
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 2,580,391 – 2,580,485 |
| Length | 94 |
| Max. P | 0.999963 |

| Location | 2,580,391 – 2,580,485 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 86.39 |
| Mean single sequence MFE | -37.16 |
| Consensus MFE | -36.60 |
| Energy contribution | -36.12 |
| Covariance contribution | -0.48 |
| Combinations/Pair | 1.11 |
| Mean z-score | -3.73 |
| Structure conservation index | 0.98 |
| SVM decision value | 4.94 |
| SVM RNA-class probability | 0.999963 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
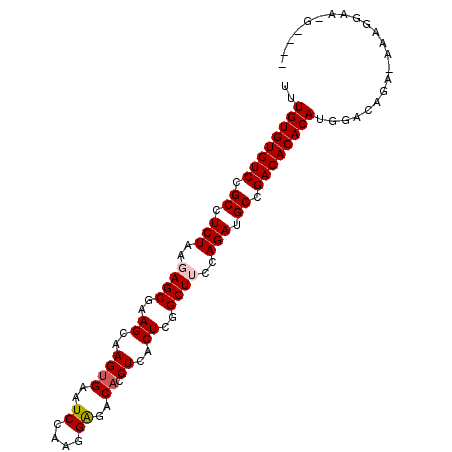
>3L_DroMel_CAF1 2580391 94 + 23771897 ----CACACCUUU-UCUGACCAUGUGUGUCGGCAUCUGGAAGCCGAGUGAGGUGUCUCCAUGGAUUCACUUGCUUUGCUCUUAGAGGCGGACACACAAA ----.........-........((((((((.((.((((..(((.((((.(((((..((....))..))))))))).)))..)))).)).)))))))).. ( -37.60) >DroSec_CAF1 7377 92 + 1 ------ACUCUUU-UCUGUACAUGUGUGUCGGCAUCUGGAAGCCGAGUGAGGUGUCUCCCUGGAUUCACUUGCUUCGCUCUUAGAGGCGGACACACAAA ------.......-........((((((((.((.((((..(((.((((.(((((..((....))..))))))))).)))..)))).)).)))))))).. ( -37.60) >DroSim_CAF1 7366 92 + 1 ------CACCUUU-UCCGUCCAUGUGUGUCGGCAUCUGGAAGCCGAGUGAGGUGUCUCCUUGGAUUCACUUGCUUCGCUCUUAGAGGCGGACACACAAA ------.......-........((((((((.((.((((..(((.((((.(((((..((....))..))))))))).)))..)))).)).)))))))).. ( -37.60) >DroEre_CAF1 4683 98 + 1 UUUUCUUUCCUUU-UCUGUGCAUGUGUGUCGGCAUCUGGCAGCCGAGUGAGGUGUCUCUUUGGACUCGCUUGCUUCGCUCUUAGAGGCGGACACACAAA .............-........((((((((.((.(((((.(((.(((..(((.((((....)))).).))..))).))).))))).)).)))))))).. ( -38.60) >DroYak_CAF1 7381 99 + 1 AUGUCUUUCCUUUUUUUGUCCAUGUGUGUCGGCAUCUGGCAGCCGAGUGAGCUGUUCCUUUGGAUUCACUUGCUUUGCUCUUAGAGGCAGACACACAAA ......................((((((((.((.(((((.(((.(((..((.((.(((...)))..))))..))).))).))))).)).)))))))).. ( -34.40) >consensus ____C_CACCUUU_UCUGUCCAUGUGUGUCGGCAUCUGGAAGCCGAGUGAGGUGUCUCCUUGGAUUCACUUGCUUCGCUCUUAGAGGCGGACACACAAA ......................((((((((.((.(((((.(((.(((..((..((((....))))...))..))).))).))))).)).)))))))).. (-36.60 = -36.12 + -0.48)
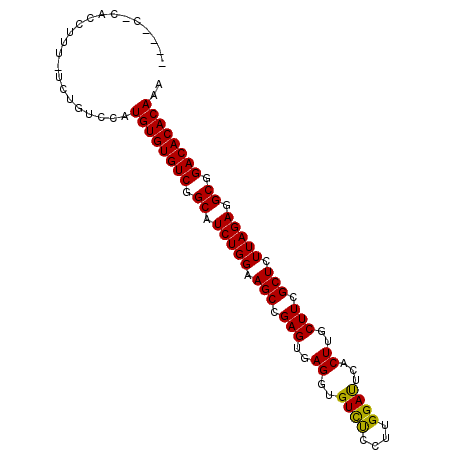


| Location | 2,580,391 – 2,580,485 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 86.39 |
| Mean single sequence MFE | -27.44 |
| Consensus MFE | -24.04 |
| Energy contribution | -24.48 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.88 |
| SVM decision value | 0.36 |
| SVM RNA-class probability | 0.706525 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 2580391 94 - 23771897 UUUGUGUGUCCGCCUCUAAGAGCAAAGCAAGUGAAUCCAUGGAGACACCUCACUCGGCUUCCAGAUGCCGACACACAUGGUCAGA-AAAGGUGUG---- ..((((((((.((.(((..((((..((...(((..((....))..)))....))..))))..))).)).))))))))........-.........---- ( -28.40) >DroSec_CAF1 7377 92 - 1 UUUGUGUGUCCGCCUCUAAGAGCGAAGCAAGUGAAUCCAGGGAGACACCUCACUCGGCUUCCAGAUGCCGACACACAUGUACAGA-AAAGAGU------ ..((((((((.((.(((....(.(((((.(((((......(....)...)))))..))))))))).)).))))))))........-.......------ ( -28.90) >DroSim_CAF1 7366 92 - 1 UUUGUGUGUCCGCCUCUAAGAGCGAAGCAAGUGAAUCCAAGGAGACACCUCACUCGGCUUCCAGAUGCCGACACACAUGGACGGA-AAAGGUG------ ..((((((((.((.(((....(.(((((.(((((......(....)...)))))..))))))))).)).))))))))........-.......------ ( -27.80) >DroEre_CAF1 4683 98 - 1 UUUGUGUGUCCGCCUCUAAGAGCGAAGCAAGCGAGUCCAAAGAGACACCUCACUCGGCUGCCAGAUGCCGACACACAUGCACAGA-AAAGGAAAGAAAA ..((((((((.((.(((..(.((..(((...(((((.....(((....))))))))))))))))).)).))))))))........-............. ( -25.60) >DroYak_CAF1 7381 99 - 1 UUUGUGUGUCUGCCUCUAAGAGCAAAGCAAGUGAAUCCAAAGGAACAGCUCACUCGGCUGCCAGAUGCCGACACACAUGGACAAAAAAAGGAAAGACAU ..((((((((.((.(((...(((..((..(((...(((...)))...)))..))..)))...))).)).))))))))...................... ( -26.50) >consensus UUUGUGUGUCCGCCUCUAAGAGCGAAGCAAGUGAAUCCAAGGAGACACCUCACUCGGCUUCCAGAUGCCGACACACAUGGACAGA_AAAGGAA_G____ ..((((((((.((.(((..((((..((..((((..((....))..)).))..))..))))..))).)).))))))))...................... (-24.04 = -24.48 + 0.44)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:52:51 2006