
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 22,321,185 – 22,321,371 |
| Length | 186 |
| Max. P | 0.986898 |

| Location | 22,321,185 – 22,321,305 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.78 |
| Mean single sequence MFE | -35.45 |
| Consensus MFE | -35.45 |
| Energy contribution | -35.45 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.39 |
| Structure conservation index | 1.00 |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.568372 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22321185 120 + 23771897 CCUGUCAGUGUCAGUUGCCCUGAGUGGAGCUCAAGUGUGCCCCGCUCCUCCUCUGGUUAUUUGCUUAAGUGCAAAUAUGAGCAACUAUUUGCUAAAUGACAGUUAUUGCUGAUUAGCUGU .((((((..((.((((((((.(((.(((((.............)))))..))).)).(((((((......)))))))...))))))....))....)))))).....(((....)))... ( -35.12) >DroSec_CAF1 355 120 + 1 CCUGUCAGUGUCAGUUGCCCUGAGUGGAGCUCGAGUGUGCCCCGCUCCCUCUCUGGUUAUUUGCUUAAGUGCAAAUAUGAGCAACUAUUUGCUAAAUGACAGUUAUUGCUGAUUAGCUGU .((((((..((.((((((((.(((.(((((.............)))))..))).)).(((((((......)))))))...))))))....))....)))))).....(((....)))... ( -35.62) >DroSim_CAF1 475 120 + 1 CCUGUCAGUGUCAGUUGCCCUGAGUGGAGCUCGAGUGUGCACCGCUCCCUCUCUGGUUAUUUGCUUAAGUGCAAAUAUGAGCAACUAUUUGCUAAAUGACAGUUAUUGCUGAUUAGCUGU .((((((..((.((((((((.(((.(((((.............)))))..))).)).(((((((......)))))))...))))))....))....)))))).....(((....)))... ( -35.62) >consensus CCUGUCAGUGUCAGUUGCCCUGAGUGGAGCUCGAGUGUGCCCCGCUCCCUCUCUGGUUAUUUGCUUAAGUGCAAAUAUGAGCAACUAUUUGCUAAAUGACAGUUAUUGCUGAUUAGCUGU .((((((..((.((((((((.(((.(((((.............)))))..))).)).(((((((......)))))))...))))))....))....)))))).....(((....)))... (-35.45 = -35.45 + -0.00)



| Location | 22,321,185 – 22,321,305 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 97.78 |
| Mean single sequence MFE | -41.15 |
| Consensus MFE | -40.11 |
| Energy contribution | -40.22 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -4.15 |
| Structure conservation index | 0.97 |
| SVM decision value | 2.06 |
| SVM RNA-class probability | 0.986898 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22321185 120 - 23771897 ACAGCUAAUCAGCAAUAACUGUCAUUUAGCAAAUAGUUGCUCAUAUUUGCACUUAAGCAAAUAACCAGAGGAGGAGCGGGGCACACUUGAGCUCCACUCAGGGCAACUGACACUGACAGG ...(((....))).....((((((.........(((((((((.(((((((......)))))))....(((..(((((.((.....))...))))).))).)))))))))....)))))). ( -40.52) >DroSec_CAF1 355 120 - 1 ACAGCUAAUCAGCAAUAACUGUCAUUUAGCAAAUAGUUGCUCAUAUUUGCACUUAAGCAAAUAACCAGAGAGGGAGCGGGGCACACUCGAGCUCCACUCAGGGCAACUGACACUGACAGG ...(((....))).....((((((.........(((((((((.(((((((......)))))))......((((((((.(((....)))..))))).))).)))))))))....)))))). ( -42.52) >DroSim_CAF1 475 120 - 1 ACAGCUAAUCAGCAAUAACUGUCAUUUAGCAAAUAGUUGCUCAUAUUUGCACUUAAGCAAAUAACCAGAGAGGGAGCGGUGCACACUCGAGCUCCACUCAGGGCAACUGACACUGACAGG ...(((....))).....((((((.........(((((((((.(((((((......)))))))......(((((((((((....)))...))))).))).)))))))))....)))))). ( -40.42) >consensus ACAGCUAAUCAGCAAUAACUGUCAUUUAGCAAAUAGUUGCUCAUAUUUGCACUUAAGCAAAUAACCAGAGAGGGAGCGGGGCACACUCGAGCUCCACUCAGGGCAACUGACACUGACAGG ...(((....))).....((((((.........(((((((((.(((((((......)))))))....(((..(((((.(((....)))..))))).))).)))))))))....)))))). (-40.11 = -40.22 + 0.11)


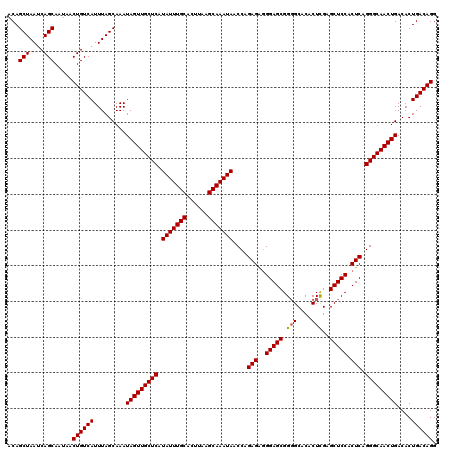
| Location | 22,321,225 – 22,321,345 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 98.33 |
| Mean single sequence MFE | -31.83 |
| Consensus MFE | -31.75 |
| Energy contribution | -31.53 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.28 |
| Structure conservation index | 1.00 |
| SVM decision value | 1.49 |
| SVM RNA-class probability | 0.958679 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
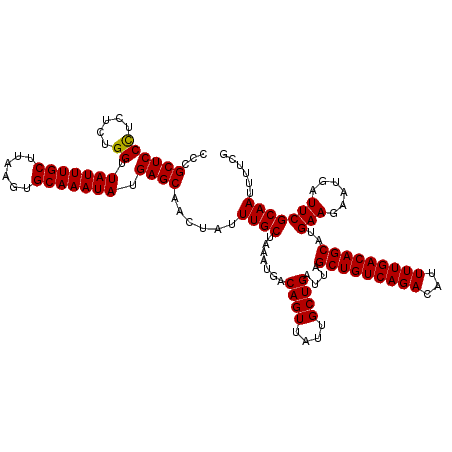
>3L_DroMel_CAF1 22321225 120 + 23771897 CCCGCUCCUCCUCUGGUUAUUUGCUUAAGUGCAAAUAUGAGCAACUAUUUGCUAAAUGACAGUUAUUGCUGAUUAGCUGUCAGACAUUUUGACAGCAUGAAGAAUGAUUCGCAAUUUUCG ...((((..((...)).(((((((......))))))).))))......((((.......((((....))))....(((((((((...)))))))))..(((......)))))))...... ( -30.30) >DroSec_CAF1 395 120 + 1 CCCGCUCCCUCUCUGGUUAUUUGCUUAAGUGCAAAUAUGAGCAACUAUUUGCUAAAUGACAGUUAUUGCUGAUUAGCUGUCAGACAUUUUGACAGCAUGAAGAAUGAUUCGCAAUUUUCG ...((((((.....)).(((((((......))))))).))))......((((.......((((....))))....(((((((((...)))))))))..(((......)))))))...... ( -32.60) >DroSim_CAF1 515 120 + 1 ACCGCUCCCUCUCUGGUUAUUUGCUUAAGUGCAAAUAUGAGCAACUAUUUGCUAAAUGACAGUUAUUGCUGAUUAGCUGUCAGACAUUUUGACAGCAUGAAGAAUGAUUCGCAAUUUUCG ...((((((.....)).(((((((......))))))).))))......((((.......((((....))))....(((((((((...)))))))))..(((......)))))))...... ( -32.60) >consensus CCCGCUCCCUCUCUGGUUAUUUGCUUAAGUGCAAAUAUGAGCAACUAUUUGCUAAAUGACAGUUAUUGCUGAUUAGCUGUCAGACAUUUUGACAGCAUGAAGAAUGAUUCGCAAUUUUCG ...((((((.....)).(((((((......))))))).))))......((((.......((((....))))....(((((((((...)))))))))..(((......)))))))...... (-31.75 = -31.53 + -0.22)

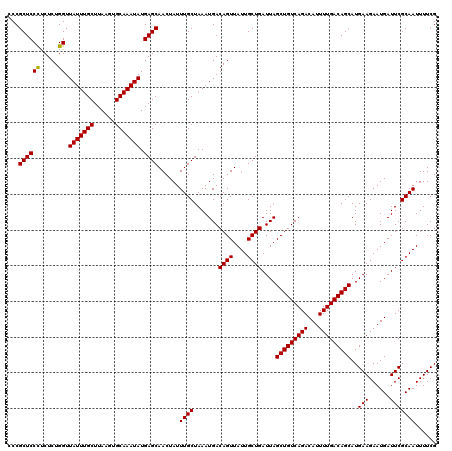
| Location | 22,321,225 – 22,321,345 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.33 |
| Mean single sequence MFE | -30.83 |
| Consensus MFE | -29.47 |
| Energy contribution | -29.80 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.28 |
| Structure conservation index | 0.96 |
| SVM decision value | 1.14 |
| SVM RNA-class probability | 0.921608 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
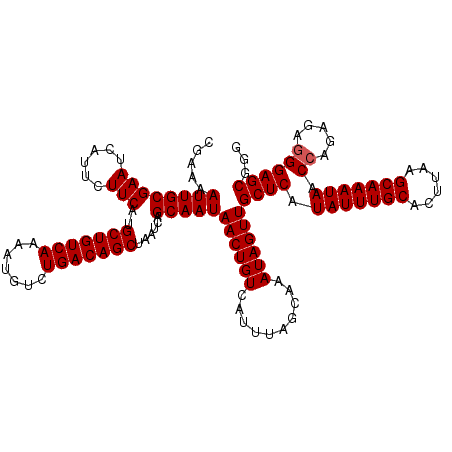
>3L_DroMel_CAF1 22321225 120 - 23771897 CGAAAAUUGCGAAUCAUUCUUCAUGCUGUCAAAAUGUCUGACAGCUAAUCAGCAAUAACUGUCAUUUAGCAAAUAGUUGCUCAUAUUUGCACUUAAGCAAAUAACCAGAGGAGGAGCGGG ......((((......((((((..(((((((.......))))))).....((((((...(((......)))....))))))..(((((((......)))))))....))))))..)))). ( -29.70) >DroSec_CAF1 395 120 - 1 CGAAAAUUGCGAAUCAUUCUUCAUGCUGUCAAAAUGUCUGACAGCUAAUCAGCAAUAACUGUCAUUUAGCAAAUAGUUGCUCAUAUUUGCACUUAAGCAAAUAACCAGAGAGGGAGCGGG .....((((((((......)))..(((((((.......)))))))......)))))((((((..........))))))((((.(((((((......))))))).((.....))))))... ( -31.40) >DroSim_CAF1 515 120 - 1 CGAAAAUUGCGAAUCAUUCUUCAUGCUGUCAAAAUGUCUGACAGCUAAUCAGCAAUAACUGUCAUUUAGCAAAUAGUUGCUCAUAUUUGCACUUAAGCAAAUAACCAGAGAGGGAGCGGU .....((((((((......)))..(((((((.......)))))))......)))))((((((..........))))))((((.(((((((......))))))).((.....))))))... ( -31.40) >consensus CGAAAAUUGCGAAUCAUUCUUCAUGCUGUCAAAAUGUCUGACAGCUAAUCAGCAAUAACUGUCAUUUAGCAAAUAGUUGCUCAUAUUUGCACUUAAGCAAAUAACCAGAGAGGGAGCGGG .....((((((((......)))..(((((((.......)))))))......)))))((((((..........))))))((((.(((((((......))))))).((.....))))))... (-29.47 = -29.80 + 0.33)


| Location | 22,321,265 – 22,321,371 |
|---|---|
| Length | 106 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.22 |
| Mean single sequence MFE | -35.00 |
| Consensus MFE | -31.78 |
| Energy contribution | -32.03 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.621681 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22321265 106 + 23771897 GCAACUAUUUGCUAAAUGACAGUUAUUGCUGAUUAGCUGUCAGACAUUUUGACAGCAUGAAGAAUGAUUCGCAAUUUUCGUUCGGCUUGGAC--------------UGGCAGCCCGGUUA ((((....))))....((.(((((...(((((...(((((((((...)))))))))(((((((.((.....)).))))))))))))...)))--------------)).))......... ( -32.20) >DroSec_CAF1 435 120 + 1 GCAACUAUUUGCUAAAUGACAGUUAUUGCUGAUUAGCUGUCAGACAUUUUGACAGCAUGAAGAAUGAUUCGCAAUUUUCGUUCGGCUUGGACUCGGCUUGGAGCCCUGGCAGCCCGGUUA ..((((..((((((......((((...(((((...(((((((((...)))))))))(((((((.((.....)).))))))))))))...)))).(((.....))).))))))...)))). ( -36.40) >DroSim_CAF1 555 120 + 1 GCAACUAUUUGCUAAAUGACAGUUAUUGCUGAUUAGCUGUCAGACAUUUUGACAGCAUGAAGAAUGAUUCGCAAUUUUCGUUCGGCUUGGACUCGGCUUGGAGCCCUGGCAGCCCGGUUA ..((((..((((((......((((...(((((...(((((((((...)))))))))(((((((.((.....)).))))))))))))...)))).(((.....))).))))))...)))). ( -36.40) >consensus GCAACUAUUUGCUAAAUGACAGUUAUUGCUGAUUAGCUGUCAGACAUUUUGACAGCAUGAAGAAUGAUUCGCAAUUUUCGUUCGGCUUGGACUCGGCUUGGAGCCCUGGCAGCCCGGUUA ..((((..((((((.......(((...(((((...(((((((((...)))))))))(((((((.((.....)).))))))))))))...)))..(((.....))).))))))...)))). (-31.78 = -32.03 + 0.25)

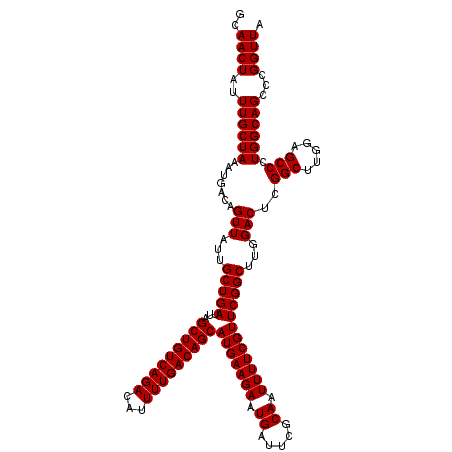

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:18:37 2006