
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 26,886,571 – 26,886,753 |
| Length | 182 |
| Max. P | 0.981894 |

| Location | 26,886,571 – 26,886,661 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 76.11 |
| Mean single sequence MFE | -31.40 |
| Consensus MFE | -20.11 |
| Energy contribution | -20.08 |
| Covariance contribution | -0.03 |
| Combinations/Pair | 1.28 |
| Mean z-score | -2.29 |
| Structure conservation index | 0.64 |
| SVM decision value | 1.90 |
| SVM RNA-class probability | 0.981894 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 26886571 90 - 27905053 ------------------UCGUCUUUGUGUUGGUGUGAAAAUGCUGAAGCAGGAGGGAUAGCAACGUGCUCGCGGUAGAAACCGUGCGAGUGCGGGCGGGACGACUAG ------------------((((((((((.(..((((....))))..).))))........((..((..((((((((....)))..)))))..)).)).)))))).... ( -34.10) >DroVir_CAF1 136656 108 - 1 AUACGUAUAGACUUGUUGGCAUGCUUGUGUUGGUGUGAAAACGCUGAAACAGUCAGGCUAGCAACGUGCUCGCUGUAGAGCCAACAAUAGUGCAAGCGAGACAACCCU .(((((.......(((((((.((.((((.(..((((....))))..).)))).)).))))))))))))((((((.....((........))...))))))........ ( -33.31) >DroSec_CAF1 94809 90 - 1 ------------------UUUUCUUUGUGUUGGUGUGAAAAUGCUGAAGCAGGAGGGAUAGCAACGUGCUCGCGGUAGAAACAGUGUGAGUGCGGGCGGGACGACUAG ------------------..((((((.(((((((((....)))))..)))).))))))..((..((..((((((.(......).))))))..)).))........... ( -27.90) >DroSim_CAF1 98482 90 - 1 ------------------UCGUCUUUGUGUUGGUGUGAAAAUGCUGAAGCAGGAGGGAUAGCAACGUGCUCGCGGUAAAAACCGUGUGAGUGCGGGCGGGACGACUAG ------------------((((((((((.(..((((....))))..).))))........((..((..((((((((....)))))..)))..)).)).)))))).... ( -32.70) >DroWil_CAF1 22409 90 - 1 ------------------UUGCAGUUGUAGUGGUGUGAAAACGCUGAAACAGUCUGGCUAGCAACGUGCUGCCUGUAUAAACAGCUCAAGAGCAAGCGAGACAUCUAU ------------------.......(((..((((((....))))))..)))((((.(((.((...(.((((..........)))).)....)).))).))))...... ( -25.70) >DroYak_CAF1 109542 90 - 1 ------------------UCGUCUUUGUGUUGGUGUGAAAAUGCUGAAGCAGGAGGGAUAGCAACGUGCUCGCGGUAGAAACCGCGUGAGUGCGGGCGAGACGACUAG ------------------((((((((((.(..((((....))))..).))).........((..((..((((((((....)))))..)))..)).))))))))).... ( -34.70) >consensus __________________UCGUCUUUGUGUUGGUGUGAAAAUGCUGAAGCAGGAGGGAUAGCAACGUGCUCGCGGUAGAAACAGCGUGAGUGCGGGCGAGACGACUAG .....................(((((((.(((((((....))))))).)))).)))....((...(..((((((((....)))))..)))..)..))........... (-20.11 = -20.08 + -0.03)


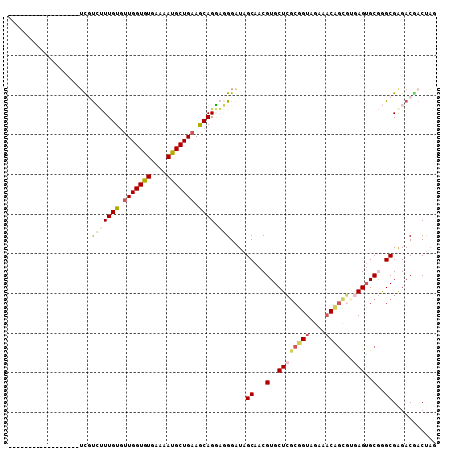
| Location | 26,886,661 – 26,886,753 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 90.32 |
| Mean single sequence MFE | -31.74 |
| Consensus MFE | -25.30 |
| Energy contribution | -27.90 |
| Covariance contribution | 2.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.80 |
| SVM decision value | 1.32 |
| SVM RNA-class probability | 0.941839 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 26886661 92 - 27905053 GAAAAAGCGGCGGUUACCACGCGCAGGAAAACUCGAAUUUCCCAACAGCAUUUCAGUG-CUCAUUGAGAGUGCGAGUGUGUGAGUGAGUGCCC ........((((.(((((((((((.(((((.......))))).....((((((((((.-...))))))).)))..))))))).)))).)))). ( -34.60) >DroSec_CAF1 94899 76 - 1 GGAAAAGCGGCGGUUACCACGCGCAGGAAAACUCGAAUUUCCCAGCAGCAUUUCAGUGUCUCAUUGAGA--------------GUGAGUG--- .((((.((.(((.......)))((.(((((.......)))))..)).)).)))).....(((((.....--------------)))))..--- ( -18.40) >DroSim_CAF1 98572 93 - 1 GGAAAAGCGGCGGUUACCACGCGCAGGAAAACUCGAAUUUCCCAGCAGCAUUUCAGUGUCUCAUUGAGCGUGCGAGUGUGUGAGUGAGUGCCC ........((((.(((((((((((.(((((.......))))).....(((((((((((...))))))).))))..))))))).)))).)))). ( -34.90) >DroEre_CAF1 103479 93 - 1 GGAAAAGCGGCGGUUACCACGCGCAGGAAAACUCGAAUUUCCCAGCAGCAUUUCAGUGUCUCAUUGAGAGUGCGAGUGUGUGAGUGAGUGCCC ........((((.(((((((((((.(((((.......))))).....(((((((((((...)))))))).)))..))))))).)))).)))). ( -35.40) >DroYak_CAF1 109632 93 - 1 GGAAAAACGGCGGUUACCACGCGCAGGAAAACUCGAAUUUCCCAACAGCAUUUCAGUGUCUCAUUGAGAGUGCGAGUGUGUGAGUGAGUGCCC ........((((.(((((((((((.(((((.......))))).....(((((((((((...)))))))).)))..))))))).)))).)))). ( -35.40) >consensus GGAAAAGCGGCGGUUACCACGCGCAGGAAAACUCGAAUUUCCCAGCAGCAUUUCAGUGUCUCAUUGAGAGUGCGAGUGUGUGAGUGAGUGCCC ........((((.(((((((((((.(((((.......))))).....(((((((((((...)))))))).)))..))))))).)))).)))). (-25.30 = -27.90 + 2.60)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:47:27 2006