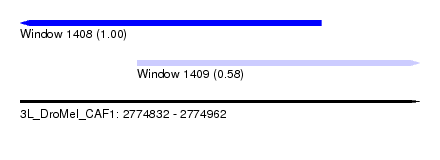
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 2,774,832 – 2,774,962 |
| Length | 130 |
| Max. P | 0.999479 |

| Location | 2,774,832 – 2,774,930 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 78.68 |
| Mean single sequence MFE | -32.18 |
| Consensus MFE | -20.31 |
| Energy contribution | -20.31 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.25 |
| Mean z-score | -3.54 |
| Structure conservation index | 0.63 |
| SVM decision value | 3.64 |
| SVM RNA-class probability | 0.999479 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 2774832 98 - 23771897 CCGGGCACUCGGAAGGGAAAUCGAUCGAAACCUAA---GG-AUCUCGCUCGAUUU-UUGUUCUGAGUGUACUUA-UUCUAGUGCCUUGCUAAUAUGAACCUCCA- ..((((((((((((.(((((((((.(((..((...---))-...))).)))))))-)).))))))))))..(((-(..((((.....))))..))))....)).- ( -34.40) >DroSec_CAF1 19990 99 - 1 CCGGUCACUCGGAAAGGAAGUCGAUUGAAAACAUACCUCG-UUCUCGCUCGAUUUUUUGUUCUGAGUGUACUUUUUUCUAGUGCCUUGCUAAUA----CCUCCA- ..((((((((((((((((((((((.(((.(((.......)-)).))).)))))))))).)))))))))..........((((.....))))..)----))....- ( -28.60) >DroSim_CAF1 10538 97 - 1 CCGGGCACUCGGAAAGGAAGUCGAUUGAAAACAUACCUCG-UUCUCGCUCGAUUU-UUGUUCUGAGUGUACUUA-UUCAAGUGCCUUGCUAAUA----CCUCCA- ...(((((((((((..((((((((.(((.(((.......)-)).))).)))))))-)..))))))))((((((.-...))))))...)))....----......- ( -31.40) >DroEre_CAF1 21772 97 - 1 CCAGGCACUCGGAA-GGAAAUCGACCGAGAUCCCA---GA-AUCCAGAUCGAUUU-UUGUUCCGGGUGUACAUA-UCCUAGUGCCUUGCUAAUAUGUACCCGCA- ....((((((((((-(((((((((.(..(((....---..-)))..).)))))))-)).))))))))(((((((-(..((((.....))))))))))))..)).- ( -32.80) >DroYak_CAF1 19404 100 - 1 CCAGGCACUCGUAACGGAAAUCGAUCAGGAUCCUA---GUUCUCCAGUUCGAUUU-UUGUUCUGAGUGUACAUA-UUCUAGUGCCUUACUAAUAUGUACCUGCAU .((((((((((.((((((((((((...(((.....---....)))...)))))))-))))).))))))((((((-(..(((((...))))))))))))))))... ( -33.70) >consensus CCGGGCACUCGGAAAGGAAAUCGAUCGAAAACAUA___GG_AUCUCGCUCGAUUU_UUGUUCUGAGUGUACUUA_UUCUAGUGCCUUGCUAAUAUG_ACCUCCA_ ....((((((((((...(((((((.(((................))).)))))))....)))))))))).........((((.....)))).............. (-20.31 = -20.31 + 0.00)

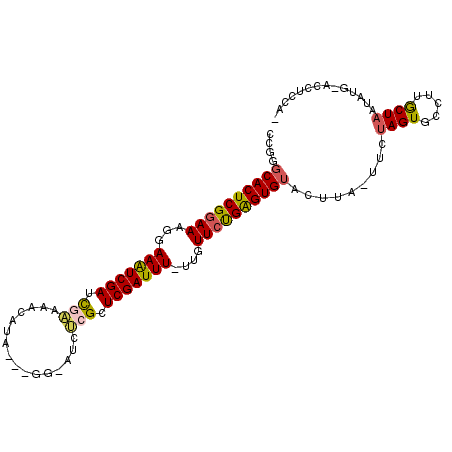
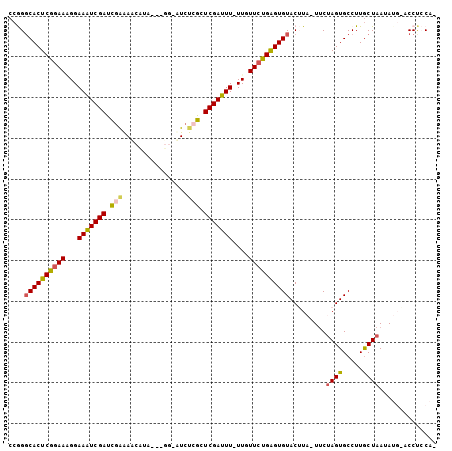
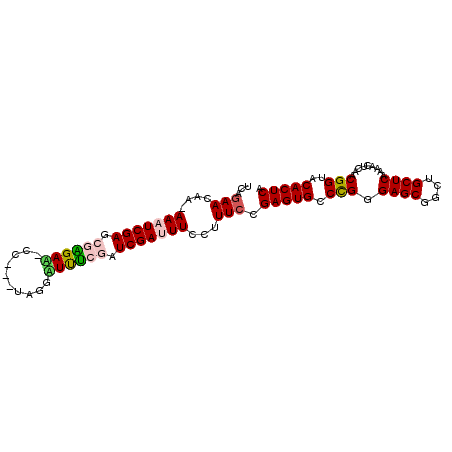
| Location | 2,774,870 – 2,774,962 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 83.35 |
| Mean single sequence MFE | -26.96 |
| Consensus MFE | -20.24 |
| Energy contribution | -20.60 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.579102 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 2774870 92 + 23771897 UCAGAACAA-AAAUCGAGCGAGAU-CC---UUAGGUUUCGAUCGAUUUCCCUUCCGAGUGCCCGGGAGCGGCUGCUCAAAACUCAACGGUACACUCA ...(((...-(((((((.((((((-(.---...))))))).)))))))...))).(((((.(((.((((....)))).........)))..))))). ( -29.00) >DroSec_CAF1 20025 96 + 1 UCAGAACAAAAAAUCGAGCGAGAA-CGAGGUAUGUUUUCAAUCGACUUCCUUUCCGAGUGACCGGGAGCGGCUGCUCAAAACUCAACGGUACACUCA .............((((..(((((-((.....)))))))..))))..........(((((((((.((((....)))).........)))).))))). ( -26.80) >DroSim_CAF1 10572 95 + 1 UCAGAACAA-AAAUCGAGCGAGAA-CGAGGUAUGUUUUCAAUCGACUUCCUUUCCGAGUGCCCGGGAGCGGCUGCUCAAAACUCAACGGUACACUCA .........-...((((..(((((-((.....)))))))..))))..........(((((.(((.((((....)))).........)))..))))). ( -23.50) >DroEre_CAF1 21810 91 + 1 CCGGAACAA-AAAUCGAUCUGGAU-UC---UGGGAUCUCGGUCGAUUUCC-UUCCGAGUGCCUGGGAGCUGUUGCUCAAAACUCAACGGUACACUCA .(((((...-(((((((((.((((-(.---...))))).)))))))))..-))))).((((((((((((....)))).....)))..)))))..... ( -32.60) >DroYak_CAF1 19443 93 + 1 UCAGAACAA-AAAUCGAACUGGAGAAC---UAGGAUCCUGAUCGAUUUCCGUUACGAGUGCCUGGGAGCGGCUGCUCAAAACUCAACGGUACACUCA ...((....-(((((((.(.(((....---.....))).).)))))))(((((..((((......((((....))))...))))))))).....)). ( -22.90) >consensus UCAGAACAA_AAAUCGAGCGAGAA_CC___UAGGAUUUCGAUCGAUUUCCUUUCCGAGUGCCCGGGAGCGGCUGCUCAAAACUCAACGGUACACUCA ...(((....(((((((.((((((..........)))))).)))))))...))).(((((.(((.((((....)))).........)))..))))). (-20.24 = -20.60 + 0.36)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:54:20 2006