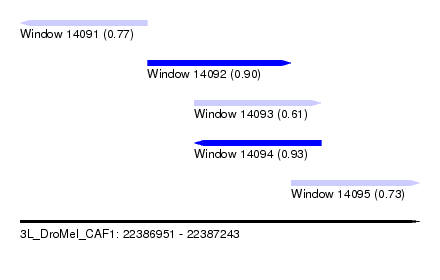
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 22,386,951 – 22,387,243 |
| Length | 292 |
| Max. P | 0.933078 |

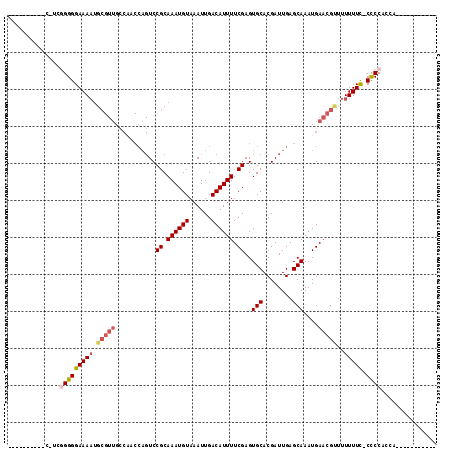
| Location | 22,386,951 – 22,387,044 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 82.43 |
| Mean single sequence MFE | -27.28 |
| Consensus MFE | -19.07 |
| Energy contribution | -20.02 |
| Covariance contribution | 0.95 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.52 |
| SVM RNA-class probability | 0.767722 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22386951 93 - 23771897 ----------C-UCUGGGGAAAAUGCGUUACCAACCAGUCCGCAAAUGUAAAUUGACAUUUUCGAGUGCACGAUUGAGCAAAUGAACGUUUUUUUC-CCCCACCC----------- ----------.-...((((((((.(((((.....((((((((.((((((......)))))).)).(....)))))).)......)))))..)))))-))).....----------- ( -24.20) >DroSec_CAF1 38076 91 - 1 ----------C-UCGGGGGAAAAUGCGUUGCCAACCAGUCCGCAAAUGUAAAUUGACAUUUUCGAGUGCACGAUUGAGCAAAUGAACGUUUUUUUC-CCCCAC------------- ----------.-..(((((((((.((((((((((...((.(((((((((......))))))....))).))..))).))).....))))..)))))-))))..------------- ( -29.10) >DroSim_CAF1 53660 93 - 1 ----------C-UCGGGGGAAAAUGCGUUGCCAACCAGUCCGCAAAUGUAAAUUGACAUUUUCGAGUGCACGAUUGAGCAAAUGAACGUUUUUUUC-CCCCACCC----------- ----------.-..(((((((((.((((((((((...((.(((((((((......))))))....))).))..))).))).....))))..)))))-))))....----------- ( -29.10) >DroEre_CAF1 54352 94 - 1 ----------C-UCGGGGGAAAAUGCGUUGCCAACCAGUCCGCAAAUGUAAAUUGACAUUUUCGAGUGCACGAUUGAGCAAAUGAACGUUUUUUUCCCCCCACCA----------- ----------.-..(((((((((.((((((((((...((.(((((((((......))))))....))).))..))).))).....))))..))))))))).....----------- ( -28.20) >DroYak_CAF1 54611 78 - 1 ----------C-UCGGGGGAAA----------------UCCGCAAAUGUAAAUUGACAUUUUCGAGUGCACGAUUGAGCAAAUGAACGUUUUUUUUCCCCCACCA----------- ----------.-..((((((((----------------...((((((((......))))))(((..(((........)))..)))..))....))))))))....----------- ( -22.40) >DroPer_CAF1 95126 115 - 1 GGCGGCGCCACACCAGGGGAAAAUGCGUUGGCUGCCAGUCCGCAAAUGUAAAUUGACAUUUUCGAGUGCACGAUUGAGCAAAUGAACGCUUUUUUC-CUCCUCCUUCUCCCUACAU (((...))).....(((((((((.(((((.(.((((((((((.((((((......)))))).)).(....)))))).)))..).)))))..)))))-).))).............. ( -30.70) >consensus __________C_UCGGGGGAAAAUGCGUUGCCAACCAGUCCGCAAAUGUAAAUUGACAUUUUCGAGUGCACGAUUGAGCAAAUGAACGUUUUUUUC_CCCCACCA___________ ..............(((((((((.(((((...........((.((((((......)))))).))..(((........)))....)))))..))))).))))............... (-19.07 = -20.02 + 0.95)


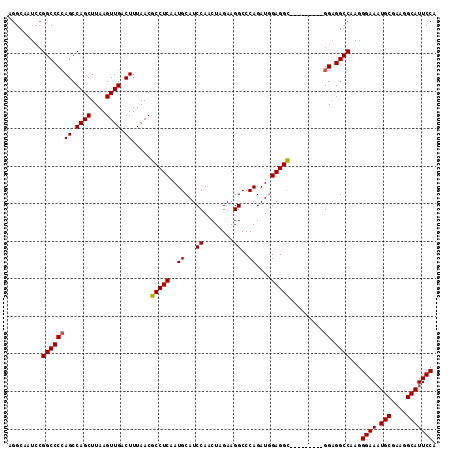
| Location | 22,387,044 – 22,387,149 |
|---|---|
| Length | 105 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 91.78 |
| Mean single sequence MFE | -36.95 |
| Consensus MFE | -31.09 |
| Energy contribution | -31.15 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.84 |
| SVM decision value | 1.02 |
| SVM RNA-class probability | 0.902200 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22387044 105 + 23771897 AGGCAAUACGGCCCCAGCCAGCUUAAGUUGACUUUAACGCCUCAAUGCAUCCAACUAGAAGGCCCAGAUGGAGGC---------GGUGGCCAAGGGAAAUGCGAAGGCAUUCCA .........((((((((.((((....)))).)).....(((((..((...((........))..))....)))))---------)).))))...((((.(((....))))))). ( -35.40) >DroSec_CAF1 38167 105 + 1 AGGCAAUCCGGCCCCAGCCAGCUUAAGUUGACUUUAACGCCUCAAUGCAUCCAACUAGAAGGCCCAGAUUGAGGC---------GGAGGCCAAGGGAAAUGCGAAGGCAUUCCA .((....))((((((((.((((....)))).)).....((((((((....((........)).....))))))))---------)).))))...((((.(((....))))))). ( -37.70) >DroEre_CAF1 54446 114 + 1 AGGCAAUCGGGCCCCAGCCAGCUUAAGUUGACUUUAACGCCUCAAUGCAUCCAACUAGAAGGCCCAGAUGGAGGCGGAGGAGGCGGAGGCCAAGGGAAAUGCAAAGGCAUUCCA .........((((((((.((((....)))).)).....(((((..(((.((((.((.(.....).)).)))).)))...))))))).))))...((((.(((....))))))). ( -41.40) >DroYak_CAF1 54689 105 + 1 AGGCAAUCCGGCCCAAGCCAGCUUAAGUUGACUUUAACGCCUCAAUGCAUCCAACUAGAAGGCCCAGAUGGAGGU---------GGAGGCCAAUGGAACUGCGAAGGCAUUCCA .((....))((((((((.((((....)))).)))...((((((..((...((........))..))....)))))---------)).))))..(((((.(((....)))))))) ( -33.30) >consensus AGGCAAUCCGGCCCCAGCCAGCUUAAGUUGACUUUAACGCCUCAAUGCAUCCAACUAGAAGGCCCAGAUGGAGGC_________GGAGGCCAAGGGAAAUGCGAAGGCAUUCCA .........((((((((.((((....)))).)).....(((((..((...((........))..))....))))).........)).))))...((((.(((....))))))). (-31.09 = -31.15 + 0.06)


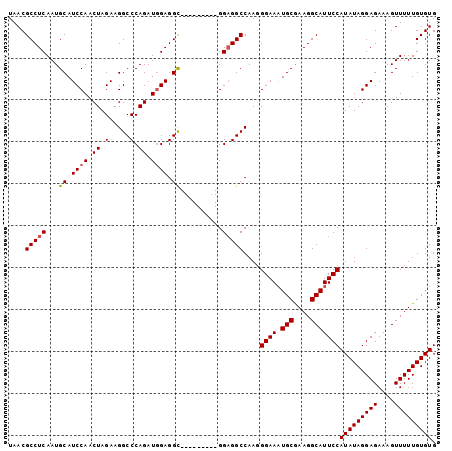
| Location | 22,387,078 – 22,387,171 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 92.31 |
| Mean single sequence MFE | -28.38 |
| Consensus MFE | -22.43 |
| Energy contribution | -22.75 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.610736 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22387078 93 + 23771897 UAACGCCUCAAUGCAUCCAACUAGAAGGCCCAGAUGGAGGC---------GGUGGCCAAGGGAAAUGCGAAGGCAUUCCAUAUAGGAGAAAGUUUUUGUGUG ....(((....(((.((((.((.(.....).)).)))).))---------)..)))....((((.(((....)))))))(((((((((....))))))))). ( -25.80) >DroSec_CAF1 38201 93 + 1 UAACGCCUCAAUGCAUCCAACUAGAAGGCCCAGAUUGAGGC---------GGAGGCCAAGGGAAAUGCGAAGGCAUUCCAUAUAGGAGAAAGUUUUUGUGUG ...(((((((((....((........)).....))))))))---------).........((((.(((....)))))))(((((((((....))))))))). ( -27.70) >DroEre_CAF1 54480 102 + 1 UAACGCCUCAAUGCAUCCAACUAGAAGGCCCAGAUGGAGGCGGAGGAGGCGGAGGCCAAGGGAAAUGCAAAGGCAUUCCAUAUAGGAGAAAGUUUUUGUGUG ...((((((..(((.((((.((.(.....).)).)))).)))...)))))).............((((((((((.((((.....))))...)))))))))). ( -32.30) >DroYak_CAF1 54723 93 + 1 UAACGCCUCAAUGCAUCCAACUAGAAGGCCCAGAUGGAGGU---------GGAGGCCAAUGGAACUGCGAAGGCAUUCCAUAUAGGAGAAAGUUUUUGUGUG ....(((((...((.((((.((.(.....).)).)))).))---------.)))))..((((((.(((....)))))))))(((.((((....)))).))). ( -27.70) >consensus UAACGCCUCAAUGCAUCCAACUAGAAGGCCCAGAUGGAGGC_________GGAGGCCAAGGGAAAUGCGAAGGCAUUCCAUAUAGGAGAAAGUUUUUGUGUG ....(((((...((.((((.((.(.....).)).)))).))..........)))))....((((.(((....)))))))(((((((((....))))))))). (-22.43 = -22.75 + 0.31)


| Location | 22,387,078 – 22,387,171 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 92.31 |
| Mean single sequence MFE | -25.60 |
| Consensus MFE | -21.35 |
| Energy contribution | -21.85 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.32 |
| Structure conservation index | 0.83 |
| SVM decision value | 1.23 |
| SVM RNA-class probability | 0.933078 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22387078 93 - 23771897 CACACAAAAACUUUCUCCUAUAUGGAAUGCCUUCGCAUUUCCCUUGGCCACC---------GCCUCCAUCUGGGCCUUCUAGUUGGAUGCAUUGAGGCGUUA ...............(((.....)))(((((((.(((........(((....---------)))((((.((((.....)))).)))))))...))))))).. ( -24.20) >DroSec_CAF1 38201 93 - 1 CACACAAAAACUUUCUCCUAUAUGGAAUGCCUUCGCAUUUCCCUUGGCCUCC---------GCCUCAAUCUGGGCCUUCUAGUUGGAUGCAUUGAGGCGUUA ...............(((.....)))(((((((.((((((..((.(((((..---------..........)))))....))..))))))...))))))).. ( -24.60) >DroEre_CAF1 54480 102 - 1 CACACAAAAACUUUCUCCUAUAUGGAAUGCCUUUGCAUUUCCCUUGGCCUCCGCCUCCUCCGCCUCCAUCUGGGCCUUCUAGUUGGAUGCAUUGAGGCGUUA .................(((...(((((((....)))))))...)))....((((((....((.((((.((((.....)))).)))).))...))))))... ( -29.00) >DroYak_CAF1 54723 93 - 1 CACACAAAAACUUUCUCCUAUAUGGAAUGCCUUCGCAGUUCCAUUGGCCUCC---------ACCUCCAUCUGGGCCUUCUAGUUGGAUGCAUUGAGGCGUUA ........(((..........(((((((((....))).))))))..(((((.---------.(.((((.((((.....)))).)))).)....)))))))). ( -24.60) >consensus CACACAAAAACUUUCUCCUAUAUGGAAUGCCUUCGCAUUUCCCUUGGCCUCC_________GCCUCCAUCUGGGCCUUCUAGUUGGAUGCAUUGAGGCGUUA ...............(((.....)))(((((((.((((((..((.((((.......................))))....))..))))))...))))))).. (-21.35 = -21.85 + 0.50)


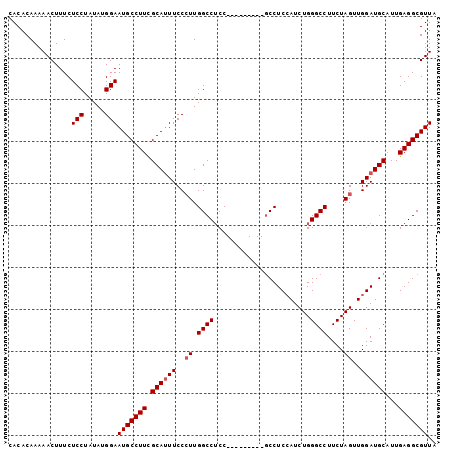
| Location | 22,387,149 – 22,387,243 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 97.04 |
| Mean single sequence MFE | -22.27 |
| Consensus MFE | -20.05 |
| Energy contribution | -20.30 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.90 |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.728319 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22387149 94 + 23771897 UAUAGGAGAAAGUUUUUGUGUGUGCAACGGGGAA-AAA-GAAAACUUUUUAUUAAAGUCAGCUGAAAUGGGUGGAACUCGUUUCGGUUUCGAAUUU ..(((.(((((((((((.....(.(....).)..-...-))))))))))).))).....((((((((((((.....))))))))))))........ ( -21.30) >DroSec_CAF1 38272 95 + 1 UAUAGGAGAAAGUUUUUGUGUGUGCAACAGGGAAAAAA-GAAAACUUUUUAUUAAAGUCAGCUGAAAUGGGUGGAACUCGUUUCGGUCUCGAAUUU ..(((.(((((((((((.....(.(.....).).....-))))))))))).)))...((((((((((((((.....)))))))))).)).)).... ( -21.20) >DroEre_CAF1 54560 96 + 1 UAUAGGAGAAAGUUUUUGUGUGUGCAACAGGGAAAAAAGGAAAACUUUUUAUUAAAGUCAGCUGAAAUGGGUGGAACUCGUUUCGGUUUCGGAUUU ..(((.(((((((((((.(.....(......).....).))))))))))).)))(((((.(((((((((((.....)))))))))))....))))) ( -23.30) >DroYak_CAF1 54794 96 + 1 UAUAGGAGAAAGUUUUUGUGUGUGCAACAGGGAAAAAAGGAAAACUUUUUAUUAAAGUCAGCUGAAAUGGGUGGAACUCGUUUCGGUUUCGGAUUU ..(((.(((((((((((.(.....(......).....).))))))))))).)))(((((.(((((((((((.....)))))))))))....))))) ( -23.30) >consensus UAUAGGAGAAAGUUUUUGUGUGUGCAACAGGGAAAAAA_GAAAACUUUUUAUUAAAGUCAGCUGAAAUGGGUGGAACUCGUUUCGGUUUCGAAUUU ..(((.(((((((((((.(.....(......).....).))))))))))).))).....((((((((((((.....))))))))))))........ (-20.05 = -20.30 + 0.25)

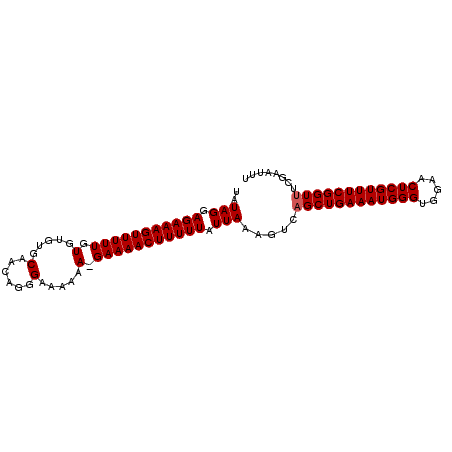
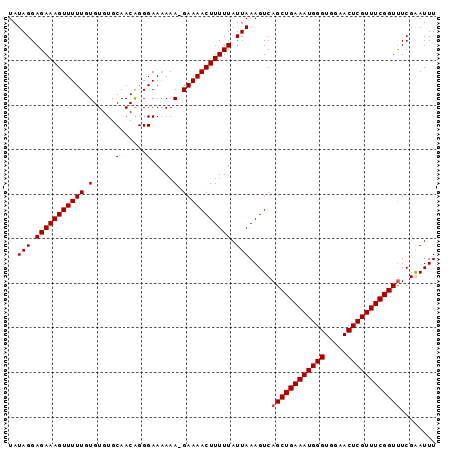
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:20:05 2006