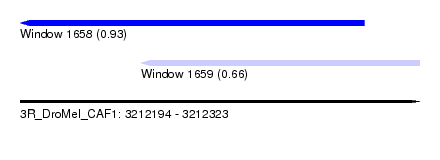
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 3,212,194 – 3,212,323 |
| Length | 129 |
| Max. P | 0.925973 |

| Location | 3,212,194 – 3,212,305 |
|---|---|
| Length | 111 |
| Sequences | 5 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 91.25 |
| Mean single sequence MFE | -31.42 |
| Consensus MFE | -24.20 |
| Energy contribution | -24.24 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.23 |
| Structure conservation index | 0.77 |
| SVM decision value | 1.17 |
| SVM RNA-class probability | 0.925973 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
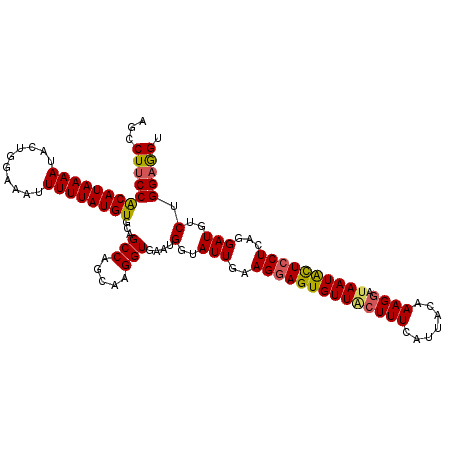
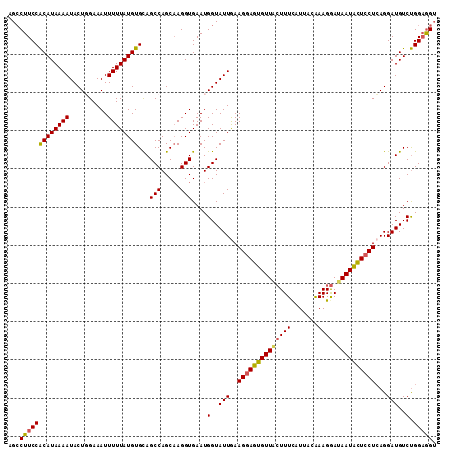
>3R_DroMel_CAF1 3212194 111 - 27905053 AGCCUUCCACAUAAAAUACUGGAAAUUUUUAUGUGUAGCCAGCAAGGUGAAUGGUAUUGAAGGAGUGUUACUUUUAUUACAAAGGAUAAUACUCCUCAGGAUGUCUGGUAG- .(((((((((((((((..........)))))))))..(((.....)))))).))).((((.((((((((((((((.....))))).)))))))))))))............- ( -29.50) >DroSec_CAF1 36873 112 - 1 AGCCUUCCGCAUAAAAUACUGGAAAUUUUUAUGUGCAGCCAGCAAGGUGAAUGGUAUUAAAGGAGUGUUACUUUCAUUACAAAGGAUAAUACUCCUCAGGAUGUCCGGAGGU .(((((((((((((((..........)))))))))..(((.....)))....((((((..((((((((((((((.......)))).))))))))))...)))).)))))))) ( -32.70) >DroSim_CAF1 38338 112 - 1 AGCCUUCCACAUAAAAUACUGGAAAUUUUUAUGUGCAGCCAGCAAGGUGAAUGGUAUUGAAGGAGUGUUACUUUCAUUACAAAGGAUAAUACUCCUCAGGAUGUCUGGAGGU .(((((((((((((((..........)))))))))..((((..........)))).((((.(((((((((((((.......)))).))))))))))))).......)))))) ( -31.00) >DroEre_CAF1 31399 111 - 1 UGCCUUCCACAUAAAAUACUUGAAAUUUUUAUGUGCAGCCAGCAAGGUGAAUGGUAUUGAAGGAGUGUUGGUUUCCUUACAAAGAAAAAUGUU-CUGAGGAUGUCUGGAGGU .(((((((((((((((..........))))))))).((((((((...(.(((...))).).....))))))))(((((....((((.....))-))))))).....)))))) ( -25.20) >DroYak_CAF1 21699 112 - 1 UGGCUUCCACAUAAAAUACUUGAAAUUUUUAUGUGCAGCCAGCCAGGUGAAUGGUAUUGAAGGAGUGUUGCUUUCAUUUCGAAGGAUAAUACUCCUCAGGAUGUCUGGAGGU (((((..(((((((((..........))))))))).))))).((((..(.......((((.((((((((((((((.....))))).)))))))))))))..)..)))).... ( -38.70) >consensus AGCCUUCCACAUAAAAUACUGGAAAUUUUUAUGUGCAGCCAGCAAGGUGAAUGGUAUUGAAGGAGUGUUACUUUCAUUACAAAGGAUAAUACUCCUCAGGAUGUCUGGAGGU ...(((((((((((((..........))))))))...(((.....)))....(..(((..((((((((((((((.......)))).))))))))))...)))..).))))). (-24.20 = -24.24 + 0.04)

| Location | 3,212,233 – 3,212,323 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 90 |
| Reading direction | reverse |
| Mean pairwise identity | 94.89 |
| Mean single sequence MFE | -22.30 |
| Consensus MFE | -17.88 |
| Energy contribution | -17.76 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.655235 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
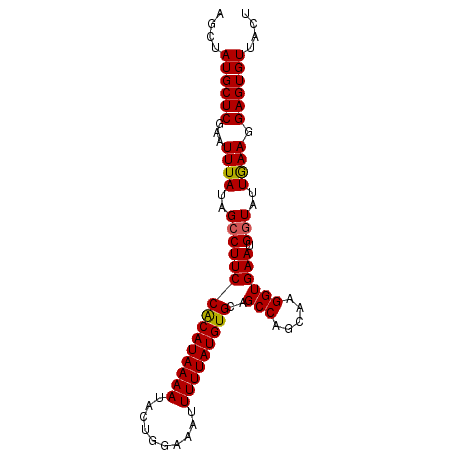
>3R_DroMel_CAF1 3212233 90 - 27905053 AGCUAUGCUCGAAUUUAUAGCCUUCCACAUAAAAUACUGGAAAUUUUUAUGUGUAGCCAGCAAGGUGAAUGGUAUUGAAGGAGUGUUACU ((..((((((...((((..(((((((((((((((..........)))))))))..(((.....)))))).)))..)))).))))))..)) ( -19.70) >DroSec_CAF1 36913 90 - 1 AGCUAUGCUCCAAUUUAUAGCCUUCCGCAUAAAAUACUGGAAAUUUUUAUGUGCAGCCAGCAAGGUGAAUGGUAUUAAAGGAGUGUUACU ((..(((((((..((((..(((((((((((((((..........)))))))))..(((.....)))))).)))..)))))))))))..)) ( -21.30) >DroSim_CAF1 38378 90 - 1 AGCUAUGCUCGAAUUUAUAGCCUUCCACAUAAAAUACUGGAAAUUUUUAUGUGCAGCCAGCAAGGUGAAUGGUAUUGAAGGAGUGUUACU ((..((((((...((((..(((((((((((((((..........)))))))))..(((.....)))))).)))..)))).))))))..)) ( -19.50) >DroEre_CAF1 31438 90 - 1 AGCUAUGCUCGAAUUUAUUGCCUUCCACAUAAAAUACUUGAAAUUUUUAUGUGCAGCCAGCAAGGUGAAUGGUAUUGAAGGAGUGUUGGU .(((((((((...((((.((((((((((((((((..........)))))))))..(((.....)))))).)))).)))).))))).)))) ( -23.00) >DroYak_CAF1 21739 90 - 1 AGCUAUGCUCGAAUUUAUUGGCUUCCACAUAAAAUACUUGAAAUUUUUAUGUGCAGCCAGCCAGGUGAAUGGUAUUGAAGGAGUGUUGCU (((.((((((........(((((..(((((((((..........))))))))).)))))((((......)))).......)))))).))) ( -28.00) >consensus AGCUAUGCUCGAAUUUAUAGCCUUCCACAUAAAAUACUGGAAAUUUUUAUGUGCAGCCAGCAAGGUGAAUGGUAUUGAAGGAGUGUUACU ....((((((...((((..(((((((((((((((..........)))))))))..(((.....)))))).)))..)))).)))))).... (-17.88 = -17.76 + -0.12)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:58:39 2006