
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 27,391,318 – 27,391,410 |
| Length | 92 |
| Max. P | 0.776867 |

| Location | 27,391,318 – 27,391,410 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 78.28 |
| Mean single sequence MFE | -21.10 |
| Consensus MFE | -15.13 |
| Energy contribution | -15.05 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.33 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.776867 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 27391318 92 + 27905053 CGAUACAAU--AGAAUGCGCGCUUUUAAAUGCACUGGA-AGACGAG-----------AUUCCAGCAUUUUAAAUCGGCGCAUUGCCAACAAAAGAACGGAAAUUCA .........--..((((((((.(((.((((((..((((-(......-----------.))))))))))).))).).)))))))....................... ( -20.40) >DroSec_CAF1 7491 92 + 1 CGAUACGAU--AUAAUGCGCGCUUUUAAAUGCACUGGA-CGACGAG-----------AUUCCAGCAUUUUAAAUCGGCGCAUUGCCAAGAAAAGAACGGAAAUUCA .........--.(((((((((.(((.((((((..((((-.......-----------..)))))))))).))).).))))))))...................... ( -18.80) >DroEre_CAF1 8088 91 + 1 UGAU-CAAU--CGAAUGCGCGCUUUUAAAUGCACUGGA-AUGCGCA-----------AUUCCAGCAUUUUAAAUCGGCGCAUUGCCAAGAAAAGAACGGAAAUUUA ....-....--..((((((((.(((.((((((..((((-((.....-----------)))))))))))).))).).)))))))....................... ( -22.10) >DroYak_CAF1 7436 104 + 1 CGAUACAAU--AGAAUGCGCGCUUUUAAAUGCACUGGAAAUGCACAGGAAAAUGGAGAUUCCAGCAUUUUAAAUCGGCGCAUUGCCAAGAAAAGAACGGAAAUUGA .....((((--..((((((((.(((.((((((.(((........))).....(((.....))))))))).))).).))))))).((...........))..)))). ( -23.50) >DroAna_CAF1 8057 97 + 1 CGAUACGAUGAAUAAUGCGCGCUUUUUAAUGCCUUGGA-AAUCCAG--------UGGUUUCCAACAUUUUAAAUCGGCACAUUGCCAAGAAAAAAACGAAAAGUUU .....(((((.....(((.((.(((..((((..(((((-((((...--------.)))))))))))))..))).)))))))))).........((((.....)))) ( -19.10) >DroPer_CAF1 46409 92 + 1 UGUCUCUCA--AGAAUGCGCGCUUUUAAAUGCCCUGGA-AAUUGAA----AA----UGUUCCAGCAUUUUGAAUCUGCGCAUUGCCAAUCA---AACGGAAAUUCU .........--..((((((((..(((((((((..((((-(..(...----.)----..)))))))).))))))..)))))))).((.....---...))....... ( -22.70) >consensus CGAUACAAU__AGAAUGCGCGCUUUUAAAUGCACUGGA_AAACGAA___________AUUCCAGCAUUUUAAAUCGGCGCAUUGCCAAGAAAAGAACGGAAAUUCA .............(((((((...((((((....(((((.....................)))))...))))))...)))))))....................... (-15.13 = -15.05 + -0.08)



| Location | 27,391,318 – 27,391,410 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 78.28 |
| Mean single sequence MFE | -21.50 |
| Consensus MFE | -16.34 |
| Energy contribution | -16.40 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.14 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.724575 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 27391318 92 - 27905053 UGAAUUUCCGUUCUUUUGUUGGCAAUGCGCCGAUUUAAAAUGCUGGAAU-----------CUCGUCU-UCCAGUGCAUUUAAAAGCGCGCAUUCU--AUUGUAUCG .((((....))))......(((.((((((((..((((((..(((((((.-----------......)-))))))...)))))).).)))))))))--)........ ( -22.30) >DroSec_CAF1 7491 92 - 1 UGAAUUUCCGUUCUUUUCUUGGCAAUGCGCCGAUUUAAAAUGCUGGAAU-----------CUCGUCG-UCCAGUGCAUUUAAAAGCGCGCAUUAU--AUCGUAUCG .((((....))))..........((((((((..((((((..((((((..-----------.......-))))))...)))))).).)))))))..--......... ( -19.40) >DroEre_CAF1 8088 91 - 1 UAAAUUUCCGUUCUUUUCUUGGCAAUGCGCCGAUUUAAAAUGCUGGAAU-----------UGCGCAU-UCCAGUGCAUUUAAAAGCGCGCAUUCG--AUUG-AUCA .....................(.((((((((..((((((..((((((((-----------.....))-))))))...)))))).).)))))))).--....-.... ( -21.40) >DroYak_CAF1 7436 104 - 1 UCAAUUUCCGUUCUUUUCUUGGCAAUGCGCCGAUUUAAAAUGCUGGAAUCUCCAUUUUCCUGUGCAUUUCCAGUGCAUUUAAAAGCGCGCAUUCU--AUUGUAUCG ...................(((.((((((((..((((((..(((((((....(((......)))...)))))))...)))))).).)))))))))--)........ ( -23.70) >DroAna_CAF1 8057 97 - 1 AAACUUUUCGUUUUUUUCUUGGCAAUGUGCCGAUUUAAAAUGUUGGAAACCA--------CUGGAUU-UCCAAGGCAUUAAAAAGCGCGCAUUAUUCAUCGUAUCG ........((((((((..(((((.....))))).....(((((((((((((.--------..)).))-))))..)))))))))))))................... ( -19.50) >DroPer_CAF1 46409 92 - 1 AGAAUUUCCGUU---UGAUUGGCAAUGCGCAGAUUCAAAAUGCUGGAACA----UU----UUCAAUU-UCCAGGGCAUUUAAAAGCGCGCAUUCU--UGAGAGACA .........(((---(..(..(.(((((((............((((((..----..----......)-))))).((........))))))))).)--..).)))). ( -22.70) >consensus UGAAUUUCCGUUCUUUUCUUGGCAAUGCGCCGAUUUAAAAUGCUGGAAU___________CUCGAAU_UCCAGUGCAUUUAAAAGCGCGCAUUCU__AUUGUAUCG .......................(((((((...(((.((((((((((.....................))))..)))))).)))..)))))))............. (-16.34 = -16.40 + 0.06)

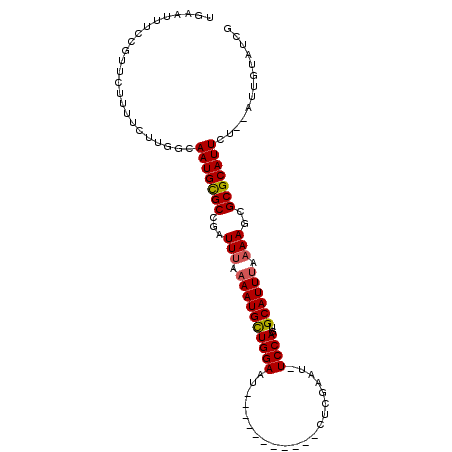

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:51:26 2006