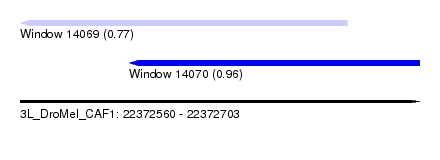
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 22,372,560 – 22,372,703 |
| Length | 143 |
| Max. P | 0.959021 |

| Location | 22,372,560 – 22,372,677 |
|---|---|
| Length | 117 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.74 |
| Mean single sequence MFE | -41.86 |
| Consensus MFE | -30.52 |
| Energy contribution | -31.88 |
| Covariance contribution | 1.36 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.52 |
| SVM RNA-class probability | 0.768700 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22372560 117 - 23771897 AGCACAAUCUUGCAC--UUUUCGCCGAAGGUGAAUGGGCAAGACAAUCAUGGCCGAGUGCCUCUGGCGGCAUUUGGGCUUGCAAACACUCCAGUGGAAUUCCUCUGCA-UGUGCUAAGAG ((((((.((((((.(--(.((((((...)))))).))))))))....((.(.(((((((((......))))))))).).)).........(((.((....)).)))..-))))))..... ( -45.00) >DroSec_CAF1 23700 117 - 1 GGCACAAUCUUGCAC--UUUUCGCCGAAGGUGAAUGGGCAAGACAAUCAUGGCCGAGUGCCUCUGGCGGCAUUUGGGCUUGCAAAUUCUCCAGUGGGAUUCCUCUGCA-UGUGCUAAGAG ((((((.((((((.(--(.((((((...)))))).)))))))).......(.(((((((((......))))))))).).((((.....(((....)))......))))-))))))..... ( -44.60) >DroSim_CAF1 38132 117 - 1 GGCACAAUCUUGCAC--UUUUCGCCGAAGGUGAAUGGGCAAGACAAUCAUGGCCGAGUGCCUCUGGCGGCAUUUGGGCUUGCAAAUUCCCCAGUGGGAUUCCUCUGCA-UGUGCUAAGAG ((((((.((((((.(--(.((((((...)))))).)))))))).......(.(((((((((......))))))))).).((((...((((....))))......))))-))))))..... ( -47.20) >DroEre_CAF1 40531 116 - 1 GGCACAAUCUUGCAAUCACUGCGCCGAAGGUGAAUGGGCAAGACCAUCAUGGCCGAGUGUCUCUGG---CAUCUGGGCUUUCAAGCAC-CCAGUGGGAUUCCCCUGCACUGGGCUAUGUG ((.....((((((..(((((........)))))....))))))))..(((((((.(((((......---...(((((..........)-)))).(((....))).))))).))))))).. ( -41.40) >DroYak_CAF1 32100 109 - 1 GGCACAGUCUUGCAC--A-UUUUCCAAAGGUGAACGGGCAAGAAAAUCAUGGCCGAGUGUCUUUGG---CAUCUGGGC----AAAUAC-CCAGUGGGAUUUCAUUGAAGUGUACUAUUUG .(((......)))((--(-((((((((((..(..(((.((.(.....).)).)))..)..))))))---((.(((((.----.....)-))))))..........)))))))........ ( -31.10) >consensus GGCACAAUCUUGCAC__UUUUCGCCGAAGGUGAAUGGGCAAGACAAUCAUGGCCGAGUGCCUCUGGCGGCAUUUGGGCUUGCAAAUAC_CCAGUGGGAUUCCUCUGCA_UGUGCUAAGAG ((((((.((((((......((((((...))))))...)))))).......(.(((((((((......))))))))).)............(((.((....)).)))...))))))..... (-30.52 = -31.88 + 1.36)

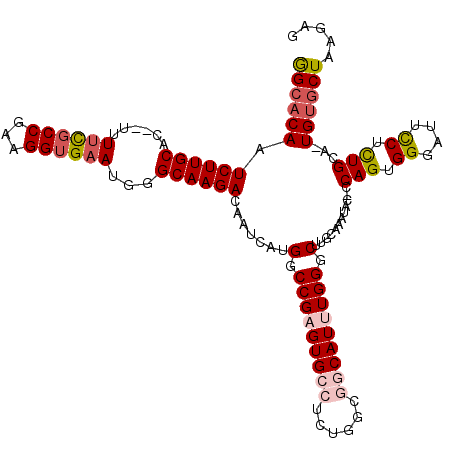
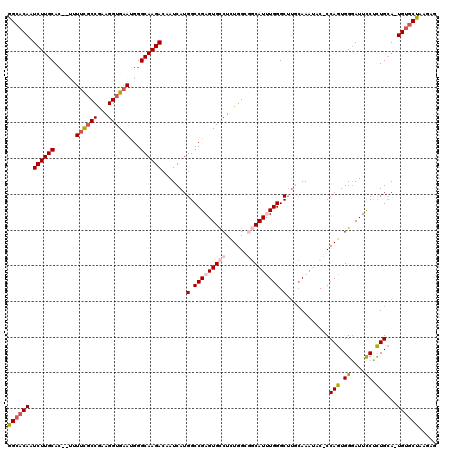
| Location | 22,372,599 – 22,372,703 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 81.61 |
| Mean single sequence MFE | -40.74 |
| Consensus MFE | -28.54 |
| Energy contribution | -30.06 |
| Covariance contribution | 1.52 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.38 |
| Structure conservation index | 0.70 |
| SVM decision value | 1.50 |
| SVM RNA-class probability | 0.959021 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22372599 104 - 23771897 GUACAGCCACUAGCUCC---ACUCCUCGGAGCACAAUCUUGCAC--UUUUCGCCGAAGGUGAAUGGGCAAGACAAUCAUGGCCGAGUGCCUCUGGCGGCAUUUGGGCUU ....((((.((((((((---.......)))))....((((((.(--(.((((((...)))))).))))))))......))).((((((((......)))))))))))). ( -41.50) >DroSec_CAF1 23739 107 - 1 GUACAGCCACUGGCCCCCCGACUCAUCGGGGCACAAUCUUGCAC--UUUUCGCCGAAGGUGAAUGGGCAAGACAAUCAUGGCCGAGUGCCUCUGGCGGCAUUUGGGCUU ....((((...((((((((((....)))))).....((((((.(--(.((((((...)))))).)))))))).......))))(((((((......))))))).)))). ( -48.10) >DroSim_CAF1 38171 107 - 1 GUACAGCCACUGGCCCCCCGACUCCUCGGGGCACAAUCUUGCAC--UUUUCGCCGAAGGUGAAUGGGCAAGACAAUCAUGGCCGAGUGCCUCUGGCGGCAUUUGGGCUU ....((((...((((((((((....)))))).....((((((.(--(.((((((...)))))).)))))))).......))))(((((((......))))))).)))). ( -48.20) >DroEre_CAF1 40570 92 - 1 GUG-----------CCC---ACUCCUCGGGGCACAAUCUUGCAAUCACUGCGCCGAAGGUGAAUGGGCAAGACCAUCAUGGCCGAGUGUCUCUGG---CAUCUGGGCUU (((-----------(((---........))))))..((((((..(((((........)))))....)))))).......((((.(((((.....)---)).)).)))). ( -35.20) >DroYak_CAF1 32137 98 - 1 AUACAGUCACUCGCCCC---ACUCUUUGGGGCACAGUCUUGCAC--A-UUUUCCAAAGGUGAACGGGCAAGAAAAUCAUGGCCGAGUGUCUUUGG---CAUCUGGGC-- .....(((....(((((---(.....)))))).(((...(((..--.-.....(((((..(..(((.((.(.....).)).)))..)..))))))---)).))))))-- ( -30.70) >consensus GUACAGCCACUGGCCCC___ACUCCUCGGGGCACAAUCUUGCAC__UUUUCGCCGAAGGUGAAUGGGCAAGACAAUCAUGGCCGAGUGCCUCUGGCGGCAUUUGGGCUU ............(((((..........)))))....((((((......((((((...))))))...)))))).......(.(((((((((......))))))))).).. (-28.54 = -30.06 + 1.52)

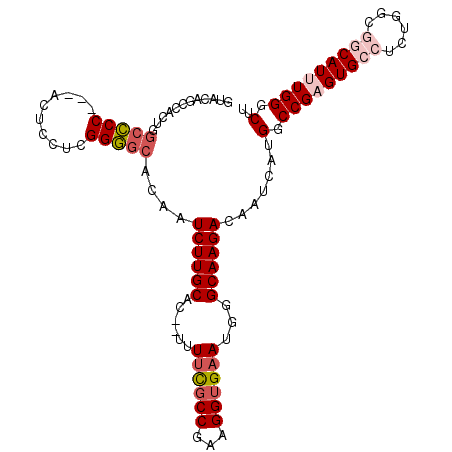
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:19:42 2006