
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 27,310,854 – 27,310,994 |
| Length | 140 |
| Max. P | 0.978830 |

| Location | 27,310,854 – 27,310,954 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.12 |
| Mean single sequence MFE | -36.29 |
| Consensus MFE | -31.65 |
| Energy contribution | -31.34 |
| Covariance contribution | -0.30 |
| Combinations/Pair | 1.21 |
| Mean z-score | -2.64 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.82 |
| SVM RNA-class probability | 0.978830 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 27310854 100 - 27905053 ACGAAAUCCAGGGGGCUUGUAAUAAAUGCGAUUCUGUAACCCAUUUCUGGGCUUUCUGGGUUAAGGGCAAGUGGGACAGCU---------------GCUCAAAUGUGUCCCACAA----- .(..(((((((..((((((((.....))))).........(((....))))))..)))))))..).....((((((((.(.---------------........)))))))))..----- ( -34.00) >DroSec_CAF1 15757 100 - 1 ACGAAAUCCAGGGGGCUUGUAAUAAAUGCGAUUCUGUAACCCAUUUCGGGGCUUUCUGGGUUAAGGGCAAGUGGGACAGCU---------------GCUCAAAUGUGUCCCACAA----- .(..(((((((..((((((((.....)))))........(((.....))))))..)))))))..).....((((((((.(.---------------........)))))))))..----- ( -36.60) >DroSim_CAF1 16213 100 - 1 ACGAAAUCCAGGGGGCUUGUAAUAAAUGCGAUUCUGUAACCCAUUUCGGGGCUUUCUGGGUUAAGGGCAAGUGGGACAGCU---------------GCUCAAAUGUGUCCCACAA----- .(..(((((((..((((((((.....)))))........(((.....))))))..)))))))..).....((((((((.(.---------------........)))))))))..----- ( -36.60) >DroEre_CAF1 16041 100 - 1 ACGAAAUCCAGGGGGCUUGUAAUAAAUGCGAUUCUGUAACCCAUUUCGGGGCUUUCUGGGUUAAGGGCAAGUGGGACAGCU---------------GCUCAAAUGUGUCCCACAA----- .(..(((((((..((((((((.....)))))........(((.....))))))..)))))))..).....((((((((.(.---------------........)))))))))..----- ( -36.60) >DroYak_CAF1 12134 100 - 1 ACGAAAUCCAGGGGGCUUGUAAUAAAUGCGAUUCUGUAACCCAUUUCGGGGCUUUCUGGGUUAAGGGCAAGUGGGACAGCU---------------GCUCAAAUGUGUCCCACAA----- .(..(((((((..((((((((.....)))))........(((.....))))))..)))))))..).....((((((((.(.---------------........)))))))))..----- ( -36.60) >DroAna_CAF1 17695 120 - 1 ACGAAACCCGGCUAGCUUGUCAUAAAUGCGAUUCUGUAACCCAUUUUGCGACUAUUUGGGUUAAGGGCAAGUGGGACAGAUCCACGGAUCACAUUGGCUUGGAUUCGUCCCACAAUUUAG ......((((((......)))...............(((((((.............))))))).)))...(((((((..(((((.((.((.....)))))))))..)))))))....... ( -37.32) >consensus ACGAAAUCCAGGGGGCUUGUAAUAAAUGCGAUUCUGUAACCCAUUUCGGGGCUUUCUGGGUUAAGGGCAAGUGGGACAGCU_______________GCUCAAAUGUGUCCCACAA_____ .(..(((((((..((((((((.....)))))........(((.....))))))..)))))))..).....(((((((.((........................)))))))))....... (-31.65 = -31.34 + -0.30)

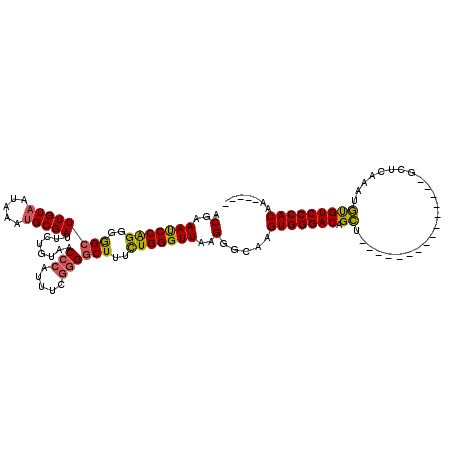

| Location | 27,310,874 – 27,310,994 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.44 |
| Mean single sequence MFE | -39.73 |
| Consensus MFE | -35.98 |
| Energy contribution | -35.98 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.25 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.752347 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 27310874 120 - 27905053 GAGCUGGCCAAAAAGCCUGACUUAUUUGUGGCCGCCAGACACGAAAUCCAGGGGGCUUGUAAUAAAUGCGAUUCUGUAACCCAUUUCUGGGCUUUCUGGGUUAAGGGCAAGUGGGACAGC ..((((.(((....((((.......(((((.........)))))(((((((..((((((((.....))))).........(((....))))))..)))))))..))))...)))..)))) ( -38.50) >DroSec_CAF1 15777 120 - 1 GAGCUGGCCAAAAAGCCUGACUUAUUUGUGGCCGCCAGACACGAAAUCCAGGGGGCUUGUAAUAAAUGCGAUUCUGUAACCCAUUUCGGGGCUUUCUGGGUUAAGGGCAAGUGGGACAGC ..((((.(((....((((.......(((((.........)))))(((((((..((((((((.....)))))........(((.....))))))..)))))))..))))...)))..)))) ( -41.10) >DroSim_CAF1 16233 120 - 1 GAGCUGGCCAAAAAGCCUGACUUAUUUGUGGCCGCCAGACACGAAAUCCAGGGGGCUUGUAAUAAAUGCGAUUCUGUAACCCAUUUCGGGGCUUUCUGGGUUAAGGGCAAGUGGGACAGC ..((((.(((....((((.......(((((.........)))))(((((((..((((((((.....)))))........(((.....))))))..)))))))..))))...)))..)))) ( -41.10) >DroEre_CAF1 16061 120 - 1 GAGCUGGCCAAAAAGCCUGACUUAUUUGUGGCCGCCAGACACGAAAUCCAGGGGGCUUGUAAUAAAUGCGAUUCUGUAACCCAUUUCGGGGCUUUCUGGGUUAAGGGCAAGUGGGACAGC ..((((.(((....((((.......(((((.........)))))(((((((..((((((((.....)))))........(((.....))))))..)))))))..))))...)))..)))) ( -41.10) >DroYak_CAF1 12154 120 - 1 GAGCUGGCCAAAAAGCCUGACUUAUUUGUGGUCGCCAGACACGAAAUCCAGGGGGCUUGUAAUAAAUGCGAUUCUGUAACCCAUUUCGGGGCUUUCUGGGUUAAGGGCAAGUGGGACAGC ..((((.(((....((((.......(((((.((....)))))))(((((((..((((((((.....)))))........(((.....))))))..)))))))..))))...)))..)))) ( -42.50) >DroAna_CAF1 17735 117 - 1 GAGCUGGUCG---AGCCUGACUUAUUUGUGGCAGCCAGACACGAAACCCGGCUAGCUUGUCAUAAAUGCGAUUCUGUAACCCAUUUUGCGACUAUUUGGGUUAAGGGCAAGUGGGACAGA ..((.(((((---....))))).((((((((((((.((...((.....)).)).)).))))))))))))...(((((..(((((((.((((((.....))))....)))))))))))))) ( -34.10) >consensus GAGCUGGCCAAAAAGCCUGACUUAUUUGUGGCCGCCAGACACGAAAUCCAGGGGGCUUGUAAUAAAUGCGAUUCUGUAACCCAUUUCGGGGCUUUCUGGGUUAAGGGCAAGUGGGACAGC ..((((.(((....((((.......(((((.........)))))(((((((..((((((((.....)))))........(((.....))))))..)))))))..))))...)))..)))) (-35.98 = -35.98 + 0.00)

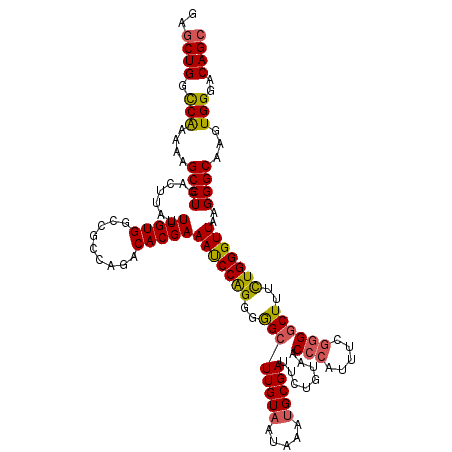
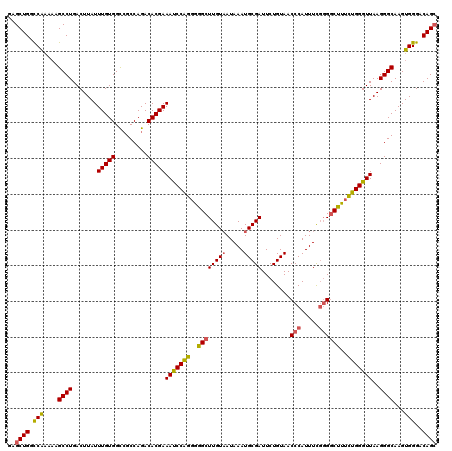
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:50:50 2006