
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 22,426,221 – 22,426,421 |
| Length | 200 |
| Max. P | 0.862506 |

| Location | 22,426,221 – 22,426,341 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 81.23 |
| Mean single sequence MFE | -29.23 |
| Consensus MFE | -23.64 |
| Energy contribution | -23.87 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.82 |
| SVM RNA-class probability | 0.859027 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22426221 120 - 23771897 UUCCAGUCGGCCAGGAGAGUUUCAGGGAAUGGCAGAACAAGUCUACUAACUGGUUGUCGGUUUUUAGUUUAAGCACACUAAAUCAAUUCCCAGGCAUUGCUCUUUAAGCAGCCAAUUCAU .(((.........)))(((((...(((((((((((....(((......)))..)))))....((((((........))))))...)))))).(((..((((.....)))))))))))).. ( -27.60) >DroEre_CAF1 4115 101 - 1 UUCCAGUCGGUCUGGAGAGUUUCAAG-------------------CCCACUGGCAGUGGGUUUUUAGUUUAAGCACACUAAAUCAAUUCCCAGGCAUUGCUCUUUAAGCAGGCAAUUCAU .....(((.((((((.(((((..(((-------------------(((((.....))))))))(((((........)))))...)))))))))))..((((.....)))))))....... ( -31.60) >DroYak_CAF1 38114 99 - 1 -UUCAGUCGGUCUGGAGAGUUUUAA--------------------CCCACUGGUAGUGGGUUUUUAGUUUAAGCACACUAAAUCAAUUCCCAGGCAUUGCUCUUUAAGCAGCCAAUUCAU -........((((((.(((((..((--------------------(((((.....)))))))((((((........))))))..))))))))))).(((((.....)))))......... ( -28.50) >consensus UUCCAGUCGGUCUGGAGAGUUUCAAG___________________CCCACUGGUAGUGGGUUUUUAGUUUAAGCACACUAAAUCAAUUCCCAGGCAUUGCUCUUUAAGCAGCCAAUUCAU .........((((((.(((((........................(((((.....)))))..((((((........))))))..))))))))))).(((((.....)))))......... (-23.64 = -23.87 + 0.23)


| Location | 22,426,261 – 22,426,381 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 78.89 |
| Mean single sequence MFE | -21.91 |
| Consensus MFE | -15.74 |
| Energy contribution | -16.63 |
| Covariance contribution | 0.89 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.862506 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
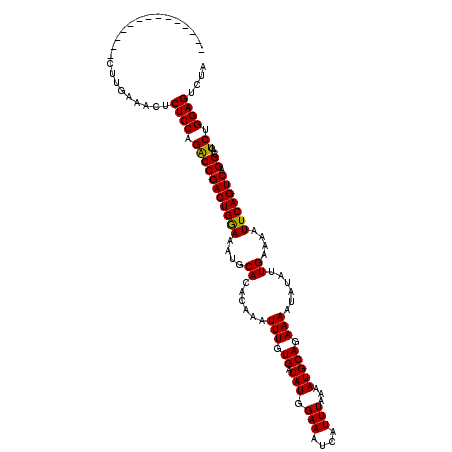
>3L_DroMel_CAF1 22426261 120 + 23771897 UAGUGUGCUUAAACUAAAAACCGACAACCAGUUAGUAGACUUGUUCUGCCAUUCCCUGAAACUCUCCUGGCCGACUGGAAAUGCACACAAAUUUGUGACAUGGAAAUCAUUUAAAAUGCA ..((((..(((((.......((.....((((((.(((((.....))))).....((.((.....))..))..))))))..(((((((......)))).)))))......))))).)))). ( -20.42) >DroEre_CAF1 4155 101 + 1 UAGUGUGCUUAAACUAAAAACCCACUGCCAGUGGG-------------------CUUGAAACUCUCCAGACCGACUGGAAAUGCACACAAAUUUCUGACAUGGAAAUCAUUUAAAAUGCA ..((((((............(((((.....)))))-------------------..........(((((.....)))))...))))))..((((((.....))))))............. ( -26.60) >DroYak_CAF1 38154 96 + 1 UAGUGUGCUUAAACUAAAAACCCACUACCAGUGGG--------------------UUAAAACUCUCCAGACCGACUGAA----CACACAAAUUUGUGACAUAGAAAUCAUUUAAAAUGCA ..(((((...........(((((((.....)))))--------------------)).........(((.....)))..----)))))......((((........)))).......... ( -18.70) >consensus UAGUGUGCUUAAACUAAAAACCCACUACCAGUGGG___________________CUUGAAACUCUCCAGACCGACUGGAAAUGCACACAAAUUUGUGACAUGGAAAUCAUUUAAAAUGCA ..((((((............(((((.....))))).............................(((((.....)))))...))))))................................ (-15.74 = -16.63 + 0.89)



| Location | 22,426,301 – 22,426,421 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 84.39 |
| Mean single sequence MFE | -22.93 |
| Consensus MFE | -17.01 |
| Energy contribution | -16.57 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.643421 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22426301 120 + 23771897 UUGUUCUGCCAUUCCCUGAAACUCUCCUGGCCGACUGGAAAUGCACACAAAUUUGUGACAUGGAAAUCAUUUAAAAUGCAGAAAAUAUAUUGAAAAUUCAGUCAGGCACUCUGGAGUCUA .......................((((.(((((((((((.(((((((......)))).)))........(((((.(((.......))).)))))..))))))).))).)...)))).... ( -23.40) >DroEre_CAF1 4190 106 + 1 --------------CUUGAAACUCUCCAGACCGACUGGAAAUGCACACAAAUUUCUGACAUGGAAAUCAUUUAAAAUGCAGAAAAUAUAUUGAAAAUUCAGUCAGGCACUCUGGAGUCUA --------------.........((((((((((((((((....((......((((((.(((.(((....)))...)))))))))......))....))))))).))...))))))).... ( -27.20) >DroYak_CAF1 38189 101 + 1 ---------------UUAAAACUCUCCAGACCGACUGAA----CACACAAAUUUGUGACAUAGAAAUCAUUUAAAAUGCAGAAAAUAUAUUGAAAAUUCAGUCAGGCACUCAGGAGUCUA ---------------.....(((((..((.(((((((((----((((......))))............(((((.(((.......))).)))))..))))))).))..))..)))))... ( -18.20) >consensus ______________CUUGAAACUCUCCAGACCGACUGGAAAUGCACACAAAUUUGUGACAUGGAAAUCAUUUAAAAUGCAGAAAAUAUAUUGAAAAUUCAGUCAGGCACUCUGGAGUCUA .......................((((.(((((((((((....((......(((.((.(((.(((....)))...))))).)))......))....))))))).))...)).)))).... (-17.01 = -16.57 + -0.44)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:20:32 2006