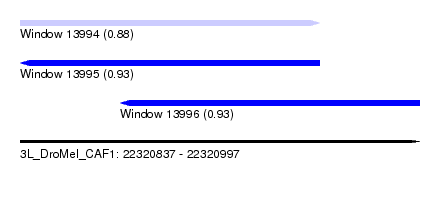
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 22,320,837 – 22,320,997 |
| Length | 160 |
| Max. P | 0.933251 |

| Location | 22,320,837 – 22,320,957 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.00 |
| Mean single sequence MFE | -40.83 |
| Consensus MFE | -38.83 |
| Energy contribution | -39.50 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.89 |
| SVM RNA-class probability | 0.875684 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22320837 120 + 23771897 CUAAUGAUGCUCUCUGGCCUGCAUCGAGAAGCCAAGGCACCAAGGCAUCAGCGGCAUCGCUGGCCGAAGCCACUCGAUGUCCAUCAUUCAUUUGGCAUCGAUGUACGAGCCAUGUUUAUG ..............(((((((((((((...((((((((.....(((..(((((....))))))))...)))....(((....)))......))))).)))))))).).))))........ ( -41.00) >DroSec_CAF1 1 114 + 1 ------AUGCUCUCUGGCCUGCAUCGAGAAGCCAAGGCACCAAGGCAUCAGCGGCAUCGCUGGCCAAAGCCACUCGAUGUCCAUCAUUCAUUUGGCUCCGAUGUACGAGCCAUGUUUAUG ------........((((((((((((.((.((((((((......))....(.(((((((.((((....))))..))))))))........))))))))))))))).).))))........ ( -40.50) >DroSim_CAF1 115 120 + 1 CUAAUGAUGCUCUCUGGCCUGCAUCGAGAAGCCAAGGCACCAAGGCAUCAGCGGCAUCGCUGGCCAAAGCCACUCGAUGUCCAUCAUUCAUUUGGCUUCGAUGUACGAGCCAUGUUUAUG ..............(((((((((((((..(((((((((......))....(.(((((((.((((....))))..))))))))........))))))))))))))).).))))........ ( -41.00) >consensus CUAAUGAUGCUCUCUGGCCUGCAUCGAGAAGCCAAGGCACCAAGGCAUCAGCGGCAUCGCUGGCCAAAGCCACUCGAUGUCCAUCAUUCAUUUGGCUUCGAUGUACGAGCCAUGUUUAUG ..............(((((((((((((..(((((((((......))....(.(((((((.((((....))))..))))))))........))))))))))))))).).))))........ (-38.83 = -39.50 + 0.67)
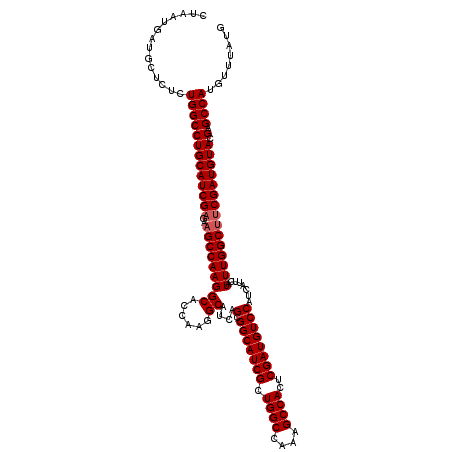


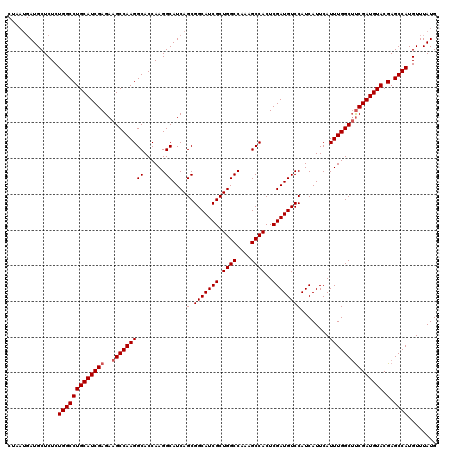
| Location | 22,320,837 – 22,320,957 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.00 |
| Mean single sequence MFE | -45.00 |
| Consensus MFE | -42.39 |
| Energy contribution | -42.17 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.94 |
| SVM decision value | 1.21 |
| SVM RNA-class probability | 0.930920 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22320837 120 - 23771897 CAUAAACAUGGCUCGUACAUCGAUGCCAAAUGAAUGAUGGACAUCGAGUGGCUUCGGCCAGCGAUGCCGCUGAUGCCUUGGUGCCUUGGCUUCUCGAUGCAGGCCAGAGAGCAUCAUUAG .........(((((((...(((((((((.........))).)))))).((((....)))))))).)))..(((((((((.(.((((..((........))))))).))).)))))).... ( -46.20) >DroSec_CAF1 1 114 - 1 CAUAAACAUGGCUCGUACAUCGGAGCCAAAUGAAUGAUGGACAUCGAGUGGCUUUGGCCAGCGAUGCCGCUGAUGCCUUGGUGCCUUGGCUUCUCGAUGCAGGCCAGAGAGCAU------ ........((((((........)))))).(((.....(((.(((((((.((((..((((((((....)))))..((....)))))..)))).)))))))....))).....)))------ ( -44.20) >DroSim_CAF1 115 120 - 1 CAUAAACAUGGCUCGUACAUCGAAGCCAAAUGAAUGAUGGACAUCGAGUGGCUUUGGCCAGCGAUGCCGCUGAUGCCUUGGUGCCUUGGCUUCUCGAUGCAGGCCAGAGAGCAUCAUUAG ........(((((((.....)).)))))(((((.((.(((.(((((((.((((..((((((((....)))))..((....)))))..)))).)))))))....))).....))))))).. ( -44.60) >consensus CAUAAACAUGGCUCGUACAUCGAAGCCAAAUGAAUGAUGGACAUCGAGUGGCUUUGGCCAGCGAUGCCGCUGAUGCCUUGGUGCCUUGGCUUCUCGAUGCAGGCCAGAGAGCAUCAUUAG ........(((((((.....))).)))).(((.....(((.(((((((.((((..((((((((....)))))..((....)))))..)))).)))))))....))).....)))...... (-42.39 = -42.17 + -0.22)

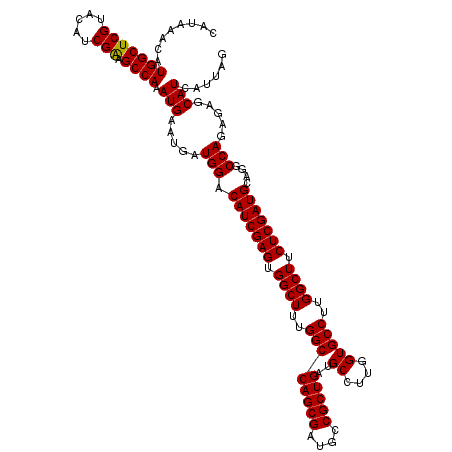
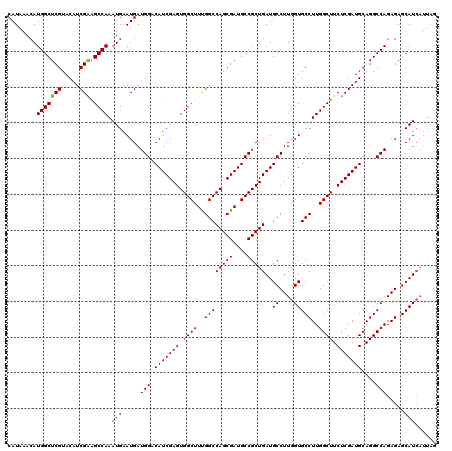
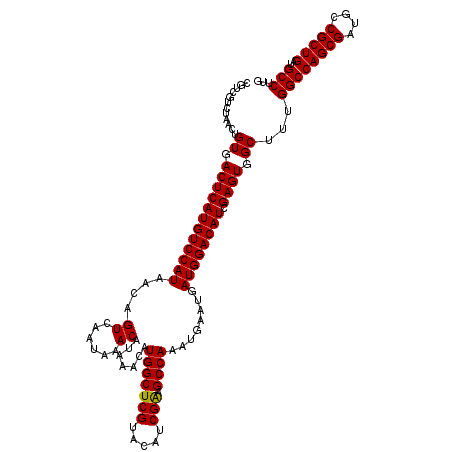
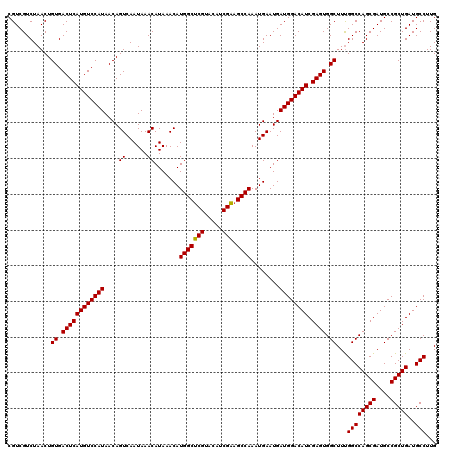
| Location | 22,320,877 – 22,320,997 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.33 |
| Mean single sequence MFE | -38.30 |
| Consensus MFE | -37.72 |
| Energy contribution | -37.50 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.98 |
| SVM decision value | 1.23 |
| SVM RNA-class probability | 0.933251 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22320877 120 - 23771897 CGUCGUCUAACUGUGACUCAUGUCCAUAACAGUCAAUAAACAUAAACAUGGCUCGUACAUCGAUGCCAAAUGAAUGAUGGACAUCGAGUGGCUUCGGCCAGCGAUGCCGCUGAUGCCUUG ............((.((((((((((((....((......)).......(((((((.....))).))))........)))))))).)))).))...((((((((....)))))..)))... ( -38.00) >DroSec_CAF1 35 120 - 1 CGUCGUCUAACUGUGACUCAUGUCCAUAACAGUCAAUAAACAUAAACAUGGCUCGUACAUCGGAGCCAAAUGAAUGAUGGACAUCGAGUGGCUUUGGCCAGCGAUGCCGCUGAUGCCUUG ............((.((((((((((((....((......)).......((((((........))))))........)))))))).)))).))...((((((((....)))))..)))... ( -39.20) >DroSim_CAF1 155 120 - 1 CGUCGUCUAACUGUGACUCAUGUCCAUAACAGUCAAUAAACAUAAACAUGGCUCGUACAUCGAAGCCAAAUGAAUGAUGGACAUCGAGUGGCUUUGGCCAGCGAUGCCGCUGAUGCCUUG ............((.((((((((((((....((......)).......(((((((.....)).)))))........)))))))).)))).))...((((((((....)))))..)))... ( -37.70) >consensus CGUCGUCUAACUGUGACUCAUGUCCAUAACAGUCAAUAAACAUAAACAUGGCUCGUACAUCGAAGCCAAAUGAAUGAUGGACAUCGAGUGGCUUUGGCCAGCGAUGCCGCUGAUGCCUUG ............((.((((((((((((....((......)).......(((((((.....))).))))........)))))))).)))).))...((((((((....)))))..)))... (-37.72 = -37.50 + -0.22)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:18:32 2006