
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 22,322,843 – 22,323,002 |
| Length | 159 |
| Max. P | 0.588498 |

| Location | 22,322,843 – 22,322,962 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.89 |
| Mean single sequence MFE | -31.03 |
| Consensus MFE | -26.62 |
| Energy contribution | -26.73 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.588498 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
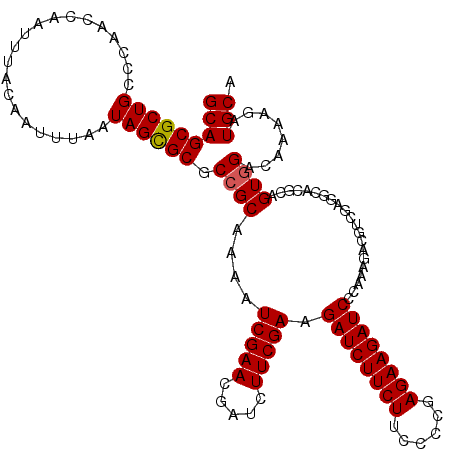
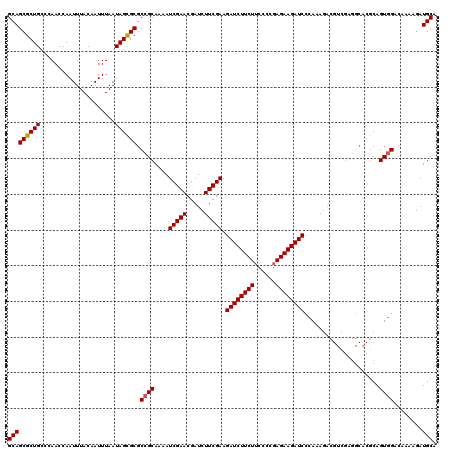
>3L_DroMel_CAF1 22322843 119 - 23771897 GCAGCGCUGCCCAACCAAUUUACAAUUUAAUAGCGCGCCGCAAAAUCGAACGAUCUUCGAAGAUCUUCUUCCCCGAGAAGAUCCCAAAGACGUCGAGGCACGCAGUGGA-AAAAGAUGCA (((((((((.....................)))))).((((....((((.((.((((.(..(((((((((....))))))))).).)))))))))).(....).)))).-......))). ( -32.70) >DroSec_CAF1 2037 120 - 1 GCAGCGCUGCCCAACCAAUUUACAAUUUAAUAGCGCGCCGCAAAAUCGAACGAUCUUCGAAGAUCUUCUUCCCCGAGAAGAUCCCAAAGACGUCGAGGCACGCAGUGGACAAAAGAUGCA (((((((((.....................))))))(((((....((((.((.((((.(..(((((((((....))))))))).).)))))))))).(....).)))).)......))). ( -33.00) >DroEre_CAF1 131 108 - 1 GCAGCGCUGCCCAACCAAUUUACAAUUUAAUAGUGCGCCGCAAAAUCGAACGAUCUUCGAAGAUCUUCUGCCCCGAGAAGAUCCCAAAC------------ACAGUAGGCAGAAGAUGCA (((((((((.....................))))))(((((....(((((.....))))).((((((((......))))))))......------------...)).)))......))). ( -27.40) >consensus GCAGCGCUGCCCAACCAAUUUACAAUUUAAUAGCGCGCCGCAAAAUCGAACGAUCUUCGAAGAUCUUCUUCCCCGAGAAGAUCCCAAAGACGUCGAGGCACGCAGUGGACAAAAGAUGCA (((((((((.....................)))))).((((....(((((.....))))).((((((((......)))))))).....................))))........))). (-26.62 = -26.73 + 0.11)

| Location | 22,322,882 – 22,323,002 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.67 |
| Mean single sequence MFE | -34.60 |
| Consensus MFE | -32.71 |
| Energy contribution | -32.93 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.95 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.519242 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22322882 120 - 23771897 GUUCAAUACGAUGUGGCAUUCAGACACGUUUGCCGCUGCAGCAGCGCUGCCCAACCAAUUUACAAUUUAAUAGCGCGCCGCAAAAUCGAACGAUCUUCGAAGAUCUUCUUCCCCGAGAAG ((((........((((((..(......)..))))))(((.((.((((((.....................)))))))).))).....))))(((((....)))))(((((....))))). ( -33.00) >DroSec_CAF1 2077 120 - 1 GUUCAAUACGAUGUGGCAUUCAGACACGUUUGCCGCUGCGGCAGCGCUGCCCAACCAAUUUACAAUUUAAUAGCGCGCCGCAAAAUCGAACGAUCUUCGAAGAUCUUCUUCCCCGAGAAG ((((........((((((..(......)..))))))((((((.((((((.....................)))))))))))).....))))(((((....)))))(((((....))))). ( -39.10) >DroEre_CAF1 159 120 - 1 GUUUAAUACGAUGUGGCAUUCAGACACGUUUGCCGGAGCGGCAGCGCUGCCCAACCAAUUUACAAUUUAAUAGUGCGCCGCAAAAUCGAACGAUCUUCGAAGAUCUUCUGCCCCGAGAAG ........((((.(((((..(......)..)))))..(((((.((((((.....................)))))))))))...))))...(((((....)))))((((......)))). ( -31.70) >consensus GUUCAAUACGAUGUGGCAUUCAGACACGUUUGCCGCUGCGGCAGCGCUGCCCAACCAAUUUACAAUUUAAUAGCGCGCCGCAAAAUCGAACGAUCUUCGAAGAUCUUCUUCCCCGAGAAG ((((........((((((..(......)..)))))).(((((.((((((.....................)))))))))))......))))(((((....)))))((((......)))). (-32.71 = -32.93 + 0.23)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:18:39 2006