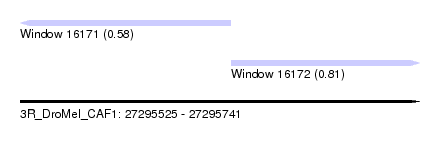
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 27,295,525 – 27,295,741 |
| Length | 216 |
| Max. P | 0.812468 |

| Location | 27,295,525 – 27,295,639 |
|---|---|
| Length | 114 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 90.76 |
| Mean single sequence MFE | -42.20 |
| Consensus MFE | -29.42 |
| Energy contribution | -29.42 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.577501 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 27295525 114 - 27905053 CAGCCCUGAGCGCCAACUGCACC---ACCAUUGUGGUGCCCUUGCGGCACCAGUGGUUCCGCUGGUCCGUCGGCCAAGUGAGUGACAGCCAUCACACUGAGCUGAAAUACUCGAACA ..((.....))(((....(((((---((....)))))))....((((.(((((((....))))))))))).)))....((((((.((((((......)).))))...)))))).... ( -45.00) >DroSec_CAF1 597 114 - 1 CAGCACGGAGCGCCAACUGCACC---GCCAUUGUGGUGCCCUUGCGCCACCAGUGGUGCCGCUGUUCCGUCGGCCAAGUGAGUGACAGCCAUCACACCGAACUGAAAUACUCUAAUA (((..((((((((.....(((((---((....(((((((....)))))))..))))))).)).))))))((((....((((.((.....)))))).)))).)))............. ( -41.20) >DroSim_CAF1 1005 114 - 1 CAGCACGGAGCGCCAACUGCACC---ACCAUUGUGGUGCCCUUGCGCCACCAGUGGUGCCGCUGGUCCGUCGGCCAAGUGAGUGACAGCCAUCACACCGAACUGAAAUACUCGAAUA (((..((((((((.((..(((((---((....)))))))..))))))..((((((....))))))))))((((....((((.((.....)))))).)))).)))............. ( -41.90) >DroEre_CAF1 744 112 - 1 CAGCCCUGAGCGCCAACUGCACCACCACCAUUGUGGUGCCCUUGCGCCACCAGUGGUUCCGCUGGUCUGUCGGUCGAGUGAGUGACAGCCAUCACAC-----UGAAAUACUCGAAUA .....((((((((.((..(((((((.......)))))))..)))))).(((((((....)))))))...))))(((((((((((..........)))-----)....)))))))... ( -40.70) >consensus CAGCACGGAGCGCCAACUGCACC___ACCAUUGUGGUGCCCUUGCGCCACCAGUGGUGCCGCUGGUCCGUCGGCCAAGUGAGUGACAGCCAUCACACCGAACUGAAAUACUCGAAUA .......(((......((((((....(((((((((((((....)))))).)))))))..((((((((....))))).))).))).)))...(((........)))....)))..... (-29.42 = -29.42 + 0.00)


| Location | 27,295,639 – 27,295,741 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 84.32 |
| Mean single sequence MFE | -23.35 |
| Consensus MFE | -16.75 |
| Energy contribution | -16.62 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.65 |
| SVM RNA-class probability | 0.812468 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 27295639 102 + 27905053 CCCGAGUCACUGGGUCAACUUGGUGAAGGCAAUGCAAUGCAGAGGGAGACAGCCAAAUCGGAUCCAAAAA-UUCAGAUAAAA-AUAU--------UUAA-----AAAUGGACAUAAA .((((.((((..((....))..)))).(((..(((...)))...(....).)))...)))).((((....-...(((((...-.)))--------))..-----...))))...... ( -24.40) >DroSec_CAF1 711 102 + 1 CCCGAGUCACUGGGUCAACUGAGUGAAGGCAAUGCAAUAUAGAGGGAGACAGCCAAAUCGGAUCCAAAAA-UGCAGGUAAAA-AUAG--------UUAC-----AAAUGCACAUAAA .((((.(((((.((....)).))))).(((....(......)..(....).)))...)))).........-((((.((((..-....--------))))-----...))))...... ( -24.40) >DroSim_CAF1 1119 102 + 1 CCCGAGUCACUGGGUCAACUGGGUGAAGGCAAUGCAAUGCAGAGGGAGACAGCCAAAUCGGAUCCAAAAA-UUCAGAUAAAA-AUAG--------UUAA-----AAAUGCACAUAAA .((((.(((((.((....)).))))).(((..(((...)))...(....).)))...)))).........-...........-....--------....-----............. ( -21.40) >DroEre_CAF1 856 112 + 1 CCCGAGUCACUGGGUCAACUGGGUGAAGGCAAUG-----CAGAGGGAGAGAGCCAAAUCGGACCCAAAAAGUUCAAAUAAAAAAUAGGUGUAGCUUUAAAAAUAAAAUGUAUGUAAA .((...(((((.((....)).))))).))..(((-----((..(((...((......))...)))..((((((((..((.....))..)).))))))..........)))))..... ( -23.20) >consensus CCCGAGUCACUGGGUCAACUGGGUGAAGGCAAUGCAAUGCAGAGGGAGACAGCCAAAUCGGAUCCAAAAA_UUCAGAUAAAA_AUAG________UUAA_____AAAUGCACAUAAA .((((.(((((.((....)).))))).(((..((.....))...(....).)))...))))........................................................ (-16.75 = -16.62 + -0.12)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:50:14 2006