
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 22,371,560 – 22,371,724 |
| Length | 164 |
| Max. P | 0.705121 |

| Location | 22,371,560 – 22,371,660 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 80.66 |
| Mean single sequence MFE | -38.23 |
| Consensus MFE | -26.36 |
| Energy contribution | -29.30 |
| Covariance contribution | 2.94 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.69 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.516999 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
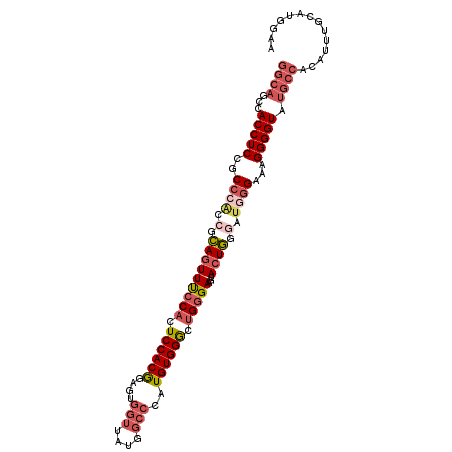
>3L_DroMel_CAF1 22371560 100 - 23771897 GGCAGUCACCUCUGCCCGCCGCAGUUUCCACUCCACA------UUAUGUUCCAUGUGGACUGGGAAGUACUGGGAUUGGAAAGGGGUAUGCCAAAUUUGGAUGGAA ((((...((((((..((.((.(((((((((.((((((------(........))))))).)))))...))))))...))..)))))).)))).............. ( -38.80) >DroSec_CAF1 22701 106 - 1 GGCAGCCACCUCCGCCCACCGCAGUUUCCACUCCACGGAGUGGUUAUGGCCCAUGUGGGCUGGGAAGUACUGGAAUGGGAAAGGGGUAUGCCACAUUUGCAUGGAA (((((((.(((...((((...((((..((((((....))))))....(((((....))))).......))))...))))..)))))).)))).............. ( -42.50) >DroSim_CAF1 37096 106 - 1 GGCAGCCACCUCCGCCCACCGCAGUUUCCACUCCACGGAGUGGUUAUGGCCCAUGUGGGCUGGGAAGUACUGGGAUGGGAAAGGGGUAUGCCACAUUUCCAUGGAA (((((((.(((...((((((.((((..((((((....))))))....(((((....))))).......)))))).))))..)))))).))))....(((....))) ( -46.70) >DroYak_CAF1 31136 100 - 1 GACUGCUACCUCUGC------UACUUCGCUCUCCACGUAGUGGUUAUGGCCUAUGUGGGCUGGGAUGUACUAAGACUGGGAAGGGGUAUACAACAUUUGCAUGGAA ...((((((((((..------(((.((.(..((((((((..(((....)))))))))))..).)).)))(((....)))..)))))))..........)))..... ( -24.90) >consensus GGCAGCCACCUCCGCCCACCGCAGUUUCCACUCCACGGAGUGGUUAUGGCCCAUGUGGGCUGGGAAGUACUGGGAUGGGAAAGGGGUAUGCCACAUUUGCAUGGAA ((((...(((((..((((.(.(((((((((.((((((....(((....)))..)))))).)))))...)))).).))))...))))).)))).............. (-26.36 = -29.30 + 2.94)



| Location | 22,371,623 – 22,371,724 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 77.83 |
| Mean single sequence MFE | -35.52 |
| Consensus MFE | -26.17 |
| Energy contribution | -25.29 |
| Covariance contribution | -0.88 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.36 |
| SVM RNA-class probability | 0.705121 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22371623 101 + 23771897 ---UGUGGAGUGGAAACUGCGGCGGGCAGAGGUGACUGCC----------------AGCUGUUUACUCGUUUCUGCUGCGGCUGCACUUCACUUUGAUUUAUCAAUAAAAGCCAAAGUGU ---.((.(((..(((((.(((((.(((((......)))))----------------.)))))......)))))..)).).)).......(((((((..((........))..))))))). ( -33.50) >DroSec_CAF1 22767 104 + 1 CUCCGUGGAGUGGAAACUGCGGUGGGCGGAGGUGGCUGCC----------------AGCUGUCUACUGGCUGCUGCUGCGGCUGCACUUCACUUUGAUUUAUUAAUAAAAGCCAAAGUUU (.((((..(((....))))))).)((((((((..((((((----------------(((.(((....))).))))..)))))..).)))).......((((....)))).)))....... ( -37.60) >DroSim_CAF1 37162 104 + 1 CUCCGUGGAGUGGAAACUGCGGUGGGCGGAGGUGGCUGCC----------------AGCUGUCUACUGGCUGCUGCUGCGGCUGCACUUCACUUUGAUUUAUCAAUAAAAGCCAAAGUUU (.((((..(((....))))))).)((((((((..((((((----------------(((.(((....))).))))..)))))..).))))...((((....)))).....)))....... ( -39.60) >DroYak_CAF1 31202 114 + 1 CUACGUGGAGAGCGAAGUA------GCAGAGGUAGCAGUCGCAGAGAUGGGAGAGAAGCUGACUACUGGCUGUUGCUGUGUAUGCACUUCACUUUGAUUUAUCAAUAAAAGACAAAGUUU ....((((((.(((.((((------((((.(((((((((..(............)..)))).)))))..)))))))).....))).)))))).((((....))))............... ( -31.40) >consensus CUCCGUGGAGUGGAAACUGCGGUGGGCAGAGGUGGCUGCC________________AGCUGUCUACUGGCUGCUGCUGCGGCUGCACUUCACUUUGAUUUAUCAAUAAAAGCCAAAGUUU ....((((((((.(..((((((((.((((.((((((.((..................)).).)))))..)))))))))))).).)))))))).((((....))))............... (-26.17 = -25.29 + -0.88)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:19:40 2006