
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 26,914,632 – 26,914,766 |
| Length | 134 |
| Max. P | 0.939845 |

| Location | 26,914,632 – 26,914,729 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 76.07 |
| Mean single sequence MFE | -22.58 |
| Consensus MFE | -11.79 |
| Energy contribution | -12.40 |
| Covariance contribution | 0.61 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.52 |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.575004 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 26914632 97 - 27905053 GCAGCAAGGCAACCAUUAGCUAAUUAAGACGCUCCGAGUUGCCUUGGAAGCCACCC--------ACGAAAAAACACACACAUUACGGGGACCGCAGACAAAGG-AC ((..(((((((((....(((..........)))....))))))))).......(((--------.((.................)))))...)).........-.. ( -22.53) >DroGri_CAF1 25376 84 - 1 UU------AGAGCCAUUAGCUAAUUAAG-UGCCC-AAUGUUUCUGGGCACCCACUC--------ACAAGGAAACU-----AGUGGAAAAACCGCAGACAAUAGCA- ..------..........((((.....(-(((((-(.......)))))))(((((.--------...((....))-----)))))...............)))).- ( -25.10) >DroSec_CAF1 17416 97 - 1 GCAGCAAGGCAACCAUUAGCUAAUUAAGACGCUCCGAGUUGCCUUGGAAGCCACCC--------ACGAAAAAACACACACAUUACGGGGACCGCAGACAAAGG-AC ((..(((((((((....(((..........)))....))))))))).......(((--------.((.................)))))...)).........-.. ( -22.53) >DroSim_CAF1 16574 97 - 1 GCAGCAAGGCAACCAUUAGCUAAUUAAGACGCUCCGAGUUGCCUUGGAAGCCACCC--------ACGAAAAAACACACACAUUACGGGGACCGCAGACAAAGG-AC ((..(((((((((....(((..........)))....))))))))).......(((--------.((.................)))))...)).........-.. ( -22.53) >DroYak_CAF1 16311 105 - 1 GCAGCAAGGCAACCAUUAGCUAAUUAAGACGCUCCGAGUUGCCUUGGAAGCCGCCCUCGAAAAAACGAAAAAACACACACAUUACGGCAGCCGCAGACAAAGG-AC ((..(((((((((....(((..........)))....)))))))))...((((...(((......)))................))))....)).........-.. ( -23.01) >DroAna_CAF1 22620 87 - 1 GUAGCAAGGCAACCAUUAGCUAAUUAAGACGCUCCGAGUUGCCUUG----CCACCC--------AC------ACCCUUAAAUUUUCGAAACCGCAGACAAAAG-AC ((.((((((((((....(((..........)))....)))))))))----).))..--------..------...(((...(((.((....)).)))...)))-.. ( -19.80) >consensus GCAGCAAGGCAACCAUUAGCUAAUUAAGACGCUCCGAGUUGCCUUGGAAGCCACCC________ACGAAAAAACACACACAUUACGGGAACCGCAGACAAAGG_AC ....(((((((((....(((..........)))....)))))))))............................................................ (-11.79 = -12.40 + 0.61)

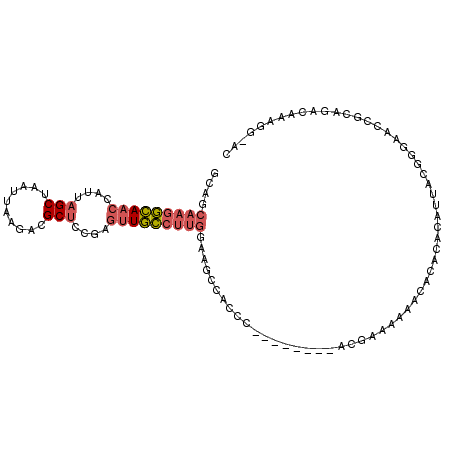

| Location | 26,914,670 – 26,914,766 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 92.91 |
| Mean single sequence MFE | -35.08 |
| Consensus MFE | -31.38 |
| Energy contribution | -31.27 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.801789 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 26914670 96 + 27905053 CGU--------GGGUGGCUUCCAAGGCAACUCGGAGCGUCUUAAUUAGCUAAUGGUUGCCUUGCUGCCUUGCGGCCUUGCAGGCACGAAAAAGGGCUUAGACUG ...--------.(((......((((((((((...(((..........)))...))))))))))..(((((.(((((.....))).))....)))))....))). ( -33.10) >DroSec_CAF1 17454 96 + 1 CGU--------GGGUGGCUUCCAAGGCAACUCGGAGCGUCUUAAUUAGCUAAUGGUUGCCUUGCUGCCUUGCGGCCUUGCAGGCACGAAAAAGGGCUUAGACUG ...--------.(((......((((((((((...(((..........)))...))))))))))..(((((.(((((.....))).))....)))))....))). ( -33.10) >DroSim_CAF1 16612 96 + 1 CGU--------GGGUGGCUUCCAAGGCAACUCGGAGCGUCUUAAUUAGCUAAUGGUUGCCUUGCUGCCUUGCGGCCUUGCAGGCACGAAAAAGGGCUUAGACUG ...--------.(((......((((((((((...(((..........)))...))))))))))..(((((.(((((.....))).))....)))))....))). ( -33.10) >DroEre_CAF1 15711 96 + 1 CGU--------GGGUGGCUUCCAAGGCGACUCGGAGCGUCUUAAUUAGCUAAUGGUUGCCUUGCUGCCUGGCGGCCUUGCAGGCACGAAAAAGGGCUUAGACUG ..(--------(((((((...((((((((((...(((..........)))...))))))))))..)))...(((((.....))).)).......)))))..... ( -32.60) >DroYak_CAF1 16349 104 + 1 CGUUUUUUCGAGGGCGGCUUCCAAGGCAACUCGGAGCGUCUUAAUUAGCUAAUGGUUGCCUUGCUGCCUUGCGGCCUUGCAGGCACGAAAAAGGGCUUAGACUG (.((((((((((((((((....(((((((((...(((..........)))...)))))))))))))))))...(((.....))).)))))))).)......... ( -40.60) >DroAna_CAF1 22653 91 + 1 -GU--------GGGUGG----CAAGGCAACUCGGAGCGUCUUAAUUAGCUAAUGGUUGCCUUGCUACCAUACGGCCUUGCAGGCACGAAAAAGAGCUUAGACGG -..--------.(((((----((((((((((...(((..........)))...)))))))))))))))...(((((.....))).))................. ( -38.00) >consensus CGU________GGGUGGCUUCCAAGGCAACUCGGAGCGUCUUAAUUAGCUAAUGGUUGCCUUGCUGCCUUGCGGCCUUGCAGGCACGAAAAAGGGCUUAGACUG ...........(((((((...((((((((((...(((..........)))...))))))))))..)))...(((((.....))).)).......))))...... (-31.38 = -31.27 + -0.11)



| Location | 26,914,670 – 26,914,766 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 92.91 |
| Mean single sequence MFE | -30.39 |
| Consensus MFE | -27.43 |
| Energy contribution | -27.79 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.90 |
| SVM decision value | 1.30 |
| SVM RNA-class probability | 0.939845 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 26914670 96 - 27905053 CAGUCUAAGCCCUUUUUCGUGCCUGCAAGGCCGCAAGGCAGCAAGGCAACCAUUAGCUAAUUAAGACGCUCCGAGUUGCCUUGGAAGCCACCC--------ACG ..((((.(((.........(((((((...(((....))).)).))))).......))).....))))((((((((....))))).))).....--------... ( -31.09) >DroSec_CAF1 17454 96 - 1 CAGUCUAAGCCCUUUUUCGUGCCUGCAAGGCCGCAAGGCAGCAAGGCAACCAUUAGCUAAUUAAGACGCUCCGAGUUGCCUUGGAAGCCACCC--------ACG ..((((.(((.........(((((((...(((....))).)).))))).......))).....))))((((((((....))))).))).....--------... ( -31.09) >DroSim_CAF1 16612 96 - 1 CAGUCUAAGCCCUUUUUCGUGCCUGCAAGGCCGCAAGGCAGCAAGGCAACCAUUAGCUAAUUAAGACGCUCCGAGUUGCCUUGGAAGCCACCC--------ACG ..((((.(((.........(((((((...(((....))).)).))))).......))).....))))((((((((....))))).))).....--------... ( -31.09) >DroEre_CAF1 15711 96 - 1 CAGUCUAAGCCCUUUUUCGUGCCUGCAAGGCCGCCAGGCAGCAAGGCAACCAUUAGCUAAUUAAGACGCUCCGAGUCGCCUUGGAAGCCACCC--------ACG ..((((.(((.........(((((((...(((....))).)).))))).......))).....))))((((((((....))))).))).....--------... ( -28.09) >DroYak_CAF1 16349 104 - 1 CAGUCUAAGCCCUUUUUCGUGCCUGCAAGGCCGCAAGGCAGCAAGGCAACCAUUAGCUAAUUAAGACGCUCCGAGUUGCCUUGGAAGCCGCCCUCGAAAAAACG ............(((((((.((..((...(((....)))..(((((((((....(((..........)))....)))))))))...)).))...)))))))... ( -32.60) >DroAna_CAF1 22653 91 - 1 CCGUCUAAGCUCUUUUUCGUGCCUGCAAGGCCGUAUGGUAGCAAGGCAACCAUUAGCUAAUUAAGACGCUCCGAGUUGCCUUG----CCACCC--------AC- .................((.(((.....)))))...(((.((((((((((....(((..........)))....)))))))))----).))).--------..- ( -28.40) >consensus CAGUCUAAGCCCUUUUUCGUGCCUGCAAGGCCGCAAGGCAGCAAGGCAACCAUUAGCUAAUUAAGACGCUCCGAGUUGCCUUGGAAGCCACCC________ACG ..((((.(((.........(((((((...(((....))).)).))))).......))).....))))((((((((....))))).)))................ (-27.43 = -27.79 + 0.36)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:47:38 2006