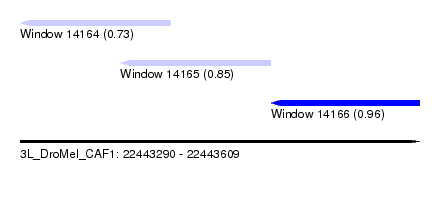
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 22,443,290 – 22,443,609 |
| Length | 319 |
| Max. P | 0.963137 |

| Location | 22,443,290 – 22,443,410 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.75 |
| Mean single sequence MFE | -35.52 |
| Consensus MFE | -34.45 |
| Energy contribution | -34.70 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.725345 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22443290 120 - 23771897 UAAAAAUAAUGAGUUUGCCGCUUCCUCUUUGGCAGCGGCUCCUGCGGAAGCGGCUUGAAAACCCGAUUAAAAAUUGAGCAGCGAGCAAGGACGUUAAGUAGAAAUGUCAAUAUGACAGCU ......(((((.....((((((.((.....)).))))))((((((....)).(((((....(.((((.....)))).)...))))).)))))))))........((((.....))))... ( -35.50) >DroSec_CAF1 14990 120 - 1 UAAAAAUAAUGAGUUUGCCGCUUCCUCUUUGGCAGCGGCUCCUGCGGAAGCGGCUUGAAAACCCGAUUAAAAAUUGAGCAGCGAGCAAGGACGUUAAGUAGAAAUGUCAAUAUGACAGCU ......(((((.....((((((.((.....)).))))))((((((....)).(((((....(.((((.....)))).)...))))).)))))))))........((((.....))))... ( -35.50) >DroSim_CAF1 19571 120 - 1 UAAAAAUAAUGAGUUUGCCGCUUCCUCUUUGGCAGCGGCUCCUGCGGAAGCGGCUUGAAAACCCGAUUAAAAAUUGAGCAGCGAGCAAGGACGUUAAGUAGAAAUGUCAAUAUGACAGCU ......(((((.....((((((.((.....)).))))))((((((....)).(((((....(.((((.....)))).)...))))).)))))))))........((((.....))))... ( -35.50) >DroEre_CAF1 21849 120 - 1 UAAAAAUAAUGAGUUUGCCGCUUCCUCUUUGGACGCGGCUCCUGCGGAAGCGGCUUGGAAACCCGAUUAAAAAUUGAGCAGCGAGCAAGGACGUUAAGUAGAAAUGUCAAUAUGACAGCU ............(((((((((.(((.....))).)))((((((((....)))).((((....)))).........)))).))))))...((((((.......))))))............ ( -35.60) >consensus UAAAAAUAAUGAGUUUGCCGCUUCCUCUUUGGCAGCGGCUCCUGCGGAAGCGGCUUGAAAACCCGAUUAAAAAUUGAGCAGCGAGCAAGGACGUUAAGUAGAAAUGUCAAUAUGACAGCU ......(((((.....((((((.((.....)).))))))((((((....)).(((((....(.((((.....)))).)...))))).)))))))))........((((.....))))... (-34.45 = -34.70 + 0.25)

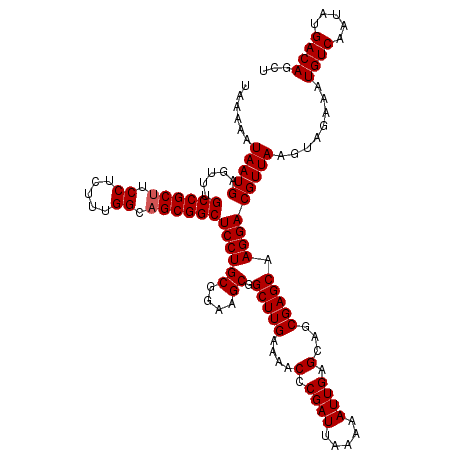
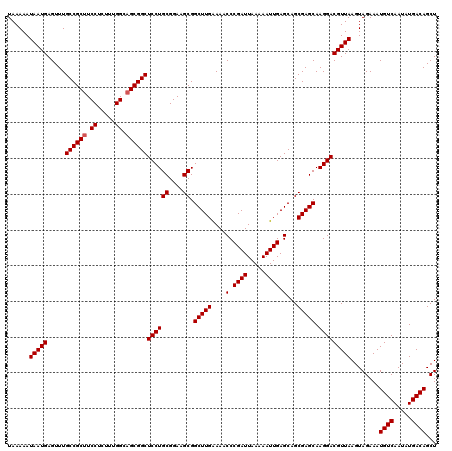
| Location | 22,443,370 – 22,443,490 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 97.64 |
| Mean single sequence MFE | -27.97 |
| Consensus MFE | -26.35 |
| Energy contribution | -26.60 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.849695 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22443370 120 - 23771897 UAAAUAUGAAAAUUAAAAUUACAAAAUGCACAAAUUGCAGCGACGUGGAGCGAGGUGAAGUGCAGCGAAAAUCAUAUUAAUAAAAAUAAUGAGUUUGCCGCUUCCUCUUUGGCAGCGGCU ..(((((((.(((....)))......(((((...(..(.((........))...)..).))))).......)))))))..................((((((.((.....)).)))))). ( -27.40) >DroSec_CAF1 15070 120 - 1 UAAAUAUGAAAAUUAAAAUUACAAAAUGCACAAAUUGCAGCGACGUGAAGCGAGGUGAAGUGCAGUGAAAAUCAUAUUAAUAAAAAUAAUGAGUUUGCCGCUUCCUCUUUGGCAGCGGCU ..(((((((.(((....))).((...(((((...(..(.((........))...)..).))))).))....)))))))..................((((((.((.....)).)))))). ( -28.60) >DroSim_CAF1 19651 120 - 1 UAAAUAUGAAAAUUAAAAUUACAAAAUGCACAAAUUGCAGCGACGUGAAGCGAGGUGAAGUGCAGUGAAAAUCAUAUUAAUAAAAAUAAUGAGUUUGCCGCUUCCUCUUUGGCAGCGGCU ..(((((((.(((....))).((...(((((...(..(.((........))...)..).))))).))....)))))))..................((((((.((.....)).)))))). ( -28.60) >DroEre_CAF1 21929 120 - 1 UAAAUAUGAAAAUUAAAAUUACAAAAUGCACAAAUUGCAGCGACGUGGAGCGAAGUGAAGUGCAGCGAAAAUCAUAUUAAUAAAAAUAAUGAGUUUGCCGCUUCCUCUUUGGACGCGGCU ..(((((((.................((((.....))))....(((...((........))...)))....)))))))..................(((((.(((.....))).))))). ( -27.30) >consensus UAAAUAUGAAAAUUAAAAUUACAAAAUGCACAAAUUGCAGCGACGUGAAGCGAGGUGAAGUGCAGCGAAAAUCAUAUUAAUAAAAAUAAUGAGUUUGCCGCUUCCUCUUUGGCAGCGGCU ..(((((((.(((....)))......(((((...(..(.((........))...)..).))))).......)))))))..................((((((.((.....)).)))))). (-26.35 = -26.60 + 0.25)

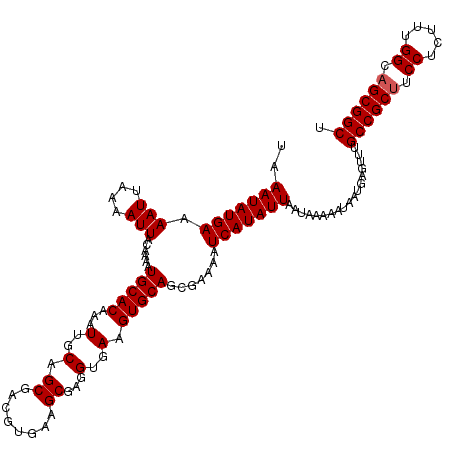

| Location | 22,443,490 – 22,443,609 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 97.08 |
| Mean single sequence MFE | -35.63 |
| Consensus MFE | -34.00 |
| Energy contribution | -34.25 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.48 |
| Structure conservation index | 0.95 |
| SVM decision value | 1.55 |
| SVM RNA-class probability | 0.963137 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
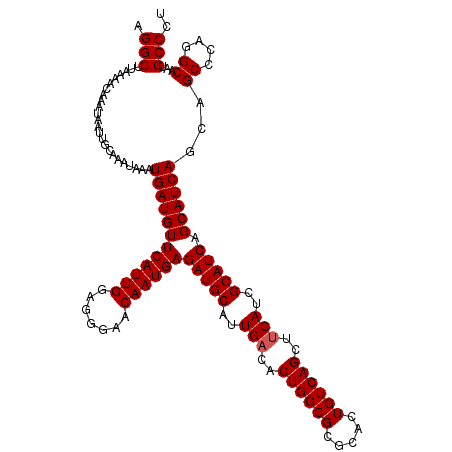
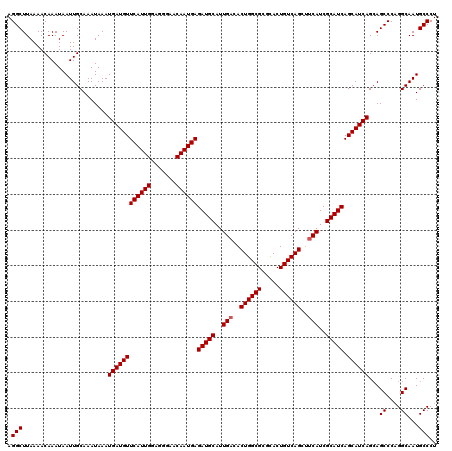
>3L_DroMel_CAF1 22443490 119 - 23771897 AGGCUUAAAACAAAUAAUUGCAAAUAAAUGAUGUUCAUUGGAGGGAACAAUGAGAUGCAUUGACACUGGCGCAUACUGUCAGCUUCAUCGCAUCAGCAUCAGGAGCCC-GGCAAUGCCCU .(((((....(((....)))........((((((((((((.......))))))(((((..(((..((((((.....))))))..)))..))))).)))))).))))).-(((...))).. ( -36.20) >DroSec_CAF1 15190 120 - 1 AGGCUUAAAACAAAUAAUUGCAAAUAAAUGAUGUUCAUUGGAGGGAACAAUGAGAUGCAUUGACACUGGCGCGCACUGUCAGCUUCAUCGCAUCAGCAUCAGCAGCCCAGGCAAUGCCCU .(((..............(((.......((((((((((((.......))))))(((((..(((..((((((.....))))))..)))..))))).)))))))))((....))...))).. ( -35.61) >DroSim_CAF1 19771 120 - 1 AGGCUUAAAACAAAUAAUUGCAAAUAAAUGAUGUUCAUUGGAGAGAACAAUGAGAUGCAUUGACACUGGCGCGCACUGUCAGCUUCAUCGCAUCAGCAUCAGCAGCCCAGGCAAUGCCCU .(((..............(((.......((((((((((((.......))))))(((((..(((..((((((.....))))))..)))..))))).)))))))))((....))...))).. ( -35.61) >DroEre_CAF1 22049 120 - 1 AGGCUUAAAACAAAUAAUUGCAAAUAAAUGAUGUUCAUUGGAGGGAACAAUGAGAUGCAUUGACACUGGCGCGCACUGUCAGCCGCAUCGCAUCAGCAUCAGCAGCCCAGGCAAUGCCCU .(((..............(((.......((((((((((((.......))))))(((((.((((((.((.....)).))))))..)))))))))))......)))((....))...))).. ( -35.12) >consensus AGGCUUAAAACAAAUAAUUGCAAAUAAAUGAUGUUCAUUGGAGGGAACAAUGAGAUGCAUUGACACUGGCGCGCACUGUCAGCUUCAUCGCAUCAGCAUCAGCAGCCCAGGCAAUGCCCU .(((........................((((((((((((.......))))))(((((..(((..((((((.....))))))..)))..))))).))))))...((....))...))).. (-34.00 = -34.25 + 0.25)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:21:13 2006