
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 27,118,808 – 27,118,901 |
| Length | 93 |
| Max. P | 0.931738 |

| Location | 27,118,808 – 27,118,901 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 63.59 |
| Mean single sequence MFE | -23.32 |
| Consensus MFE | -10.31 |
| Energy contribution | -11.50 |
| Covariance contribution | 1.19 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.86 |
| Structure conservation index | 0.44 |
| SVM decision value | 0.52 |
| SVM RNA-class probability | 0.769130 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 27118808 93 + 27905053 UUUCCACAUAGGCGACAUAUACAUACAU--AUGUACAUAUUGGCAAGUGGCCAAAGUGAACCGCAGUGGAAGUUGUUGGCGAUGGAAGGGGCAAA .((((((....(((((((((......))--)))).(((.(((((.....))))).)))...))).))))))((((....))))............ ( -25.20) >DroSec_CAF1 20831 85 + 1 UUUCCACAUAGGCGACACAUACAAAC----------AUAUUGGCAAGUGGCCAAAGUGAGCCGCAGUGGAAGUUGUUGGCGGUGGAAGGGGCAAA (((((((...(.((((((.......(----------((.(((((.....))))).)))..((.....))..).))))).).)))))))....... ( -25.10) >DroSim_CAF1 22525 77 + 1 UUUCCACAUAGGCGACAC------------------AUAUUGGCAAGUGGCCAAAGUGGGCCGCAGUGGAAGUUGUUGGCGGUGGAAGGGGCAAA (((((((...(.(((((.------------------((.((.((..((((((......)))))).)).)).))))))).).)))))))....... ( -28.90) >DroWil_CAF1 30539 74 + 1 AUUUCAUAUUUGCU---UGU----UUUUCUGUCAUCAUAUUGGCAAGUGUCCAA-GUGAGCCAAAAAUGAAGA-------------AACACUGGA .((((((.((((((---..(----((..((((((......)))).)).....))-)..)).)))).)))))).-------------......... ( -14.10) >consensus UUUCCACAUAGGCGACACAU____AC__________AUAUUGGCAAGUGGCCAAAGUGAGCCGCAGUGGAAGUUGUUGGCGGUGGAAGGGGCAAA .((((((...(((......................(((.(((((.....))))).))).)))...))))))........................ (-10.31 = -11.50 + 1.19)



| Location | 27,118,808 – 27,118,901 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 63.59 |
| Mean single sequence MFE | -20.46 |
| Consensus MFE | -8.46 |
| Energy contribution | -10.15 |
| Covariance contribution | 1.69 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.43 |
| Structure conservation index | 0.41 |
| SVM decision value | 1.22 |
| SVM RNA-class probability | 0.931738 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 27118808 93 - 27905053 UUUGCCCCUUCCAUCGCCAACAACUUCCACUGCGGUUCACUUUGGCCACUUGCCAAUAUGUACAU--AUGUAUGUAUAUGUCGCCUAUGUGGAAA ........................((((((.((((......(((((.....)))))(((((((((--....)))))))))))))....)))))). ( -23.40) >DroSec_CAF1 20831 85 - 1 UUUGCCCCUUCCACCGCCAACAACUUCCACUGCGGCUCACUUUGGCCACUUGCCAAUAU----------GUUUGUAUGUGUCGCCUAUGUGGAAA ........................((((((.(((((.((..(((((.....)))))(((----------....))))).)))))....)))))). ( -20.00) >DroSim_CAF1 22525 77 - 1 UUUGCCCCUUCCACCGCCAACAACUUCCACUGCGGCCCACUUUGGCCACUUGCCAAUAU------------------GUGUCGCCUAUGUGGAAA ........................((((((.(((((.((..(((((.....)))))..)------------------).)))))....)))))). ( -23.60) >DroWil_CAF1 30539 74 - 1 UCCAGUGUU-------------UCUUCAUUUUUGGCUCAC-UUGGACACUUGCCAAUAUGAUGACAGAAAA----ACA---AGCAAAUAUGAAAU ....((.((-------------((((((((.(((((....-..........)))))...))))).))))).----)).---.............. ( -14.84) >consensus UUUGCCCCUUCCACCGCCAACAACUUCCACUGCGGCUCACUUUGGCCACUUGCCAAUAU__________GU____AUAUGUCGCCUAUGUGGAAA ........................((((((...(((.(((.(((((.....))))).....................)))..)))...)))))). ( -8.46 = -10.15 + 1.69)

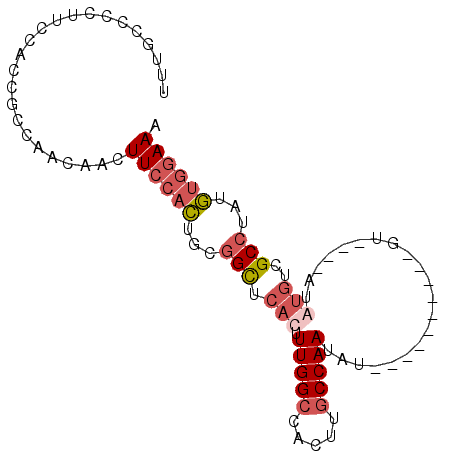
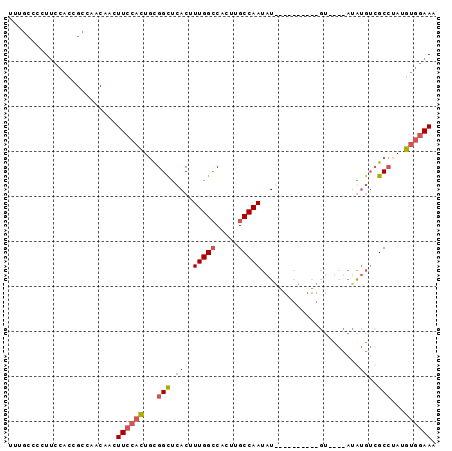
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:49:09 2006