
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 27,146,089 – 27,146,181 |
| Length | 92 |
| Max. P | 0.960911 |

| Location | 27,146,089 – 27,146,181 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 78.53 |
| Mean single sequence MFE | -20.24 |
| Consensus MFE | -9.04 |
| Energy contribution | -9.00 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.21 |
| Mean z-score | -2.49 |
| Structure conservation index | 0.45 |
| SVM decision value | 1.52 |
| SVM RNA-class probability | 0.960911 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 27146089 92 + 27905053 GCAAA-CUU-UGCAGUAGCAGUGGCAAAAUGGCAAACUGGCAAAAAA------GAAAAUGAAAAGCGAGGAAAGGCCACAAUGAGAGCUGCUAAUAGCUG .....-...-.((..(((((((..((...((((...((.((......------...........)).)).....))))...))...)))))))...)).. ( -22.73) >DroSim_CAF1 50166 92 + 1 GAAAA-CUG-UGCAGUAGCAUUGGCCAAAUGGCAAACUGGCAAAAAA------GAAAAUGAAAAGCGAGGAAAGGCCACAAUGAGAGCUGCUAAUAGCUG .....-(((-((((((..((((((((....)))....((((......------.....(....)..........)))))))))...)))))..))))... ( -22.20) >DroEre_CAF1 48479 97 + 1 GAAAA-CUU-UGCAGUAGCAGUGGCAAAAUGGAA-AAUGGCAAAAAAGAAGUUGAAAAUGAAUAGCGAGGAAAGGCCACAAUGAGAGCUGCUAAUAGCUG .....-...-.((..(((((((..((...(....-).((((.........((((........))))........))))...))...)))))))...)).. ( -17.43) >DroYak_CAF1 50647 84 + 1 GAAAA-CUUUUCCAGUAGCAGUGGCAAAAU--------GGCAAAAA-------GAAAAUGAAAAGCGAGGAAAGGCCACAAUGAGAGCUGCUAAUAGCUG .....-......((((((((((..((...(--------(((.....-------.....(....)..........))))...))...))))))....)))) ( -16.85) >DroAna_CAF1 52731 83 + 1 AAAAAGUUU-UCCAGUAGCGGCAGCAG----UUAUAGCCGCUAAAAA------U------AAAACUGAGGAAAGGCAACAAUGAGAGUUGCUAAUAGCUG ....(((((-(....(((((((.....----.....)))))))....------.------)))))).......((((((.......))))))........ ( -22.00) >consensus GAAAA_CUU_UGCAGUAGCAGUGGCAAAAUGGCA_ACUGGCAAAAAA______GAAAAUGAAAAGCGAGGAAAGGCCACAAUGAGAGCUGCUAAUAGCUG ............((((((((((.((..............))...........................((.....)).........))))))....)))) ( -9.04 = -9.00 + -0.04)

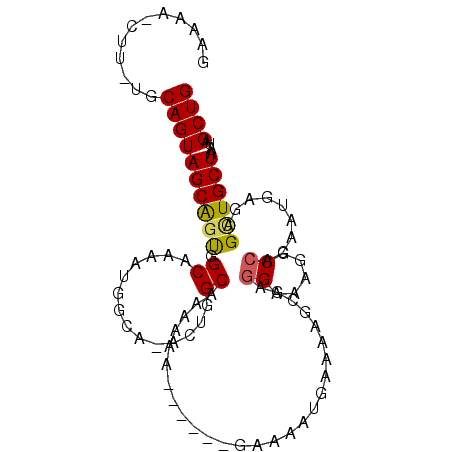

| Location | 27,146,089 – 27,146,181 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 78.53 |
| Mean single sequence MFE | -16.96 |
| Consensus MFE | -6.05 |
| Energy contribution | -6.49 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.24 |
| Structure conservation index | 0.36 |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.861098 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 27146089 92 - 27905053 CAGCUAUUAGCAGCUCUCAUUGUGGCCUUUCCUCGCUUUUCAUUUUC------UUUUUUGCCAGUUUGCCAUUUUGCCACUGCUACUGCA-AAG-UUUGC ..((...((((((....((..(((((.....((.((...........------......)).))...)))))..))...))))))..)).-...-..... ( -18.83) >DroSim_CAF1 50166 92 - 1 CAGCUAUUAGCAGCUCUCAUUGUGGCCUUUCCUCGCUUUUCAUUUUC------UUUUUUGCCAGUUUGCCAUUUGGCCAAUGCUACUGCA-CAG-UUUUC ..((...(((((....(((..(((((.....((.((...........------......)).))...))))).)))....)))))..)).-...-..... ( -16.43) >DroEre_CAF1 48479 97 - 1 CAGCUAUUAGCAGCUCUCAUUGUGGCCUUUCCUCGCUAUUCAUUUUCAACUUCUUUUUUGCCAUU-UUCCAUUUUGCCACUGCUACUGCA-AAG-UUUUC ..((...((((((....((..((((.........((.......................))....-..))))..))...))))))..)).-...-..... ( -12.96) >DroYak_CAF1 50647 84 - 1 CAGCUAUUAGCAGCUCUCAUUGUGGCCUUUCCUCGCUUUUCAUUUUC-------UUUUUGCC--------AUUUUGCCACUGCUACUGGAAAAG-UUUUC ...(((.((((((....((..(((((.....................-------.....)))--------))..))...)))))).))).....-..... ( -15.77) >DroAna_CAF1 52731 83 - 1 CAGCUAUUAGCAACUCUCAUUGUUGCCUUUCCUCAGUUUU------A------UUUUUAGCGGCUAUAA----CUGCUGCCGCUACUGGA-AAACUUUUU .........(((((.......)))))........((((((------.------....(((((((.....----.....)))))))....)-))))).... ( -20.80) >consensus CAGCUAUUAGCAGCUCUCAUUGUGGCCUUUCCUCGCUUUUCAUUUUC______UUUUUUGCCAGU_UGCCAUUUUGCCACUGCUACUGCA_AAG_UUUUC (((....(((((.........(((((.................................................)))))))))))))............ ( -6.05 = -6.49 + 0.44)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:49:19 2006