
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 26,921,112 – 26,921,210 |
| Length | 98 |
| Max. P | 0.984381 |

| Location | 26,921,112 – 26,921,210 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 90.71 |
| Mean single sequence MFE | -25.78 |
| Consensus MFE | -22.26 |
| Energy contribution | -21.22 |
| Covariance contribution | -1.04 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.97 |
| SVM RNA-class probability | 0.984381 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 26921112 98 + 27905053 UUUCUUCUUGGCAACAUCUGCUGGCAGUAGUAAAUCAAAUGCCAGAAGCCCCUUAAGUAACUUUAUAUGCCACAGCUGUUUCGUAAAGUAAACAGCAG ........(((((.......((((((.............))))))..((.......)).........)))))..((((((((.....).))))))).. ( -22.52) >DroSec_CAF1 23847 95 + 1 ---CUUCUUGGCAACAGCUGCUGGCAGUAGUAAAGCAAAUGCCAGAAGCCCCUUAAGUAACUUUAUAUGCCACAGCUGUUUCGUAGAGUAAACAGCAG ---.....(((((...(((.((((((...(.....)...)))))).))).......(((....))).)))))..((((((((.....).))))))).. ( -27.10) >DroSim_CAF1 23020 98 + 1 UUUCUUCUUGGCAACAGCUGCUGGCAGUAGUAAAUCAAAUGCCAGAAGCCCCUUAAGUAACUUUAUGUGCCAUAGCUGUUUCGUAGAGUAAACAGCAG ........(((((.(((((.((((((.............)))))).)))....((((....)))))))))))..((((((((.....).))))))).. ( -26.92) >DroEre_CAF1 22130 98 + 1 UUUCUUCUGGGCAACAGCUGCUGGCAGUAGUAAAUCAAAUGCAAGAAGCCCUUUAAGUAAGUUUAUAUGCCACAGCUGUUCCCUUAAGCAAACAGCAG ...(((..(((.((((((((.(((((...((((((.......(((.....))).......)))))).))))))))))))))))..))).......... ( -29.54) >DroYak_CAF1 22796 98 + 1 UUUCUUCUGGGCAACAGCUGCUGGCAGUAGUAAAUCAAAUGCUAGAAGCCCUUUAAGUAACUCUAUAUGCCACAGCUGCUCUCUAAAGCAAACAGCAG ...(((..(((...((((((.(((((((((.........((((.(((....))).))))...)))).)))))))))))..)))..))).......... ( -22.80) >consensus UUUCUUCUUGGCAACAGCUGCUGGCAGUAGUAAAUCAAAUGCCAGAAGCCCCUUAAGUAACUUUAUAUGCCACAGCUGUUUCGUAAAGUAAACAGCAG ...(((..(((.((((((((.(((((((((..........((.....)).............)))).))))))))))))))))..))).......... (-22.26 = -21.22 + -1.04)

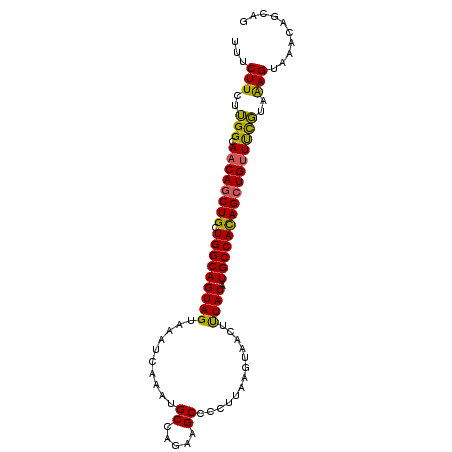

| Location | 26,921,112 – 26,921,210 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 90.71 |
| Mean single sequence MFE | -26.85 |
| Consensus MFE | -22.50 |
| Energy contribution | -22.30 |
| Covariance contribution | -0.20 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.854251 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 26921112 98 - 27905053 CUGCUGUUUACUUUACGAAACAGCUGUGGCAUAUAAAGUUACUUAAGGGGCUUCUGGCAUUUGAUUUACUACUGCCAGCAGAUGUUGCCAAGAAGAAA ..(((((((........)))))))..((((((((.((((..(....)..))))((((((.............))))))...))).)))))........ ( -24.32) >DroSec_CAF1 23847 95 - 1 CUGCUGUUUACUCUACGAAACAGCUGUGGCAUAUAAAGUUACUUAAGGGGCUUCUGGCAUUUGCUUUACUACUGCCAGCAGCUGUUGCCAAGAAG--- ..(((((((........)))))))..((((...(((......)))((.((((.((((((.............)))))).)))).)))))).....--- ( -26.62) >DroSim_CAF1 23020 98 - 1 CUGCUGUUUACUCUACGAAACAGCUAUGGCACAUAAAGUUACUUAAGGGGCUUCUGGCAUUUGAUUUACUACUGCCAGCAGCUGUUGCCAAGAAGAAA ..(((((((........)))))))..((((((.....))......((.((((.((((((.............)))))).)))).))))))........ ( -26.82) >DroEre_CAF1 22130 98 - 1 CUGCUGUUUGCUUAAGGGAACAGCUGUGGCAUAUAAACUUACUUAAAGGGCUUCUUGCAUUUGAUUUACUACUGCCAGCAGCUGUUGCCCAGAAGAAA ..........(((..((((((((((((((((......(((.....))).((.....))..............))))).)))))))).)))..)))... ( -29.50) >DroYak_CAF1 22796 98 - 1 CUGCUGUUUGCUUUAGAGAGCAGCUGUGGCAUAUAGAGUUACUUAAAGGGCUUCUAGCAUUUGAUUUACUACUGCCAGCAGCUGUUGCCCAGAAGAAA ..........((((.(.((((((((((((((.....(((...(((((..((.....)).)))))...)))..))))).)))))))).).).))))... ( -27.00) >consensus CUGCUGUUUACUUUACGAAACAGCUGUGGCAUAUAAAGUUACUUAAGGGGCUUCUGGCAUUUGAUUUACUACUGCCAGCAGCUGUUGCCAAGAAGAAA ..(((((((........)))))))...(((...(((......)))((.((((.((((((.............)))))).)))).)))))......... (-22.50 = -22.30 + -0.20)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:47:41 2006