
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 22,530,490 – 22,530,678 |
| Length | 188 |
| Max. P | 0.865936 |

| Location | 22,530,490 – 22,530,604 |
|---|---|
| Length | 114 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.89 |
| Mean single sequence MFE | -27.48 |
| Consensus MFE | -22.23 |
| Energy contribution | -23.35 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.56 |
| SVM RNA-class probability | 0.780956 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22530490 114 + 23771897 GAAAGU------AAAGACCAAAAGCAAAUAUAUUAACGUAGAACUCCAACGGAAGGGUAUCUUAGCCAAAUGCUGAUGUGCUCCUUUAAAUAAUUCGCUGGACAAAAUUGCAGUCUUAUG ......------.(((((.....((((.(((......)))....((((.(((((((((((.(((((.....))))).)))).)))........)))).)))).....)))).)))))... ( -22.10) >DroSec_CAF1 13923 114 + 1 AAAAGU------AAAGACCAAAAAGCAAUAUAUUAACGUGGAUCUCCCACGGAAGGGUAUCUUAGCCAAAUGCUGAUGUGCUCCUUUAAAUAAUUCGCUGGACAAAAUUGCAGUCUUAUG ......------.(((((......(((((.......(((((.....)))))(((((((((.(((((.....))))).)))).)))))...................))))).)))))... ( -28.40) >DroSim_CAF1 14814 114 + 1 AAAAGU------AAAGACCAAGAAGCAAUAUAUUAACGUGGAUCUCCCACGGAAGGGUAUCUUAGCCAAAUGCUGAUGUGCUCCUUUAAAUAAUUCGCUGGACAAAAUUGCAGUCUUAUG ......------.......((((.(((((.......(((((.....)))))(((((((((.(((((.....))))).)))).)))))...................)))))..))))... ( -28.50) >DroEre_CAF1 12915 120 + 1 UAAAGUUGAGCUUAAGAGCAAAAAGCGAUGUAUUUACAAGGAUCUACCACGGAAGGGUAUCUCAGCCAAAUGCUGAUGUGCUCCUUUAAAUAAUUCGCUGGACAAAAUUGCAGUCUUAUG .........(((....)))....(((((..((((((.(((((..((((.(....)))))..(((((.....))))).....)))))))))))..)))))((((.........)))).... ( -30.90) >consensus AAAAGU______AAAGACCAAAAAGCAAUAUAUUAACGUGGAUCUCCCACGGAAGGGUAUCUUAGCCAAAUGCUGAUGUGCUCCUUUAAAUAAUUCGCUGGACAAAAUUGCAGUCUUAUG .............(((((......(((((.......(((((.....)))))(((((((((.(((((.....))))).)))).)))))...................))))).)))))... (-22.23 = -23.35 + 1.12)

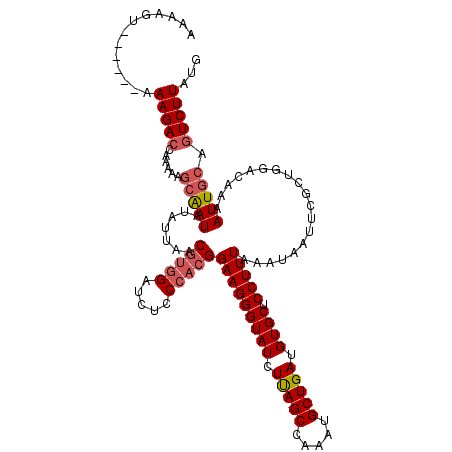

| Location | 22,530,524 – 22,530,644 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.92 |
| Mean single sequence MFE | -36.38 |
| Consensus MFE | -35.25 |
| Energy contribution | -35.38 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.85 |
| SVM RNA-class probability | 0.865936 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22530524 120 + 23771897 GAACUCCAACGGAAGGGUAUCUUAGCCAAAUGCUGAUGUGCUCCUUUAAAUAAUUCGCUGGACAAAAUUGCAGUCUUAUGGGAAAAUUCCCCACAUGCCGCAGACUUUGCGGUAUUGGUC ....(((....(((((((((.(((((.....))))).)))).)))))........((..((((.........))))..))))).......(((.(((((((((...)))))))))))).. ( -33.60) >DroSec_CAF1 13957 120 + 1 GAUCUCCCACGGAAGGGUAUCUUAGCCAAAUGCUGAUGUGCUCCUUUAAAUAAUUCGCUGGACAAAAUUGCAGUCUUAUGGGAAAAUUCCCCACAUGCCGCAGACUUUGCGGUAUUGGUC ....(((((..(((((((((.(((((.....))))).)))).))))).........((((.(......).))))....))))).......(((.(((((((((...)))))))))))).. ( -37.10) >DroSim_CAF1 14848 120 + 1 GAUCUCCCACGGAAGGGUAUCUUAGCCAAAUGCUGAUGUGCUCCUUUAAAUAAUUCGCUGGACAAAAUUGCAGUCUUAUGGGAAAAUUCCCCACAUGCCGCAGACUUUGCGGUAUUGGUC ....(((((..(((((((((.(((((.....))))).)))).))))).........((((.(......).))))....))))).......(((.(((((((((...)))))))))))).. ( -37.10) >DroEre_CAF1 12955 120 + 1 GAUCUACCACGGAAGGGUAUCUCAGCCAAAUGCUGAUGUGCUCCUUUAAAUAAUUCGCUGGACAAAAUUGCAGUCUUAUGGGAAAAUUCCCCACAUGCCGCAGACUCUGCGGUAUUGGUC ..........((((((((((.(((((.....))))).))))))..........(((.(.((((.........))))...).)))..))))(((.(((((((((...)))))))))))).. ( -37.70) >consensus GAUCUCCCACGGAAGGGUAUCUUAGCCAAAUGCUGAUGUGCUCCUUUAAAUAAUUCGCUGGACAAAAUUGCAGUCUUAUGGGAAAAUUCCCCACAUGCCGCAGACUUUGCGGUAUUGGUC ....(((((..(((((((((.(((((.....))))).)))).))))).........((((.(......).))))....))))).......(((.(((((((((...)))))))))))).. (-35.25 = -35.38 + 0.12)



| Location | 22,530,564 – 22,530,678 |
|---|---|
| Length | 114 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.65 |
| Mean single sequence MFE | -32.18 |
| Consensus MFE | -23.06 |
| Energy contribution | -25.50 |
| Covariance contribution | 2.44 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.49 |
| SVM RNA-class probability | 0.757203 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22530564 114 - 23771897 C-AACAGC-----CCCAUUACAAGUUUUUCGCUUUCGGUCGACCAAUACCGCAAAGUCUGCGGCAUGUGGGGAAUUUUCCCAUAAGACUGCAAUUUUGUCCAGCGAAUUAUUUAAAGGAG .-......-----.((.(((..(((..((((((...(((........)))(((((((.((((((.((((((((...)))))))).).))))))))))))..))))))..)))))).)).. ( -32.20) >DroPse_CAF1 37012 103 - 1 CGAGCAGCUACUGCUCAUCAGAAGUGCUCUGUUUUUGGGCUACCAAUACCGCAAUGUCUGCGGCAUGUGGAGAAUUUUCCCAUAAGACUGCAAUUUUAUCCUU----------------- .((((((...))))))...((((.(((...(((((((((((.(((...(((((.....)))))....)))))......)))).))))).))).))))......----------------- ( -30.50) >DroSec_CAF1 13997 114 - 1 C-AACAGC-----CCCAUUACAAGUUUUUCGCUUUCGGUCGACCAAUACCGCAAAGUCUGCGGCAUGUGGGGAAUUUUCCCAUAAGACUGCAAUUUUGUCCAGCGAAUUAUUUAAAGGAG .-......-----.((.(((..(((..((((((...(((........)))(((((((.((((((.((((((((...)))))))).).))))))))))))..))))))..)))))).)).. ( -32.20) >DroSim_CAF1 14888 114 - 1 C-AACAGC-----CCCAUUACAAGUUUUUCGCUUUCGGUCGACCAAUACCGCAAAGUCUGCGGCAUGUGGGGAAUUUUCCCAUAAGACUGCAAUUUUGUCCAGCGAAUUAUUUAAAGGAG .-......-----.((.(((..(((..((((((...(((........)))(((((((.((((((.((((((((...)))))))).).))))))))))))..))))))..)))))).)).. ( -32.20) >DroEre_CAF1 12995 114 - 1 C-AACAGC-----CCCAUUACAAGUUUUUCGCUUUGGGUCGACCAAUACCGCAGAGUCUGCGGCAUGUGGGGAAUUUUCCCAUAAGACUGCAAUUUUGUCCAGCGAAUUAUUUAAAGGAG .-......-----.((.(((..(((..((((((((((.....))))....(((((((.((((((.((((((((...)))))))).).))))))))))))..))))))..)))))).)).. ( -33.80) >consensus C_AACAGC_____CCCAUUACAAGUUUUUCGCUUUCGGUCGACCAAUACCGCAAAGUCUGCGGCAUGUGGGGAAUUUUCCCAUAAGACUGCAAUUUUGUCCAGCGAAUUAUUUAAAGGAG .....................((((..((((((...((....))......(((((((.((((((.((((((((...)))))))).).))))))))))))..))))))..))))....... (-23.06 = -25.50 + 2.44)

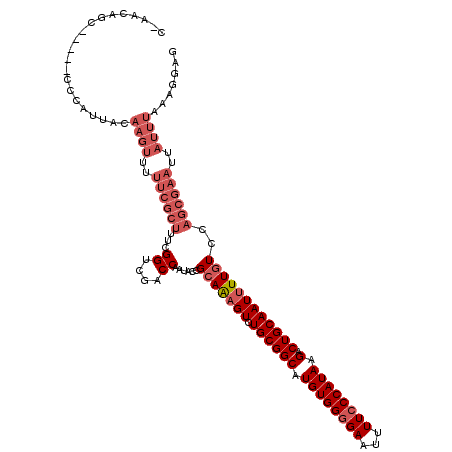
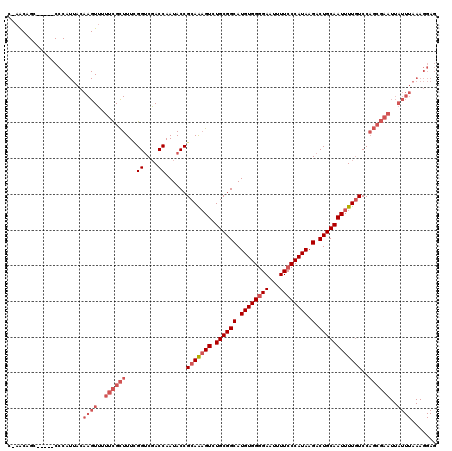
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:22:20 2006