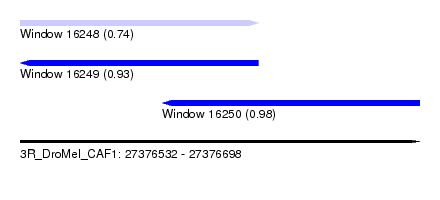
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 27,376,532 – 27,376,698 |
| Length | 166 |
| Max. P | 0.975066 |

| Location | 27,376,532 – 27,376,631 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 87.43 |
| Mean single sequence MFE | -33.50 |
| Consensus MFE | -24.48 |
| Energy contribution | -26.28 |
| Covariance contribution | 1.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.46 |
| SVM RNA-class probability | 0.743437 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 27376532 99 + 27905053 CUGCAGACAGUCUGCGGCGGCAGCGUUUGUCU----GCCAGAUUCUUCAGCUCUCUGCAGCUC--------UGCAGCUUGGCUUUUUGCUGCCUUCCUGUCUUGGCCAAUU ..((((((((...(((((((((((....).))----))).........((((..(((((....--------)))))...))))....)))))....))))))..))..... ( -34.30) >DroSec_CAF1 121912 107 + 1 CUGCAGACAGUCUGCGGCGGCAGCGUUUGUCU----GCCAGAUUCUUCAGCUCUCUGCAGCUCAGCAGCUUUGCAGCUUGGCUUUUUGCUGCCUUCCUGUCUUGGCCAAUU ..((((((((...(((((((((((....).))----))).........((((..((((((((....)))..)))))...))))....)))))....))))))..))..... ( -35.40) >DroSim_CAF1 86134 107 + 1 CUGCAGACAGUCUGCGGCGGCAGCGUUUGUCU----GCCAGAUUCUUCAGCUCUCUGCAGCUCAGCAGCUUUGCAGCUUGGCUUUUUGCUGCCUUCCUGUCUUGGCCAAUU ..((((((((...(((((((((((....).))----))).........((((..((((((((....)))..)))))...))))....)))))....))))))..))..... ( -35.40) >DroEre_CAF1 89703 95 + 1 CUGCAGACAGUCUGCGGCGGCAGCGUUUGUCUGCCUGCCAGAUUCUUCAGCGCUCU----------------GCAGCUUGGCUUUUUACUGCCUUCCUGUCUUGGCCAAUU ..((((((((...((.((((.((((((.(((((.....))))).....))))))))----------------)).))..(((........)))...))))))..))..... ( -31.50) >DroYak_CAF1 79770 95 + 1 CUGCAGACAGUCUGCGGCGGCAGCGUUUGUCUGCCUGCCAGAUUCUUCAGCGCUCU----------------GCAGCUUAGCUUUUUGCUGCCUUCCUGUCUUGGCCAAUU ..((((((((...(((((((((((....).))))).))).((....)).)).....----------------(((((..........)))))....))))))..))..... ( -30.90) >consensus CUGCAGACAGUCUGCGGCGGCAGCGUUUGUCU____GCCAGAUUCUUCAGCUCUCUGCAGCUC________UGCAGCUUGGCUUUUUGCUGCCUUCCUGUCUUGGCCAAUU ..((((.....))))((((((((((((((.(.....).))))).....((((..((((((..........))))))...))))....))))))...(......)))).... (-24.48 = -26.28 + 1.80)



| Location | 27,376,532 – 27,376,631 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 87.43 |
| Mean single sequence MFE | -35.04 |
| Consensus MFE | -25.02 |
| Energy contribution | -25.76 |
| Covariance contribution | 0.74 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.64 |
| Structure conservation index | 0.71 |
| SVM decision value | 1.20 |
| SVM RNA-class probability | 0.930222 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 27376532 99 - 27905053 AAUUGGCCAAGACAGGAAGGCAGCAAAAAGCCAAGCUGCA--------GAGCUGCAGAGAGCUGAAGAAUCUGGC----AGACAAACGCUGCCGCCGCAGACUGUCUGCAG .....((..((((((....((.((....(((....(((((--------....)))))...))).........(((----((.(....)))))))).))...)))))))).. ( -33.80) >DroSec_CAF1 121912 107 - 1 AAUUGGCCAAGACAGGAAGGCAGCAAAAAGCCAAGCUGCAAAGCUGCUGAGCUGCAGAGAGCUGAAGAAUCUGGC----AGACAAACGCUGCCGCCGCAGACUGUCUGCAG ....(((..((.(((....(((((..........)))))....)))))..)))(((((.(((((.......((((----((.(....)))))))...))).)).))))).. ( -37.40) >DroSim_CAF1 86134 107 - 1 AAUUGGCCAAGACAGGAAGGCAGCAAAAAGCCAAGCUGCAAAGCUGCUGAGCUGCAGAGAGCUGAAGAAUCUGGC----AGACAAACGCUGCCGCCGCAGACUGUCUGCAG ....(((..((.(((....(((((..........)))))....)))))..)))(((((.(((((.......((((----((.(....)))))))...))).)).))))).. ( -37.40) >DroEre_CAF1 89703 95 - 1 AAUUGGCCAAGACAGGAAGGCAGUAAAAAGCCAAGCUGC----------------AGAGCGCUGAAGAAUCUGGCAGGCAGACAAACGCUGCCGCCGCAGACUGUCUGCAG .....((..((((((...(((........)))...((((----------------(((...(....)..)))(((.(((((.(....))))))))))))).)))))))).. ( -34.40) >DroYak_CAF1 79770 95 - 1 AAUUGGCCAAGACAGGAAGGCAGCAAAAAGCUAAGCUGC----------------AGAGCGCUGAAGAAUCUGGCAGGCAGACAAACGCUGCCGCCGCAGACUGUCUGCAG .....((((.((((((...(((((..........)))))----------------....).))).....))))))..(((((((....((((....))))..))))))).. ( -32.20) >consensus AAUUGGCCAAGACAGGAAGGCAGCAAAAAGCCAAGCUGCA________GAGCUGCAGAGAGCUGAAGAAUCUGGC____AGACAAACGCUGCCGCCGCAGACUGUCUGCAG ....((((......)...((((((....(((....(((((............)))))...)))......(((.......))).....)))))))))((((.....)))).. (-25.02 = -25.76 + 0.74)

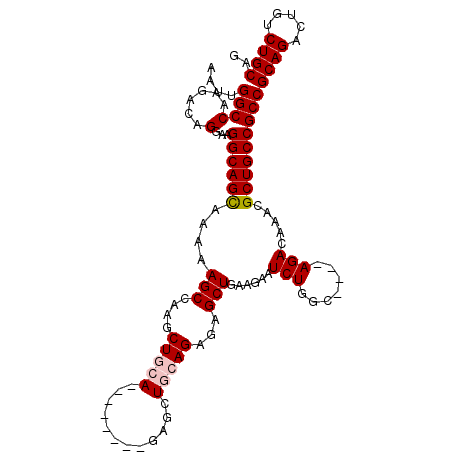
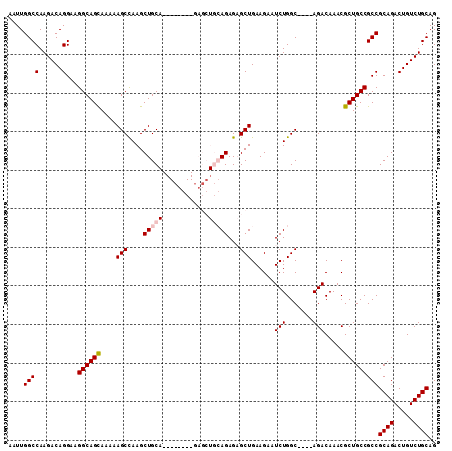
| Location | 27,376,591 – 27,376,698 |
|---|---|
| Length | 107 |
| Sequences | 5 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 97.75 |
| Mean single sequence MFE | -34.92 |
| Consensus MFE | -33.86 |
| Energy contribution | -34.10 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.97 |
| SVM decision value | 1.74 |
| SVM RNA-class probability | 0.975066 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 27376591 107 - 27905053 GGCUUUGCAAGCCUUAUGAUUUCGCCUUUGCCGACGUUGCGCUAAUUUGUUGGCAUCCAUCGAGUGGAAUUGGCCAAGACAGGAAGGCAGCAAAAAGCCAAGCUGCA ((((((((..(((((.((.(((.(((..((((((((...........))))))))(((((...)))))...))).))).))..))))).)))..)))))........ ( -36.70) >DroSec_CAF1 121979 107 - 1 GGCUUUGCAAGCCUUAUGAUUUCGCCUUUGCCGACGUUGCGCUAAAUUGUUGGCAUCCAUCGAGUGGAAUUGGCCAAGACAGGAAGGCAGCAAAAAGCCAAGCUGCA ((((((((..(((((.((.(((.(((..((((((((...........))))))))(((((...)))))...))).))).))..))))).)))..)))))........ ( -36.70) >DroSim_CAF1 86201 107 - 1 GGCUUUGCAAGCCUUAUGAUUUCGCCUUUGCCGACGUUGCGCUAAUUUGUUGGCAUCCAUCGAGUGGAAUUGGCCAAGACAGGAAGGCAGCAAAAAGCCAAGCUGCA ((((((((..(((((.((.(((.(((..((((((((...........))))))))(((((...)))))...))).))).))..))))).)))..)))))........ ( -36.70) >DroEre_CAF1 89759 106 - 1 UGCUUUGCAAGCCUUAUGAUUUCGCCUUUGCCGACGUUGCGCUAAUUUGUUGGCAUCCAUCGAGUGGAAUUGGCCAAGACAGGAAGGCAGUAAAAAGCCAAGCUGC- .(((((....(((((.((.(((.(((..((((((((...........))))))))(((((...)))))...))).))).))..))))).....)))))........- ( -31.60) >DroYak_CAF1 79826 106 - 1 UGCUUUGCAAGCCUUAUGAUUUCGCCUUUGCCGACGUUGCGCUAAUUUGUUGGCAUCCAUCGAGUGGAAUUGGCCAAGACAGGAAGGCAGCAAAAAGCUAAGCUGC- .(((((((..(((((.((.(((.(((..((((((((...........))))))))(((((...)))))...))).))).))..))))).)))..))))........- ( -32.90) >consensus GGCUUUGCAAGCCUUAUGAUUUCGCCUUUGCCGACGUUGCGCUAAUUUGUUGGCAUCCAUCGAGUGGAAUUGGCCAAGACAGGAAGGCAGCAAAAAGCCAAGCUGCA ((((((((..(((((.((.(((.(((..((((((((...........))))))))(((((...)))))...))).))).))..))))).)))..)))))........ (-33.86 = -34.10 + 0.24)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:51:21 2006