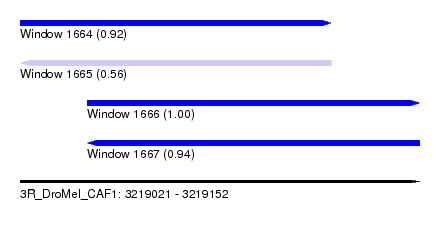

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 3,219,021 – 3,219,152 |
| Length | 131 |
| Max. P | 0.997916 |

| Location | 3,219,021 – 3,219,123 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 96.67 |
| Mean single sequence MFE | -23.36 |
| Consensus MFE | -21.68 |
| Energy contribution | -22.12 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.45 |
| Structure conservation index | 0.93 |
| SVM decision value | 1.13 |
| SVM RNA-class probability | 0.919765 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3219021 102 + 27905053 CAGCGAAAAUAUAUCAAUUGCAUGGCGCAAAGUAUUUCGCCAUCGAAACUUCAGUUCGGCAACUUUCUCAUCGAGCACUUAGAUGGGACUUGCCAAACUUUU .................(((.((((((.((....)).)))))))))......((((.(((((...(((((((.........))))))).))))).))))... ( -25.70) >DroSec_CAF1 43754 102 + 1 CAUCGAAAAUAUAUCAAUUGCAUGGCGCAAAGUAUUUCGCCAUUGAAACUUCAGUUCGGCAACUUUCUCAUCGAGCACUUAGAUGGGACUUGCCAAACUUUU .............(((.....((((((.((....)).)))))))))......((((.(((((...(((((((.........))))))).))))).))))... ( -23.90) >DroSim_CAF1 45103 102 + 1 CAUCGAAAAUAUAUCAAUUGCAUGGCGCAAAGUAUUUCGCCAUCGAAACUUCAGUUCGGCAACUUUCUCAUCGAGCACUUAGAUGGGACUUGCCAAACUUUU .................(((.((((((.((....)).)))))))))......((((.(((((...(((((((.........))))))).))))).))))... ( -25.70) >DroEre_CAF1 38069 102 + 1 CUGCGAAAAUAUAUCAAUUGCAUGGCGCAAAGUAUUUCGCCAUCGAAACUUUAGUUCGGCAACUUUCACAUCGAGCACUUAGAUGGGACUUGGCAAACUUUU .(((.((....((((..(((.((((((.((....)).))))))))).......((((((...........)))))).....))))....)).)))....... ( -20.50) >DroYak_CAF1 28394 102 + 1 CUGCGAAAAUAUAUCAAUUGCAUGGCGCAAAGUAUUUCGCCAUCGAAACUUUAGUUCGGCAACUUUCUCAUCGAGCACUUAAAUGGGACUUGCCAAACUUUU .................(((.((((((.((....)).)))))))))......((((.(((((...((((((...........)))))).))))).))))... ( -21.00) >consensus CAGCGAAAAUAUAUCAAUUGCAUGGCGCAAAGUAUUUCGCCAUCGAAACUUCAGUUCGGCAACUUUCUCAUCGAGCACUUAGAUGGGACUUGCCAAACUUUU .................(((.((((((.((....)).)))))))))......((((.(((((...(((((((.........))))))).))))).))))... (-21.68 = -22.12 + 0.44)

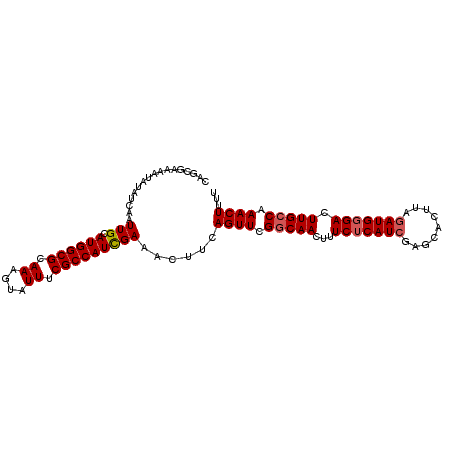
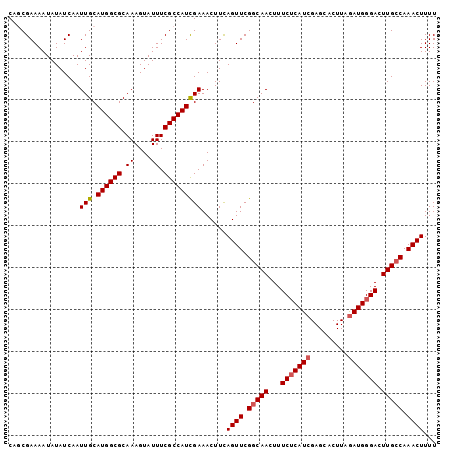
| Location | 3,219,021 – 3,219,123 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 96.67 |
| Mean single sequence MFE | -25.57 |
| Consensus MFE | -24.10 |
| Energy contribution | -24.14 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.11 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.562249 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3219021 102 - 27905053 AAAAGUUUGGCAAGUCCCAUCUAAGUGCUCGAUGAGAAAGUUGCCGAACUGAAGUUUCGAUGGCGAAAUACUUUGCGCCAUGCAAUUGAUAUAUUUUCGCUG ...((((((((((.((.((((.........)))).))...))))))))))...........(((((((....((((.....)))).........))))))). ( -26.92) >DroSec_CAF1 43754 102 - 1 AAAAGUUUGGCAAGUCCCAUCUAAGUGCUCGAUGAGAAAGUUGCCGAACUGAAGUUUCAAUGGCGAAAUACUUUGCGCCAUGCAAUUGAUAUAUUUUCGAUG ...((((((((((.((.((((.........)))).))...)))))))))).........((((((.((....)).))))))...(((((.......))))). ( -27.80) >DroSim_CAF1 45103 102 - 1 AAAAGUUUGGCAAGUCCCAUCUAAGUGCUCGAUGAGAAAGUUGCCGAACUGAAGUUUCGAUGGCGAAAUACUUUGCGCCAUGCAAUUGAUAUAUUUUCGAUG ...((((((((((.((.((((.........)))).))...)))))))))).........((((((.((....)).))))))...(((((.......))))). ( -27.90) >DroEre_CAF1 38069 102 - 1 AAAAGUUUGCCAAGUCCCAUCUAAGUGCUCGAUGUGAAAGUUGCCGAACUAAAGUUUCGAUGGCGAAAUACUUUGCGCCAUGCAAUUGAUAUAUUUUCGCAG .........................(((..((((((..(((((((((((....).))))((((((.((....)).))))))))))))..))))))...))). ( -21.50) >DroYak_CAF1 28394 102 - 1 AAAAGUUUGGCAAGUCCCAUUUAAGUGCUCGAUGAGAAAGUUGCCGAACUAAAGUUUCGAUGGCGAAAUACUUUGCGCCAUGCAAUUGAUAUAUUUUCGCAG ...((((((((((.((.((((.........)))).))...))))))))))............((((((....((((.....)))).........)))))).. ( -23.72) >consensus AAAAGUUUGGCAAGUCCCAUCUAAGUGCUCGAUGAGAAAGUUGCCGAACUGAAGUUUCGAUGGCGAAAUACUUUGCGCCAUGCAAUUGAUAUAUUUUCGCUG ...((((((((((.((.((((.........)))).))...))))))))))..((((.(.((((((.((....)).))))))).))))............... (-24.10 = -24.14 + 0.04)

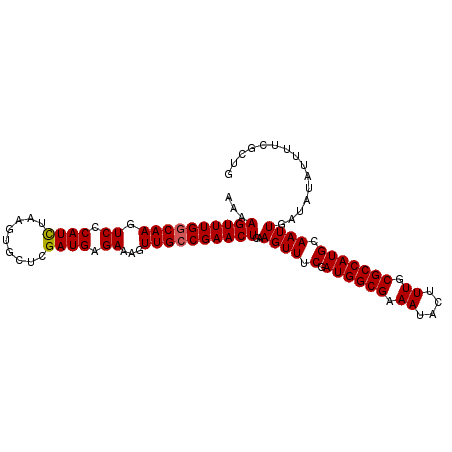
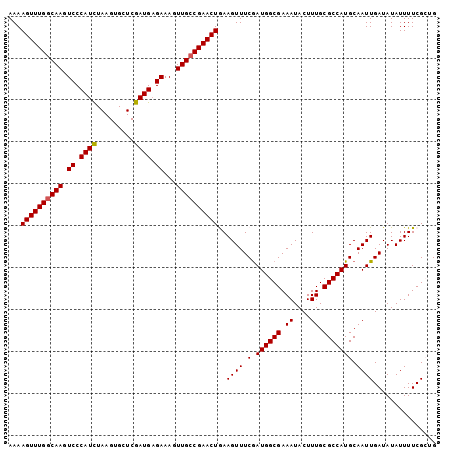
| Location | 3,219,043 – 3,219,152 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 97.61 |
| Mean single sequence MFE | -28.26 |
| Consensus MFE | -27.28 |
| Energy contribution | -27.72 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.97 |
| SVM decision value | 2.96 |
| SVM RNA-class probability | 0.997916 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3219043 109 + 27905053 UGGCGCAAAGUAUUUCGCCAUCGAAACUUCAGUUCGGCAACUUUCUCAUCGAGCACUUAGAUGGGACUUGCCAAACUUUUCGAUUUUGCGGCAACUUAAAAAUUUGAAU ((.((((((((.(((((....)))))....((((.(((((...(((((((.........))))))).))))).)))).....)))))))).))................ ( -31.00) >DroSec_CAF1 43776 109 + 1 UGGCGCAAAGUAUUUCGCCAUUGAAACUUCAGUUCGGCAACUUUCUCAUCGAGCACUUAGAUGGGACUUGCCAAACUUUUCGAUUUUGCGGCAACUUAAAAAUUUGAAU ((.((((((((.(((((....)))))....((((.(((((...(((((((.........))))))).))))).)))).....)))))))).))................ ( -29.70) >DroSim_CAF1 45125 109 + 1 UGGCGCAAAGUAUUUCGCCAUCGAAACUUCAGUUCGGCAACUUUCUCAUCGAGCACUUAGAUGGGACUUGCCAAACUUUUCGAUUUUGCGGCAACUUAAAAAUUUGAAU ((.((((((((.(((((....)))))....((((.(((((...(((((((.........))))))).))))).)))).....)))))))).))................ ( -31.00) >DroEre_CAF1 38091 109 + 1 UGGCGCAAAGUAUUUCGCCAUCGAAACUUUAGUUCGGCAACUUUCACAUCGAGCACUUAGAUGGGACUUGGCAAACUUUUCGAUUUUGCGGCAACUUCAAAAUUUGAAU ((.((((((((.(((((....)))))....((((.(.(((..(((.((((.........))))))).))).).)))).....)))))))).))..((((.....)))). ( -23.30) >DroYak_CAF1 28416 109 + 1 UGGCGCAAAGUAUUUCGCCAUCGAAACUUUAGUUCGGCAACUUUCUCAUCGAGCACUUAAAUGGGACUUGCCAAACUUUUCGAUUUUGCGGCAACUUAAAAAUUUGAAU ((.((((((((.(((((....)))))....((((.(((((...((((((...........)))))).))))).)))).....)))))))).))................ ( -26.30) >consensus UGGCGCAAAGUAUUUCGCCAUCGAAACUUCAGUUCGGCAACUUUCUCAUCGAGCACUUAGAUGGGACUUGCCAAACUUUUCGAUUUUGCGGCAACUUAAAAAUUUGAAU ((.((((((((.(((((....)))))....((((.(((((...(((((((.........))))))).))))).)))).....)))))))).))................ (-27.28 = -27.72 + 0.44)

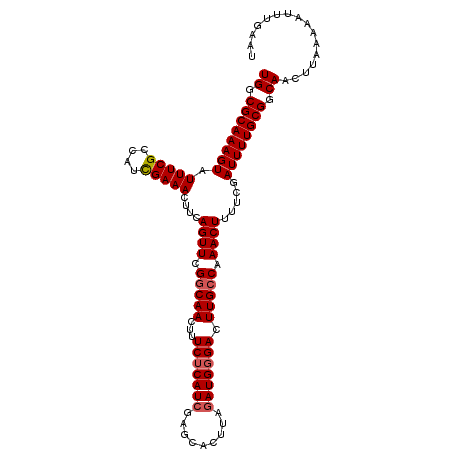
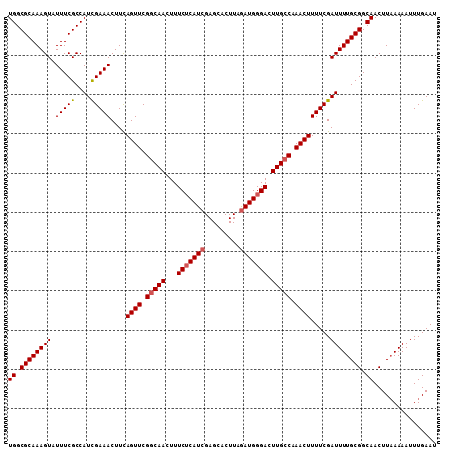
| Location | 3,219,043 – 3,219,152 |
|---|---|
| Length | 109 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 97.61 |
| Mean single sequence MFE | -27.58 |
| Consensus MFE | -26.68 |
| Energy contribution | -26.92 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.97 |
| SVM decision value | 1.32 |
| SVM RNA-class probability | 0.942198 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 3219043 109 - 27905053 AUUCAAAUUUUUAAGUUGCCGCAAAAUCGAAAAGUUUGGCAAGUCCCAUCUAAGUGCUCGAUGAGAAAGUUGCCGAACUGAAGUUUCGAUGGCGAAAUACUUUGCGCCA ............((((...(((...(((((((((((((((((.((.((((.........)))).))...))))))))))....))))))).)))....))))....... ( -29.70) >DroSec_CAF1 43776 109 - 1 AUUCAAAUUUUUAAGUUGCCGCAAAAUCGAAAAGUUUGGCAAGUCCCAUCUAAGUGCUCGAUGAGAAAGUUGCCGAACUGAAGUUUCAAUGGCGAAAUACUUUGCGCCA ................((.((((((.......((((((((((.((.((((.........)))).))...))))))))))...(((((......)))))..)))))).)) ( -26.90) >DroSim_CAF1 45125 109 - 1 AUUCAAAUUUUUAAGUUGCCGCAAAAUCGAAAAGUUUGGCAAGUCCCAUCUAAGUGCUCGAUGAGAAAGUUGCCGAACUGAAGUUUCGAUGGCGAAAUACUUUGCGCCA ............((((...(((...(((((((((((((((((.((.((((.........)))).))...))))))))))....))))))).)))....))))....... ( -29.70) >DroEre_CAF1 38091 109 - 1 AUUCAAAUUUUGAAGUUGCCGCAAAAUCGAAAAGUUUGCCAAGUCCCAUCUAAGUGCUCGAUGUGAAAGUUGCCGAACUAAAGUUUCGAUGGCGAAAUACUUUGCGCCA ...........(((((...(((...(((((((((((((.(((.((.((((.........)))).))...))).))))))....))))))).)))....)))))...... ( -24.30) >DroYak_CAF1 28416 109 - 1 AUUCAAAUUUUUAAGUUGCCGCAAAAUCGAAAAGUUUGGCAAGUCCCAUUUAAGUGCUCGAUGAGAAAGUUGCCGAACUAAAGUUUCGAUGGCGAAAUACUUUGCGCCA ............((((...(((...(((((((((((((((((.((.((((.........)))).))...))))))))))....))))))).)))....))))....... ( -27.30) >consensus AUUCAAAUUUUUAAGUUGCCGCAAAAUCGAAAAGUUUGGCAAGUCCCAUCUAAGUGCUCGAUGAGAAAGUUGCCGAACUGAAGUUUCGAUGGCGAAAUACUUUGCGCCA ............((((...(((...(((((((((((((((((.((.((((.........)))).))...))))))))))....))))))).)))....))))....... (-26.68 = -26.92 + 0.24)
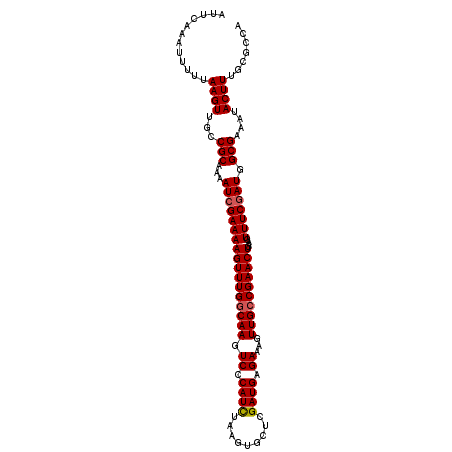


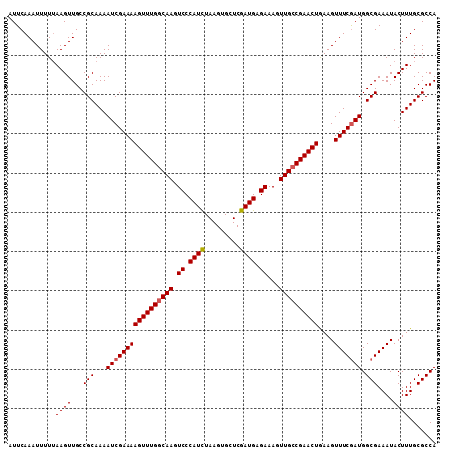
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:58:46 2006