
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 38,109 – 38,213 |
| Length | 104 |
| Max. P | 0.997828 |

| Location | 38,109 – 38,213 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 79.61 |
| Mean single sequence MFE | -38.17 |
| Consensus MFE | -23.46 |
| Energy contribution | -24.34 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.08 |
| Mean z-score | -3.20 |
| Structure conservation index | 0.61 |
| SVM decision value | 2.94 |
| SVM RNA-class probability | 0.997828 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 38109 104 + 23771897 AGUGAUGCGAGUGCAAGUGAGAAUGCGAGUGAUAGUGGCUGUAGCAGGAGC---UAUCGUGCAGACAGUAGGAGUAGCAGCCGCAGUCGCACAUAUUUU-UGCCUUGU------------ ......(((((.((((....(((((.(.(((((.(((((((((((....))---).((.(((.....))).))...)))))))).))))).).))))))-))))))))------------ ( -36.10) >DroSec_CAF1 27648 104 + 1 AGUGAUGCGAGUGCAAAUGAGAAUGCGAGUGAUAGUGGCUGUAGCAGGAGC---UGUCAUGCAGACAGUAGGAGUAGCAGCCGCAGUCGCACAUAUUUU-CGACGUGU------------ .((..(((....)))..((((((((.(.(((((.((((((((.((....((---((((.....))))))....)).)))))))).))))).).))))))-))))....------------ ( -38.40) >DroSim_CAF1 29251 104 + 1 AGUGAUGCGAGUGCAAAUGAGAAUGCGAGUGAUAGUGGCUGUAGCAGGAGC---UGUCAUGCAGACAGUAGGAGUAGCAGCCGCAGUCGCACAUAUUUU-CCACGUGU------------ .(((.(((....)))...(((((((.(.(((((.((((((((.((....((---((((.....))))))....)).)))))))).))))).).))))))-))))....------------ ( -40.10) >DroWil_CAF1 103234 114 + 1 AAUGAUGCGAAUGCGAAUGGC------UAUGAUAGUGGCUAUAACAGGAGCUGCUGUCAUGCAGACAAUAUGAAUAGCAGCCGCAGUCGCACAUUUUUCUCGAUAAGUUGAUUUCAAUGG ((((.(((((.((((.(((((------(((....)))))))).......((((((((((((.......)))).)))))))))))).)))))))))...........((((....)))).. ( -38.10) >consensus AGUGAUGCGAGUGCAAAUGAGAAUGCGAGUGAUAGUGGCUGUAGCAGGAGC___UGUCAUGCAGACAGUAGGAGUAGCAGCCGCAGUCGCACAUAUUUU_CGACGUGU____________ .(((.(((((.(((..........((........))((((((.((.........((((.....))))......)).))))))))).)))))))).......................... (-23.46 = -24.34 + 0.88)

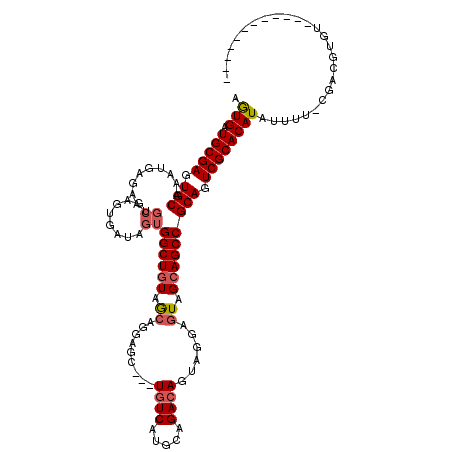
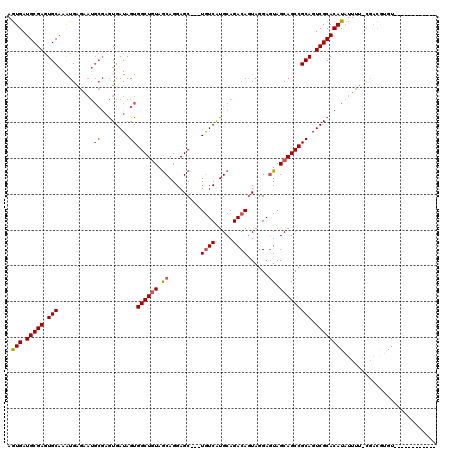
| Location | 38,109 – 38,213 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 79.61 |
| Mean single sequence MFE | -25.83 |
| Consensus MFE | -20.66 |
| Energy contribution | -20.35 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.03 |
| Structure conservation index | 0.80 |
| SVM decision value | 1.42 |
| SVM RNA-class probability | 0.952046 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 38109 104 - 23771897 ------------ACAAGGCA-AAAAUAUGUGCGACUGCGGCUGCUACUCCUACUGUCUGCACGAUA---GCUCCUGCUACAGCCACUAUCACUCGCAUUCUCACUUGCACUCGCAUCACU ------------.....(((-(......((((((.((.(((((.........(((((.....))))---).........))))).....)).))))))......))))............ ( -23.27) >DroSec_CAF1 27648 104 - 1 ------------ACACGUCG-AAAAUAUGUGCGACUGCGGCUGCUACUCCUACUGUCUGCAUGACA---GCUCCUGCUACAGCCACUAUCACUCGCAUUCUCAUUUGCACUCGCAUCACU ------------........-.......((((((....(((((.........(((((.....))))---).........)))))..........(((........)))..)))))).... ( -24.97) >DroSim_CAF1 29251 104 - 1 ------------ACACGUGG-AAAAUAUGUGCGACUGCGGCUGCUACUCCUACUGUCUGCAUGACA---GCUCCUGCUACAGCCACUAUCACUCGCAUUCUCAUUUGCACUCGCAUCACU ------------.((((((.-....)))))).((.((((((((.........(((((.....))))---).........)))))..........(((........)))....)))))... ( -26.87) >DroWil_CAF1 103234 114 - 1 CCAUUGAAAUCAACUUAUCGAGAAAAAUGUGCGACUGCGGCUGCUAUUCAUAUUGUCUGCAUGACAGCAGCUCCUGUUAUAGCCACUAUCAUA------GCCAUUCGCAUUCGCAUCAUU .........................(((((((((.(((((..(((((..(((.((...(((((((((......))))))).)))).))).)))------))...))))).))))).)))) ( -28.20) >consensus ____________ACACGUCG_AAAAUAUGUGCGACUGCGGCUGCUACUCCUACUGUCUGCAUGACA___GCUCCUGCUACAGCCACUAUCACUCGCAUUCUCAUUUGCACUCGCAUCACU ............................((((((.((((((((..........((((.....))))...((....))..)))))..........(......)....))).)))))).... (-20.66 = -20.35 + -0.31)

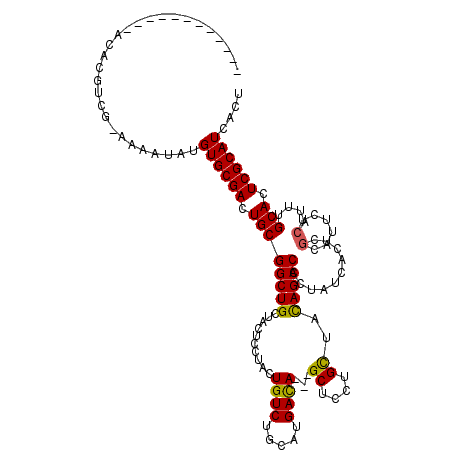
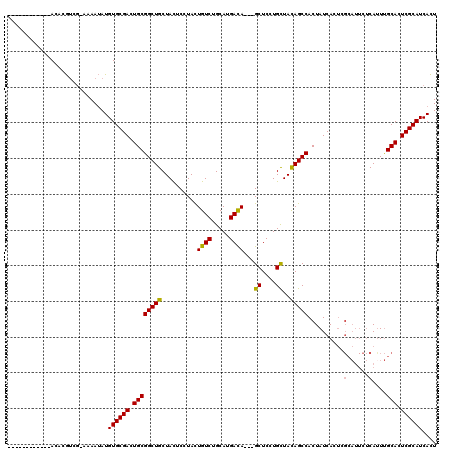
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:29:48 2006