
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 26,798,134 – 26,798,230 |
| Length | 96 |
| Max. P | 0.938672 |

| Location | 26,798,134 – 26,798,230 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 84.89 |
| Mean single sequence MFE | -27.69 |
| Consensus MFE | -19.01 |
| Energy contribution | -19.21 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.64 |
| Structure conservation index | 0.69 |
| SVM decision value | 1.29 |
| SVM RNA-class probability | 0.938672 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 26798134 96 + 27905053 GCCCCCACCACCGCCUACUCUUCCGCCCACCAGCACUUGACGCUCUUCGAAAUGCGAAACAAAUAAACAAAGGGCAGCUAAAAGGCGGUGGCGCCU .....(.(((((((((........((((...(((.......))).((((.....)))).............)))).......))))))))).)... ( -29.16) >DroSec_CAF1 6905 96 + 1 GCCCCCGCCACCGCCUACUCUUCCGCCCACCAGCACUUGACGCUCUUCAAAAUGCGAAACAAAUAAACAAAGGGCAGCUAAAAGGCGGUGGCGCCU .....(((((((((((........((((..(((...))).(((..........)))...............)))).......)))))))))))... ( -32.56) >DroSim_CAF1 6891 96 + 1 GCCCCCGCCACCGCCUACUCUUCCGCCCACCAGCACUUGACGCUCUUCGAAAUGCGAAACAAAUAAACAAAGGGCAGCUAAAAGGCGGUGGCGCCU .....(((((((((((........((((...(((.......))).((((.....)))).............)))).......)))))))))))... ( -34.56) >DroEre_CAF1 9012 89 + 1 GCCUACA---CAUCCAAC----CAGCCCACCAGCACUUGACGCUCUCCGCAAUGCGAAACAAAUAAACAAAGGGCAGCUAAAAGGCGGUGGCGCCU .......---........----..(((((((.((.(((...(((.(((.......................))).)))...)))))))))).)).. ( -21.10) >DroYak_CAF1 9198 74 + 1 ----------------------ACGCCCACCAGCACUUGACGCUCUUCGGAAUGCGAAACAAAUAAACAAAGGGCAGCUAAAAGGCGGUGGCGCCU ----------------------..(((((((.((.(((..(((((....))..))).............))).)).(((....)))))))).)).. ( -21.06) >consensus GCCCCCACCACCGCCUACUCUUCCGCCCACCAGCACUUGACGCUCUUCGAAAUGCGAAACAAAUAAACAAAGGGCAGCUAAAAGGCGGUGGCGCCU ........................(((((((..........((((((......................)))))).(((....)))))))).)).. (-19.01 = -19.21 + 0.20)

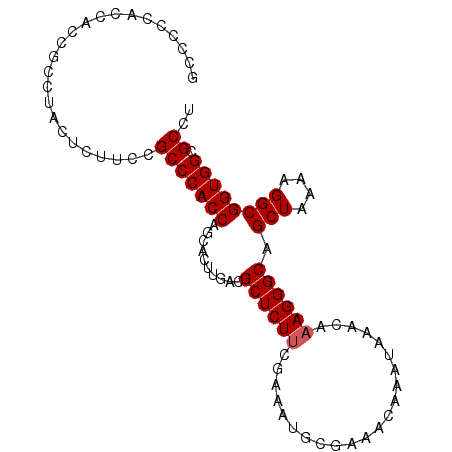

| Location | 26,798,134 – 26,798,230 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 84.89 |
| Mean single sequence MFE | -35.48 |
| Consensus MFE | -26.56 |
| Energy contribution | -30.20 |
| Covariance contribution | 3.64 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.77 |
| SVM RNA-class probability | 0.846589 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
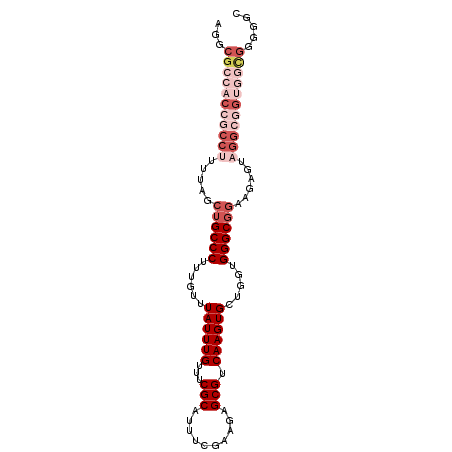
>3R_DroMel_CAF1 26798134 96 - 27905053 AGGCGCCACCGCCUUUUAGCUGCCCUUUGUUUAUUUGUUUCGCAUUUCGAAGAGCGUCAAGUGCUGGUGGGCGGAAGAGUAGGCGGUGGUGGGGGC ...((((((((((((((..((((((......((((((...(((..........))).)))))).....))))))..))).)))))))))))..... ( -42.10) >DroSec_CAF1 6905 96 - 1 AGGCGCCACCGCCUUUUAGCUGCCCUUUGUUUAUUUGUUUCGCAUUUUGAAGAGCGUCAAGUGCUGGUGGGCGGAAGAGUAGGCGGUGGCGGGGGC ...((((((((((((((..((((((......((((((...(((..........))).)))))).....))))))..))).)))))))))))..... ( -44.00) >DroSim_CAF1 6891 96 - 1 AGGCGCCACCGCCUUUUAGCUGCCCUUUGUUUAUUUGUUUCGCAUUUCGAAGAGCGUCAAGUGCUGGUGGGCGGAAGAGUAGGCGGUGGCGGGGGC ...((((((((((((((..((((((......((((((...(((..........))).)))))).....))))))..))).)))))))))))..... ( -44.00) >DroEre_CAF1 9012 89 - 1 AGGCGCCACCGCCUUUUAGCUGCCCUUUGUUUAUUUGUUUCGCAUUGCGGAGAGCGUCAAGUGCUGGUGGGCUG----GUUGGAUG---UGUAGGC ....(((..(((..(((((((((((............((((((...))))))((((.....))))...)))).)----)))))).)---))..))) ( -26.60) >DroYak_CAF1 9198 74 - 1 AGGCGCCACCGCCUUUUAGCUGCCCUUUGUUUAUUUGUUUCGCAUUCCGAAGAGCGUCAAGUGCUGGUGGGCGU---------------------- ..((.((((((((((...(((...(((((......((.....))...))))))))...))).)).)))))))..---------------------- ( -20.70) >consensus AGGCGCCACCGCCUUUUAGCUGCCCUUUGUUUAUUUGUUUCGCAUUUCGAAGAGCGUCAAGUGCUGGUGGGCGGAAGAGUAGGCGGUGGCGGGGGC ...(((((((((((.....((((((......((((((...(((..........))).)))))).....))))))......)))))))))))..... (-26.56 = -30.20 + 3.64)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:47:03 2006