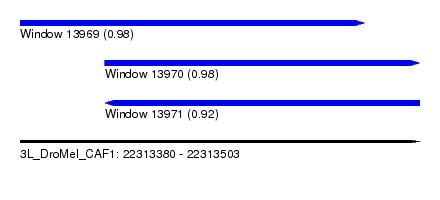

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 22,313,380 – 22,313,503 |
| Length | 123 |
| Max. P | 0.983542 |

| Location | 22,313,380 – 22,313,486 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 83.78 |
| Mean single sequence MFE | -30.62 |
| Consensus MFE | -22.52 |
| Energy contribution | -24.03 |
| Covariance contribution | 1.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.69 |
| Structure conservation index | 0.74 |
| SVM decision value | 1.92 |
| SVM RNA-class probability | 0.982643 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22313380 106 + 23771897 UCCCACG--------------GCCCCCAGCCCGAGAAACCCACUGCCUCUUGGCAUCCGCAGCCACAAAUUACAAAUUGAAAGUAGUUUGUAUCUGCUGUCUCUUCUGCUGCAGCAACUC .......--------------((...((((..((((.((....((((....))))...((((..(((((((((.........)))))))))..)))).)).))))..))))..))..... ( -30.10) >DroSec_CAF1 72092 90 + 1 UCCCACG--------------GCC----------------CACUGCCUCCUGGCAGCCGCAGCCACAAAUUACAAAUUGAAAGUAGUUUGUAUCUGCUGUCUCUUCUGCUGGAGCAACUC ......(--------------((.----------------...((((....)))))))((((..(((((((((.........)))))))))..))))((.((((......)))))).... ( -27.80) >DroEre_CAF1 70423 120 + 1 UCCCACGGCCCACGGCUCACGGCCCCCAGCCCGAGAAACCGACUGCCUCCUGGCUGCCGCAGCCACAAAUUACAAAUUGAAAGUAGUUUGUAUCUGCUGUCUCUUGUGCUGCAGCAACUC ......((((..........))))..((((.(((((....(((.(((....)))....((((..(((((((((.........)))))))))..)))).)))))))).))))......... ( -35.80) >DroYak_CAF1 50561 106 + 1 UCCCACG--------------GCCCCCAACCCGAGAAACCCACUGCCUCCUGGCAGCCGCAGCCACAAAUUACAAAUUGAAAGUAGUUUGUAUCUGCUGUCUCUUCUGCUGCAGCAACUC ......(--------------(........))((((......(((((....)))))..((((..(((((((((.........)))))))))..))))..))))...(((....))).... ( -28.80) >consensus UCCCACG______________GCCCCCAGCCCGAGAAACCCACUGCCUCCUGGCAGCCGCAGCCACAAAUUACAAAUUGAAAGUAGUUUGUAUCUGCUGUCUCUUCUGCUGCAGCAACUC ................................((((......(((((....)))))..((((..(((((((((.........)))))))))..))))..))))...(((....))).... (-22.52 = -24.03 + 1.50)

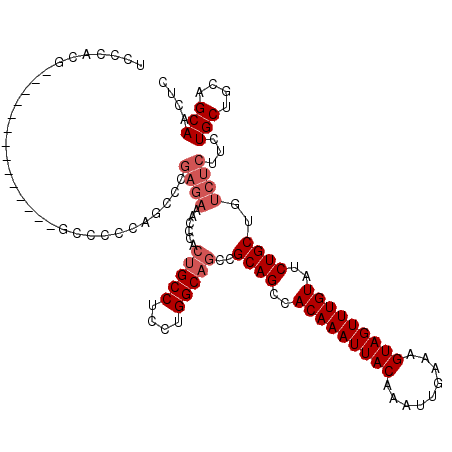
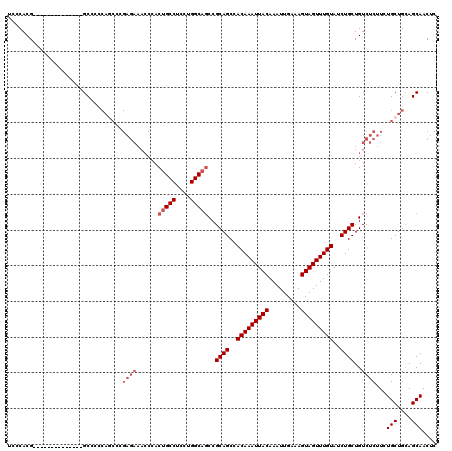
| Location | 22,313,406 – 22,313,503 |
|---|---|
| Length | 97 |
| Sequences | 4 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 96.91 |
| Mean single sequence MFE | -31.27 |
| Consensus MFE | -29.92 |
| Energy contribution | -29.93 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.42 |
| Structure conservation index | 0.96 |
| SVM decision value | 1.95 |
| SVM RNA-class probability | 0.983542 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22313406 97 + 23771897 CACUGCCUCUUGGCAUCCGCAGCCACAAAUUACAAAUUGAAAGUAGUUUGUAUCUGCUGUCUCUUCUGCUGCAGCAACUCCCGCCGGCAGUAAAAAA .((((((....(((....((((..(((((((((.........)))))))))..)))).........(((....)))......)))))))))...... ( -30.00) >DroSec_CAF1 72102 97 + 1 CACUGCCUCCUGGCAGCCGCAGCCACAAAUUACAAAUUGAAAGUAGUUUGUAUCUGCUGUCUCUUCUGCUGGAGCAACUCCCGCCGGCAGUAAAAAA .((((((....((((((...((..(((((((((.........)))))))))..))))))))......((.((((...)))).)).))))))...... ( -32.40) >DroEre_CAF1 70463 97 + 1 GACUGCCUCCUGGCUGCCGCAGCCACAAAUUACAAAUUGAAAGUAGUUUGUAUCUGCUGUCUCUUGUGCUGCAGCAACUCCCGCCGGCAGUAAAAAA .((((((....(((((...)))))(((((((((.........)))))))))...((((((..(....)..)))))).........))))))...... ( -31.70) >DroYak_CAF1 50587 97 + 1 CACUGCCUCCUGGCAGCCGCAGCCACAAAUUACAAAUUGAAAGUAGUUUGUAUCUGCUGUCUCUUCUGCUGCAGCAACUCCCGCCGGCAGUAAAAAA ..(((((....)))))..((((..(((((((((.........)))))))))..)))).........((((((.((.......))..))))))..... ( -31.00) >consensus CACUGCCUCCUGGCAGCCGCAGCCACAAAUUACAAAUUGAAAGUAGUUUGUAUCUGCUGUCUCUUCUGCUGCAGCAACUCCCGCCGGCAGUAAAAAA .((((((....(((....((((..(((((((((.........)))))))))..)))).........(((....)))......)))))))))...... (-29.92 = -29.93 + 0.00)

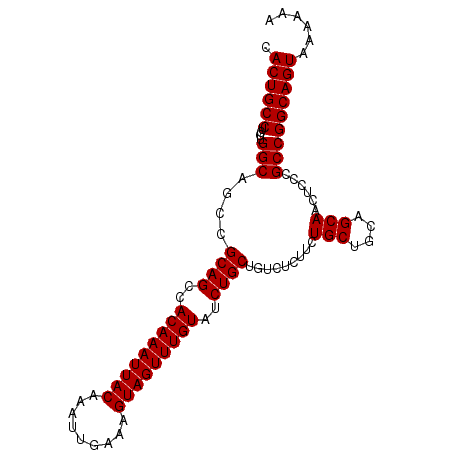
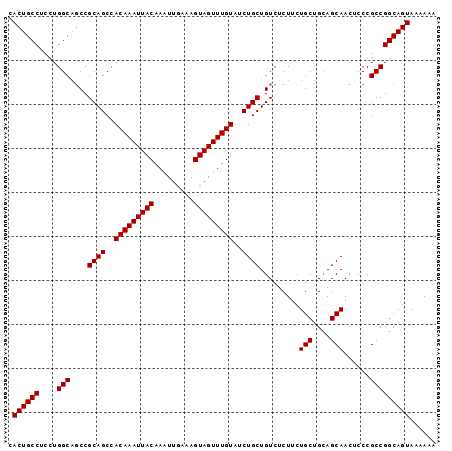
| Location | 22,313,406 – 22,313,503 |
|---|---|
| Length | 97 |
| Sequences | 4 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 96.91 |
| Mean single sequence MFE | -33.80 |
| Consensus MFE | -31.00 |
| Energy contribution | -31.50 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.33 |
| Structure conservation index | 0.92 |
| SVM decision value | 1.16 |
| SVM RNA-class probability | 0.924782 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22313406 97 - 23771897 UUUUUUACUGCCGGCGGGAGUUGCUGCAGCAGAAGAGACAGCAGAUACAAACUACUUUCAAUUUGUAAUUUGUGGCUGCGGAUGCCAAGAGGCAGUG .....(((((((....((.....(((((((..........(((((((((((..........))))).)))))).)))))))...))....))))))) ( -31.10) >DroSec_CAF1 72102 97 - 1 UUUUUUACUGCCGGCGGGAGUUGCUCCAGCAGAAGAGACAGCAGAUACAAACUACUUUCAAUUUGUAAUUUGUGGCUGCGGCUGCCAGGAGGCAGUG ((((((.((((.....((((...)))).))))))))))..((((.((((((.(((.........))).)))))).)))).((((((....)))))). ( -32.90) >DroEre_CAF1 70463 97 - 1 UUUUUUACUGCCGGCGGGAGUUGCUGCAGCACAAGAGACAGCAGAUACAAACUACUUUCAAUUUGUAAUUUGUGGCUGCGGCAGCCAGGAGGCAGUC ......((((((..(..(.((((((((((((((((..((((..((............))...))))..))))).))))))))))))..).)))))). ( -36.20) >DroYak_CAF1 50587 97 - 1 UUUUUUACUGCCGGCGGGAGUUGCUGCAGCAGAAGAGACAGCAGAUACAAACUACUUUCAAUUUGUAAUUUGUGGCUGCGGCUGCCAGGAGGCAGUG ((((((.((((..((((......)))).))))))))))..((((.((((((.(((.........))).)))))).)))).((((((....)))))). ( -35.00) >consensus UUUUUUACUGCCGGCGGGAGUUGCUGCAGCAGAAGAGACAGCAGAUACAAACUACUUUCAAUUUGUAAUUUGUGGCUGCGGCUGCCAGGAGGCAGUG ......((((((....((....((((((((..........(((((((((((..........))))).)))))).))))))))..))....)))))). (-31.00 = -31.50 + 0.50)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:18:08 2006